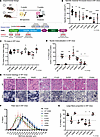
Angiotensin signaling is essential for stress erythropoiesis but causes retention of dysfunctional mitochondria in RBCs
Rai et al. report that angiotensin signaling sustains erythroid expansion but also promotes retention of dysfunctional mitochondria in erythroid cells during stress erythropoiesis. The cover image is a collage of nucleated (purple) and enucleated cells from wild-type and ATR1-deficient mice. Mitochondria are highlighted in green. Image credit: Parul Rai and Madison Rice.
In-Press Preview - More
Abstract
Ischemia/reperfusion (IR) enhances oxidative stress, leading to myocardial injury. Although Perm1 promotes cytoprotective mechanisms, the underlying mechanisms are poorly understood. Cysteine oxidation of Keap1 alleviates Cul3-mediated ubiquitination/degradation of Nrf2 and promotes antioxidant transcription. Here we show that Perm1 activates Nrf2 through cysteine oxidation of Keap1 and stabilization of Nrf2. Endogenous Perm1 was downregulated during IR, whereas the rescue of Perm1 reduced IR injury. Downregulation of Perm1 exacerbated oxidative stress, whereas upregulation of Perm1 alleviated it, accompanied by downregulation and upregulation of Nrf2-regulated antioxidant genes, respectively. Perm1 promoted oxidation of cysteine residues in Keap1, possibly through thiol-disulfide exchange reactions, which decreases Keap1-Nrf2 interaction and inhibits Cul3-mediated degradation of Nrf2. We identified Cys121 and Cys746 in Perm1 as critical for Keap1 oxidation and cardioprotection. Thus, Perm1 induces cysteine oxidation of Keap1, thereby conferring myocardial resistance to IR injury by inducing Nrf2 stabilization and transcriptional activation of antioxidant genes.
Authors
Shin-ichi Oka, Chun-Yang Huang, Masato Matsushita, Allen Sam Titus, Yasuki Nakada, Risa Mukai, Samta Veera, Youssef Mourad, Ghassan Yehia, Peter Romanienko, Yimin Tian, Peiyong Zhai, Junichi Sadoshima
Abstract
Kawasaki disease (KD) is an acute febrile systemic vasculitis of unknown etiology and the leading cause of acquired heart disease among children. Complement activation has long been observed in patients with acute KD, however, its contribution to disease development remains unknown. Here, using publicly available datasets, we showed that patients with acute KD exhibited higher expression of complement products in whole blood, consistent with the activation of the complement pathway. Similarly, in the Lactobacillus casei cell wall extract (LCWE) murine model of KD, LCWE injection induced increased expression of complement products in cardiovascular tissues, suggestive of activation of the complement pathways. C3-deficient mice or WT mice treated with the complement C5a Receptor 1 (C5ar1) antagonist developed significantly more severe LCWE-induced cardiovascular lesions and vasculitis. Furthermore, we observed that LCWE binds to serum C3, an opsonizing factor that labels microbial targets for clearance, and LCWE deposition in the liver was significantly higher in C3-deficient mice compared to WT mice. Overall, our data indicate that blocking the complement system significantly exacerbates LCWE-induced KD vasculitis, likely by impairing C3-mediated clearance of LCWE. These data suggest that the complement pathway may play a protective role in KD pathogenesis by promoting clearance of potential bacterial or viral trigger of KD.
Authors
Asli E. Atici, Begüm Kocatürk, Benjamin L. Ross, Emily A. Aubuchon, Rebecca A. Porritt, Thacyana T. Carvalho, Takahiro Namba, Youngho Lee, Magali Noval Rivas, Moshe Arditi
Abstract
Antibody production by B cells has emerged as an important factor in regulating anti-tumor immunity with both suppressive and promotive roles in cancer. However, the specific impact of antibody deficiency during development of pancreatic ductal adenocarcinoma (PDAC) has not been explored. To address this question, we crossed the well-established KPC mouse model to mice lacking all circulating immunoglobulin (Ig) due to genetic ablation of both Ig secretion and Ig class switching (KPC-μSAID mice). KPC-μSAID mice exhibited a two-fold acceleration in tumor formation, a two-fold reduction in median survival, and increased liver metastases versus KPC-WT control mice. Immunofluorescence analysis of pancreatic tissues from antibody-sufficient KC- and KPC-WT mice showed that IgG was predominantly localized within extracellular matrix (ECM). Furthermore, in both KC- and KPC-μSAID mice, ECM density and podoplanin+ cancer-associated fibroblasts (CAFs) were significantly reduced. In the KPC-μSAID tumor microenvironment (TME), intratumoral myeloid-derived suppressor cells (MDSC) were also increased, while CD4+ and CD8+ T cells decreased, relative to tumor-bearing KPC-WT mice, with macrophage exhibiting a mixed polarization phenotype. These findings were recapitulated in antibody-subclass-deficient, KPC-AID mice, suggesting a potentially novel function of IgG in suppressing PDAC progression, by directly or indirectly regulating pancreatic fibrosis and the density of the ECM.
Authors
Jeremy B. Foote, Sujith Sarvesh, Sameer Al Diffalha, David K. Crossman, Changde Cheng, Myng-Hee Kim, Cherlene Hardy, Julienne L. Carstens, Kyoko Kojima, Bart J. Rose, Christopher A. Klug
Abstract
Because older donor age is a major concern when considering kidneys for potential transplantation, we explored the actual impact of donor age on the features of kidneys that have been transplanted. We studied the correlations of donor age with molecular injury and rejection scores in 4502 kidney transplant biopsies assessed by microarrays, as well as function and postbiopsy survival. We used multivariable analyses to correct for the correlations of donor age with other predictive variables: recipient age, time of biopsy posttransplant, and deceased vs. living donors. Older donor age correlated with lower GFR and increased acute and chronic injury transcripts, but had no effect on rejection, which anti-correlated with recipient age. Acute injury transcripts peaked immediately posttransplant and regressed. Older donor age had little effect on acute molecular injury immediately posttransplant but strongly increased molecular injury scores at later times, peaking about 1-year posttransplant, indicating that older age does not increase molecular injury but increases failed repair post-injury. As expected, older donor age correlated with increased chronic injury and lower GFR, evident from the earliest time posttransplant, pre-transplant aging. However, despite significant age-related effects, the quantitative contribution of donor aging to molecular injury, function, and survival was very small.
Authors
Katelynn Madill-Thomsen, Martina Mackova, Jessica Chang, Enver Akalin, Tarek Alhamad, Sanjiv Anand, Miha Arnol, Rajendra Baliga, Mirosław Banasik, Christopher Blosser, Georg Böhmig, Daniel Brennan, Jonathan Bromberg, Klemens Budde, Andrzej Chamienia, Kevin V Chow, Michał Ciszek, Declan de Freitas, Dominika Dęborska-Materkowska, Alicja Dębska-Ślizień, Arjang Djamali, Leszek Domański, Magdalena Durlik, Gunilla Einecke, Farsad Eskandary, Richard Fatica, Iman Bajjoka-Francis, Justyna Fryc, John Gill, Jagbir Gill, Maciej Glyda, Sita Gourishankar, Marta Gryczman, Gaurav Gupta, Petra Hruba, Peter Hughes, Arskarapuk Jittirat, Zeljka Jurekovic, Layla Kamal, Mahmoud Kamel, Sam Kant, Nika Kojc, Joanna Konopa, James Lan, Roslyn Mannon, Arthur Matas, Joanna Mazurkiewicz, Marius Miglinas, Thomas Mueller, Marek Myślak, Beata Naumnik, Anita Patel, Agnieszka Perkowska-Ptasińska, Michael Picton, Grzegorz Piecha, Emillio Poggio, Silvie Rajnochova Bloudickova, Thomas Schachtner, Sung Shin, Soroush Shojai, Majid Sikosana, Janka Slatinská, Katarzyna Smykal-Jankowiak, Ashish Solanki, Zeljka Veceric Haler, Ondrej Viklicky, Ksenija Vucur Simic, Matthew R. Weir, Andrzej Wiecek, Zbigniew Włodarczyk, Ziad Zaky, Philip F. Halloran
Abstract
Skeletal muscle pathology is a critical but poorly understood contributor to neuromuscular degeneration in spinal and bulbar muscular atrophy (SBMA), a CAG/polyglutamine (polyQ) expansion disorder caused by mutation in the androgen receptor (AR). Using a gene-targeted SBMA mouse model, we applied single-nucleus RNA sequencing to identify a disease-specific population of skeletal muscle myonuclei that replaced normal myonuclear subtypes. This transition was associated with dysregulation of the pathway governed by PGC-1α, a central regulator of myofiber specification and metabolic identity. PGC-1α dysfunction in SBMA muscle was age-, hormone-, and polyQ length–dependent and was partially rescued by subcutaneous delivery of AR-targeted antisense oligonucleotides. Integrated ChIP-seq and RNA-seq analyses revealed that aberrant PGC-1α activity promoted the expression of a distinct set of myofiber specification genes while downregulating those that define healthy Type IIb and Type IIx myonuclei. We propose a model in which this dysfunction arose downstream of polyQ-mediated sequestration of PGC-1α cofactors MEF2, CREB, and CBP, leading to transcriptional reprogramming and cellular dysfunction. These findings implicated PGC-1α dysregulation as a key event linking AR polyQ expansion to skeletal muscle degeneration and suggested a shared mechanism for polyQ-mediated muscle pathology across related neurodegenerative diseases.
Authors
Curtis J. Kuo, Laura B. Chopp, Zhigang Yu, Luhan Ni, Hien T. Zhao, Janghoo Lim, Andrew P. Lieberman