Virology
Abstract
Authors
Natalie E. Andresen, Heehwa G. Son, Joongho J. Joh, Shadmehr Demehri
Abstract
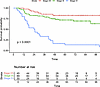
Merkel cell carcinoma (MCC) is a neuroendocrine carcinoma of the skin characterized by poor prognosis. This study aimed to explore the relationship between genetic alterations, tumor mutational burden (TMB), and MCC-specific survival (MCC-SS) in patients who underwent genomic profiling of tumors with OncoPanel. Univariate and multivariable analysis were used to assess the impact of genetic alterations on MCC-SS. Of the 188 identified patients, 164 were included in the analysis. The cohort had a mean age of 72.4 years (SD = 11.03), including 61.6% male. The median TMB was 5.32 (IQR = 3.04–25.53). Kaplan-Meier curves by high versus low TMB were significantly different (log-rank test, P = 0.017). PIK3CA (adjusted P = 0.003), SETBP1 (adjusted P = 0.002), KDR (adjusted P = 0.028), and RET (adjusted P = 0.033) were selected for multivariable analysis. In the multivariable regressions, only PIK3CA (HR = 2.07 [95% CI, 1.10–3.88]; P = 0.024) remained significant. PIK3CA remained significant across prespecified sensitivity analyses. In this study, high TMB and PIK3CA alterations were associated with poor MCC-SS. Identifying a higher-risk subgroup may inform risk stratification and motivate further evaluation of PI3K pathway targeting in future studies.
Authors
Matheus Lobo, Furkan Bahar, Julia L. Schnabel, Joao P. Duprat Neto, Aniket Shetty, Karam Khaddour, Manisha Thakuria, Ann W. Silk, James A. DeCaprio
Abstract
Antiretroviral therapy (ART) has prolonged the life expectancy of persons living with HIV, the majority of whom are now older than 50 years. Aging people with HIV are at increased risk for cardiovascular events driven by HIV-related inflammation and hypercoagulation. Apixaban is a factor Xa inhibitor that reduces cardiovascular risks and treats stroke, deep vein thrombosis, and pulmonary embolism. We assessed apixaban’s impact on key parameters of HIV/SIV pathogenesis in SIV-infected, aged rhesus macaques (RMs) receiving ART. Inflammation, coagulation, T cell subsets, B cells, and macrophages and their immune activation status were monitored throughout the study. We found no significant differences between the apixaban-treated and control groups for virus replication or CD4+ T cell recovery in blood and tissues after ART. Apixaban did not significantly affect D-dimer, immune activation, or inflammation of SIV-infected, ART-treated RMs. Apixaban-treated RMs experienced multiple bleeding episodes, tissue hemorrhages, and myocardial infarctions, as demonstrated by pathological examination of necropsy-collected tissues. Given apixaban’s lack of effect on immune activation, CD4+ T cell restoration, and inflammation, along with increased risk of hemorrhage, factor Xa inhibition may not be an efficient or safe option to target and prevent cardiovascular events in aging people with HIV.
Authors
Cuiling Xu, Haritha Annapureddy, Lilly Carson, Vansh Khurana, Ranjit Sivanandham, Sindhuja Sivanandham, Tianyu He, Kevin D. Raehtz, Janet Kim, Christie Biber, Norma Arbujas-Silva, Mohammed Daira, Sudhapriya Kandasamy, Matthew J. Feinstein, Irini Sereti, Cristian Apetrei, Ivona Pandrea
Abstract
Dengue virus (DENV) vaccines should be designed to induce balanced protective immunity against all four dengue serotype to mitigate the risk of vaccine-mediated enhanced dengue disease. The first tetravalent vaccine (Dengvaxia) tested in humans was efficacious in children who were partially immune to DENV at baseline. In DENV-naive children, the vaccine was not efficacious and placed some naïve children at risk of experiencing more severe wild-type DENV breakthrough infections. To define dengue vaccine responses at the individual subject level and their relationship to mild and severe dengue infections, we prospectively studied a cohort of DENV-naive children who received one dose of Dengvaxia. The vaccine stimulated variable responses that neutralized 0, 1 (monotypic), or 2+ (multitypic) serotypes in individual children. We used a logistic regression model to evaluate whether vaccine status and serotype-specific NAb status at the end of study period 1 influenced the probability of experiencing a virologically confirmed dengue disease (VCD) case thereafter (months 20 - 60). Vaccinated children with NAb response to only one serotype were at greater risk of being a case compared to the DENV-naïve control group (Odds Ratio 5.07). This risk was not observed in vaccinated children with no NAb or NAb to 2 or more serotypes. We propose that individuals with durable NAb to one serotype have an abundance of serotype cross-reactive, non-neutralizing Abs implicated in the enhanced replication of heterologous serotypes. We discuss the implications of our findings for flagging vaccine candidates that are likely to pose a special risk to seronegative subjects.
Authors
Laura J. White, Lindsay D. Hein, Maria Abad Fernandez, Cameron Adams, Elizabeth Adams, Emily Freeman, Ruby Shah, Lakshmanane Premkumar, Kristal An Agrupis, Maria Vinna Crisostomo, Jedas Veronica Daag, Michelle Ylade, Jacqueline Deen, Ana Lena Lopez, Leah Katzelnick, Aravinda M. de Silva
Abstract
Introduction: BK polyomavirus (BKV) infection is associated with injury and subsequent graft loss due to the extent of injury or rejection. However, the molecular mechanisms driving injury and subsequent adverse outcomes remain poorly understood. Methods: In a cross-sectional study, single-cell RNA sequencing from kidney allograft biopsies was used to assess cell type-specific responses between uninfected controls and two distinct phases of BKV infection: peaking (increasing viral blood titers) and resolving (decreasing viral titers following immunosuppression reduction). Results: Genes upregulated in BK viral nephropathy (BKVN) were enriched for polyomavirus infection hallmarks, including ribosome biogenesis, translation, and energy restructuring. Additionally enriched pathways included wound healing, cellular stress, antigen presentation and immune signaling. Even without BKVN (peaking BK viremia alone), epithelial cells expressed signatures for wound healing, cellular stress, and extracellular matrix remodeling. In vivo tubular cell responses at single-cell resolution were validated against single cell transcriptomic data of BKV infected cells in a cell culture model. Despite similarities, in vivo tubular cells underwent metabolic adaptation favoring fatty acid oxidation and proinflammatory responses not observed in culture models likely due to an absent innate and adaptive immune system. Despite lymphopenia and immunosuppressive therapies, the proportion of recipient derived intrarenal adaptive immune cells was increased in biopsies associated with peaking viremia alongside activation of innate immune responses. Adaptive immune cells exhibited persistent inflammatory signaling and remodeling of energy metabolism during the resolving phase of infection. Conclusion: These not previously reported insights into BKV-associated injury may have implications for clinical management and improved allograft outcomes.
Authors
Tess Marvin, Rachel Sealfon, Phillip J. McCown, Fadhl AlAkwaa, Evan A. Farkash, Edgar A. Otto, Felix Eichinger, Ping An, Rajasree Menon, Celine C. Berthier, Tavis J. Reed, Paula Arrowsmith, Lalita Subramanian, Kelly J. Shaffer, Silas P. Norman, Ramnika Gumber, Michael J. Imperiale, James M. Pipas, Olga G. Troyanskaya, Matthias Kretzler, Chandra L. Theesfeld, Abhijit S. Naik
Abstract
How SARS-CoV-2 causes a wide range of clinical manifestations and disease severity remains poorly understood. SARS-CoV-2 encodes 2 proteases (3CLPro and PLPro), vital for viral production, but also promiscuous with respect to host protein targets. Pharmacological inhibition of 3CLPro markedly reduced hospitalization and death in Phase 2/3 clinical studies. Here, we develop a bioinformatic algorithm, leveraging experimental data from SARS-CoV, to predict host cleavage targets of 3CLPro. We capture targets of 3CLPro described previously for SARS-CoV-2, as well as thousands of putative targets. We validate numerous targets cleaved during infection, including the giant sarcomeric protein obscurin and the innate immune protein OAS1. A long form of OAS1, p46, has been associated in numerous GWAS studies with lesser COVID disease severity. We show that 3CLPro cleaves p46 OAS1 immediately upstream of a known prenylation domain, relocalizing OAS1 from subcellular membranes to the cytosol, rendering it akin to the nonprotective, cytosolic p42 isoform. Similar OAS1 relocalization occurs upon infection by SARS-CoV-2. Our data provide a high-throughput resource to identify putative host cleavage targets of 3CLPro and reveal a mechanism by which SARS-CoV-2 antagonizes host innate immunity in individuals with the protective p46 isoform of OAS1.
Authors
Nora Yucel, Silvia Marchiano, Evan Tchelepi, Germana Paterlini, Ivan A. Kuznetsov, Kristina Li, Quentin McAfee, Nehaar Nimmagadda, Andy Ren, Sam Shi, Alyssa Grogan, Aikaterini Kontrogianni-Konstantopoulos, Charles Murry, Zoltan Arany
Abstract
Antibody-dependent enhancement (ADE) of infection is a well-described phenomenon for several viruses, including dengue, Ebola, respiratory syncytial virus, and HIV. ADE occurs when virus-antibody complexes engage Fc receptors (FcRs) and virus-specific receptors, enhancing infection under conditions of incomplete neutralization. The Coronavirus Immunotherapeutic Consortium (CoVIC) assembled a comprehensive dataset of functional properties for over 400 mAbs, enabling direct comparison of neutralization, Fc-mediated functions, receptor binding, and infection of immune cells. Infection rates in most primary human immune cell types were low, with modest increases observed for some mAbs. In contrast, macrophages were more susceptible to SARS-CoV-2 and exhibited substantial ADE with select mAbs. ADE was completely inhibited by FcR blockade and significantly reduced by antibody- or ceftazidime-mediated blocking of angiotensin-converting enzyme 2 (ACE2). Neutralization potency did not correlate with ADE, as both strongly and weakly neutralizing antibodies induced enhancement. Instead, ADE magnitude depended on an antibody’s ability to block spike protein binding to ACE2. Importantly, ADE resulted in productive infection with release of infectious virus. Evaluation of antibodies against the BA.1 (Omicron) variant revealed reduced or lost ADE for most mAbs, with increased ADE observed for several mAbs relative to the USA-WA1/2020 strain.
Authors
Natalia A. Kuzmina, Sivakumar Periasamy, Kritika Kedarinath, Keziah Hernandez, Caroline Atyeo, S. Moses Dennison, Kan Li, Daniel Bedinger, Sharon L. Schendel, Georgia D. Tomaras, Hanif Ali, Galit Alter, Erica Ollmann Saphire, Alexander Bukreyev
Abstract
Recombinant hepatitis B surface antigen (rHBsAg) vaccine with various adjuvants fails to break T and B cell tolerance in hosts with chronic hepatitis B (CHB). This study aims to explore the mechanisms to break immune tolerance that allows the host to respond to rHBsAg, achieving a cure for CHB. We engineered an anti–PD-L1–IFN-α (aPD-L1–IFN-α) heterodimeric fusion protein to allow rHBsAg to rejuvenate T and B cell responses in hepatitis B virus–tolerant (HBV-tolerant) mice. S.c. coimmunization with aPD-L1–IFN-α and rHBsAg significantly enhanced antigen uptake and maturation of both macrophage and dendritic cell (DC) subsets in draining lymph nodes. Macrophages drove early B cell activation, while cDC1s primed CD8+ T cells, breaking tolerance and leading to both B cell and cytotoxic T lymphocyte (CTL) differentiation. This strategy elicited not only anti-HBsAg neutralizing antibodies but also HBsAg-specific CD8+ T cell responses, achieving a functional cure without systemic toxicity. The efficacy of the aPD-L1–IFN-α adjuvant depended on both PD-L1 cis-targeting and IFN-α receptor signaling in antigen-presenting cells. These findings establish aPD-L1–IFN-α as a translatable adjuvant to break the strong tolerance induced by CHB, providing a dual-pathway strategy to induce HBV-specific T and B cell responses.
Authors
Chao-Yang Meng, Yong Liang, Longxin Xu, Hongjia Li, Jingya Guo, Hairong Xu, Fan Wang, Yang-Xin Fu, Hua Peng
Abstract
BACKGROUND Among people living with HIV (PLWH), immunological nonresponders (INR) fail to adequately restore CD4+ T cell counts despite effective antiretroviral therapy (ART), placing them at greater risk for adverse outcomes and reduced vaccine efficacy. We aimed to study the robustness and longevity of vaccine-induced virus-specific cellular immune responses in INR.METHODS Virus-specific CD8+ T cell responses were analyzed in INR (CD4+ T cell count < 300 cells/μL) and immunological responders (IR) (CD4+ T cell count > 500 cells/μL), receiving ART, and HIV-uninfected controls following COVID-19 mRNA vaccination and infection. Virus-specific CD8+ T cells were characterized using peptide-loaded MHC I tetramer technology, after in vitro expansion and cytokine production assays. Virus-specific CD4+ T cells and IgG levels were determined by activation-induced marker (AIM) assay and ELISA, respectively.RESULTS We demonstrated that, while long-lasting virus-specific cellular immune responses were generated in INR, CD8+ T cell immunity remained limited compared with robust CD4+ T cell reactivity. CD8+ T cell responses in INR exhibited reduced breadth and frequency, accompanied by altered memory differentiation and suboptimal activation and effector response upon antigen exposure. This deficiency correlated with low CD4+ T cell counts, independent of other disease markers, highlighting the pivotal role of CD4+ T cells in orchestrating vaccine-induced immunity. Notably, repeated booster vaccinations enhanced virus-specific CD8+ T cell responses.CONCLUSION INR elicit limited vaccine-induced virus-specific CD8+ T cell immunity, but booster vaccinations can enhance these responses, suggesting better immune outcomes with tailored vaccination strategies.FUNDING Helmholtz Society, German Research Foundation, Federal Ministry of Education and Research.
Authors
Vivien Karl, Anne Graeser, Anastasia Kremser, Liane Bauersfeld, Florian Emmerich, Nadine Herkt, Siegbert Rieg, Susanne Usadel, Bertram Bengsch, Tobias Boettler, Hendrik Luxenburger, Christoph Neumann-Haefelin, Matthias C. Müller, Robert Thimme, Maike Hofmann
Abstract
Elite controllers (ECs) maintain undetectable levels of plasma viremia in the absence of treatment, but small reservoirs of replication-competent proviruses persist in the vast majority of these persons. We longitudinally studied paired blood and inguinal lymph node samples (LNMC) from two ECs to better characterize distinguishing features of viral reservoir cell dynamics in ECs. In both participants, we observed a 7- to 10-fold lower frequency of intact proviruses in LNMC samples relative to reservoir cells circulating in blood. The landscape of intact proviruses in both tissue compartments was dominated by shared large clones that were frequently integrated in non-coding DNA regions, but the frequency and diversity of intact proviruses was more limited in LNMCs. Of note, over 9-10 years of longitudinal follow-up, a 3- to 18-fold reduction of intact proviruses was observed. Together, these data support the hypothesis that viral reservoirs in ECs’ blood and lymphoid tissues are under strong, likely immune-mediated selection pressure.
Authors
Samantha K. Marzi, Chloé M. Naasz, Leah Carrere, Carmen Gasca-Capote, Isabelle C. Roseto, Ce Gao, Matthias Cavassini, Andrea Mastrangelo, Mathias Lichterfeld, Matthieu Perreau, Xu G. Yu
No posts were found with this tag.