Immunology
Citation Information: JCI Insight. 2025. https://doi.org/10.1172/jci.insight.199679.
Abstract
The two main subgroups of autoimmune myasthenia gravis, a neuromuscular junction disorder associated with muscle weakness, are the early and late-onset forms, defined by onset before or after 50 years of age. Both carry acetylcholine-receptor autoantibodies, but differ in sex ratios, genetics and occurrence of disease-specific thymus inflammation. By applying multimodal techniques, including deep spectral cytometric phenotyping and single cell sequencing to peripheral blood and thymic lymphocyte samples we explored the possibility to discriminate the two forms by cellular immune phenotyping. Analyzing two independent cohorts we identified distinct immunological differences driven by three main lymphocyte populations. Lower frequencies of mucosa-associated invariant T cells and naïve CD8 T cells were observed in late-onset myasthenia, suggesting enhanced immune senescence. Further, a highly differentiated, canonical natural killer cell population was reduced in early-onset myasthenia, which was negatively correlated with the degree of thymic inflammation. Using only the frequency of these three populations, correct myasthenia subgroup assignment could be predicted with an accuracy of 90%. The NK cell population negatively associated to early-onset disease had a similar association to thymic hyperlasia, whereas the two T-cell populations point to enhanced immune senescence in late-onset myasthenia gravis. These distinct immunocellular endophenotypes for early- and late onset disease suggest differences in the immunopathogenic processes. Together with demographic factors and other disease subgroup-specific features, the frequency of the identified cell subpopulations may improve clinical classification, in turn of relevance for channeling to interventions.
Authors
Jakob Theorell, Nicolas Ruffin, Andrew Fower, Chiara Sorini, Philip Ambrose, Valentina Damato, Lahiru Handunnetthi, Isabel Leite, Sarosh R. Irani, Susanna Brauner, Adam E. Handel, Fredrik Piehl
Citation Information: JCI Insight. 2025. https://doi.org/10.1172/jci.insight.198203.
Abstract
Immune checkpoint inhibitors (ICIs) such as anti-PD-1 and anti-CTLA-4 antibodies are used to induce an immune response against many types of tumors. However, ICIs often also induce autoimmune responses, referred to as immune-related adverse events (irAEs), which occur unpredictably and at varying levels of severity in ICI-treated patients. The immunologic factors that predispose patients to the development of severe irAE are largely unclear. Here, we utilized high dimensional mass cytometry immunophenotyping of longitudinal blood samples from patients with metastatic melanoma treated with combination anti-PD-1/CTLA4 ICI therapy in the context of a clinical trial to characterize alterations in immune profiles induced by combination ICI therapy and to identify immune features associated with development of severe irAEs. Deep T cell profiling highlighted that ICI therapy induces prominent expansions of activated, CD38hi CD4+ and CD8+ T cells, which are frequently bound by the therapeutic anti-PD-1 antibody, as well as substantial changes in regulatory T cell phenotypes. However, neither the baseline frequency nor the extent of expansion of these cell populations was associated with development of severe irAEs. Rather, single cell-association testing revealed naïve CD4+ T cell abundance pre-treatment as significantly associated with the development of severe irAEs. Biaxial gating of naïve CD4+ T cells confirmed a significant positive association of naïve CD4+ T cell proportion and development of a severe irAE and with the number of irAEs developed in this cohort. Results from this broad profiling study indicate the abundance of naïve CD4+ T cells as a predictive feature for the development of severe irAEs following combination anti-PD-1/CTLA4 ICI therapy.
Authors
Kathryne E. Marks, Alice Horisberger, Mehreen Elahee, Ifeoluwakiisi A. Adejoorin, Nilasha Ghosh, Michael A. Postow, Laura Donlin, Anne R. Bass, Deepak A. Rao
Citation Information: JCI Insight. 2025. https://doi.org/10.1172/jci.insight.196605.
Abstract
Metabolic inflammation is closely linked to dynamic changes in circulating monocyte populations, yet how nutritional signals regulate this process remains unclear. ANGPTL8, a hepatokine rapidly induced by refeeding, emerged as a key regulator of postprandial monocyte dynamics. We examined ANGPTL8 expression in human and murine fasting-refeeding models and manipulated ANGPTL8 expression specifically in hepatocytes to assess its role in metabolic inflammation and insulin resistance in obese mice. ANGPTL8 overexpression elevated circulating monocytes and proinflammatory cytokines, while its deletion reduced these parameters and conferred metabolic benefits. Mechanistically, recombinant ANGPTL8 stimulated CCL5 production in bone marrow-derived macrophages via P38 signaling activation, promoting monocyte recruitment and proinflammatory macrophage polarization. These effects were mitigated by CCR5 antagonism. Rescue experiments demonstrated that CCL5 supplementation in Angptl8-deficient mice restored monocyte levels and inflammatory responses. Functionally, ANGPTL8 worsened insulin resistance and glucose intolerance in obese mice, effects that were reversed by its deletion and recapitulated by CCL5 administration. These findings suggest that ANGPTL8 functions as a nutritional checkpoint linking feeding status to monocyte-mediated inflammation through the CCL5-CCR5 axis. By driving monocytosis and proinflammatory macrophage activation, ANGPTL8 exacerbates metabolic dysfunction. Targeting the ANGPTL8-CCL5-CCR5 pathway may therefore offer a promising therapeutic strategy for managing obesity-related metabolic diseases.
Authors
Ran-Ran Kan, Si-Yi Wang, Xiao-Yu Meng, Li Huang, Yu-Xi Xiang, Bei-Bei Mao, Hua-Jie Zou, Ya-Ming Guo, Li-Meng Pan, Pei-Qiong Luo, Yan Yang, Zhe-Long Liu, De-Lin Ma, Wen-Jun Li, Yong Chen, Dan-Pei Li, Xue-Feng Yu
Citation Information: JCI Insight. 2025. https://doi.org/10.1172/jci.insight.198743.
Abstract
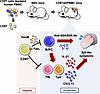
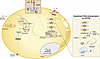
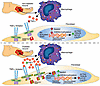
Cytomegalovirus (CMV) is a prevalent β-herpesvirus that persists asymptomatically in immunocompetent hosts. In people with HIV-1 (PWH), CMV is associated with HIV-1 persistence and particular inflammatory-related co-morbidities. The true causative role of CMV in HIV-associated pathologies however remains unclear given that nearly all PWH are coinfected with CMV. In this study, we examined acute phase immune and virological dynamics in cohorts of SIV-infected rhesus macaques (RMs) that were naturally seropositive or -negative for rhesus CMV (RhCMV). We observed prior to SIV, RhCMV expanded a polyclonal population of target CCR5+CD4+ T cells in gut and lymph nodes (LN) that expressed the chemotactic receptor CXCR3 and were largely not specific for RhCMV. Upon SIV infection, RhCMV+ RMs exhibited higher peak viremia and elevated levels of SIV DNA in the upper and lower intestine. Greater seeding of SIV DNA was associated with a maintenance of CCR5-expressing CD4+ T cells that were enriched within the RhCMV+ gut along a CXCR3-CXCL9 chemotactic axis. Overall, the data suggest that RhCMV can promote SIV susceptibility within a diverse, polyclonal pool of CD4 T cells that are not entirely RhCMV-specific.
Authors
Chrysostomos Perdios, Naveen Suresh Babu, Celeste D. Coleman, Anna T. Brown, Shevon N. Alexander, Matilda J. Moström, Carolina Allers, Lara Doyle-Meyers, Christine M. Fennessey, Lori A. Rowe, Brandon F. Keele, Amitinder Kaur, Michael L. Freeman, Joseph C. Mudd
Citation Information: JCI Insight. 2025;10(22):e191700. https://doi.org/10.1172/jci.insight.191700.
Abstract
Peripheral helper T (Tph) and follicular helper T (Tfh) cells are key regulators of B cell differentiation and antibody production, making them promising targets for autoimmune disease treatment. However, their differentiation mechanisms differ significantly between humans and mice, limiting drug validation in mouse models. Here, we present a simple and effective method for in vivo proliferation of human Tph/Tfh and B cells. We discovered that after depleting CD8+ T cells of human peripheral blood mononuclear cell–transferred immunodeficient mice (CD8TΔhPBMC mice), human Tph/Tfh cells and B cells proliferated markedly in the spleen compared with those in human PBMC–transferred immunodeficient mice (hPBMC mice). Transcriptome analysis confirmed proliferating cells’ close resemblance to human Tph/Tfh cells. Furthermore, multicolor flow cytometry revealed CXCL13+ Tph cells infiltrating Sjögren’s syndrome–associated (SjS-associated) organs, such as salivary glands. Single-cell RNA sequencing identified IL-21+CXCL13+IFN-γ+ICOS+TIGIT+GPR56+ Tph cells in the salivary glands. These findings are consistent with reduced saliva volume and elevated SjS markers, such as anti-SSA antibody, in these mice, which were both ameliorated by immunosuppressants. In vitro, CD8+ T cells from hPBMC mice induced B cell apoptosis and inhibited Tph/Tfh differentiation. This model advances understanding of human Tph/Tfh cell biology and offers a valuable platform for studying SjS and therapeutic targets.
Authors
Mariam Piruzyan, Sota Fujimori, Ryota Sato, Yuki Imura, Sachiko Mochiduki, Kana Takemoto, Akiko Nishidate, Yuzo Koda
Citation Information: JCI Insight. 2025;10(22):e185914. https://doi.org/10.1172/jci.insight.185914.
Abstract
Processes that promote white adipocyte inflammatory function remain incompletely defined. Here, we demonstrated that type I interferon–dependent (IFN-I–dependent) skewing of adipocyte glycolysis, nicotinamide adenine dinucleotide (NAD+) utilization, and pyruvate kinase isozyme M2 (PKM2) function may contribute to increased systemic and tissue inflammation and disease severity in obesity. Notably, chemical and/or genetic inhibition of glycolysis, the NAD+ salvage pathway, or PKM2 restricted IFN-I–dependent increase in adipocyte inflammatory cytokine production. Further, genetic or small molecule targeting of PKM2 function in vivo was sufficient to reduce systemic and tissue inflammation and metabolic disease severity in obese mice, in an adipocyte PKM2-dependent manner. Further, white adipose tissue of individuals living with obesity and metabolic disease, compared with metabolically healthy individuals with obesity, showed an increase in expression of inflammatory and metabolic genes, while small molecule targeting of PKM2 function contributed to reduced IFN-I–driven inflammatory cytokine production by primary human adipocytes. Together, our findings invoke the IFN-I/PKM2 axis as a potential target for modulating adipocyte dysregulated inflammation.
Authors
Michelle S.M.A. Damen, Pablo C. Alarcon, Calvin C. Chan, Traci E. Stankiewicz, Hak Chung, Keisuke Sawada, Cassidy J. Ulanowicz, John Eom, Jarren R. Oates, Jennifer L. Wayland, Jessica R. Doll, Rajib Mukherjee, Miki Watanabe-Chailland, Lindsey Romick-Rosendale, Sara Szabo, Michael A. Helmrath, Joan Sanchez-Gurmaches, Maria E. Moreno-Fernandez, Senad Divanovic
Citation Information: JCI Insight. 2025;10(22):e190836. https://doi.org/10.1172/jci.insight.190836.
Abstract
Fibroblast to myofibroblast transition is a critical event required for effective tissue repair. In pathologic wound repair processes, such as type 2 diabetes (T2D), fibroblast to myofibroblast transition is impaired. The exact factors that control this transition in wounds are unclear. Here, using human tissue and murine transgenic models, we show that the histone methyltransferase SETDB2 is elevated in diabetic wound fibroblasts and TNF-α represses fibroblast to myofibroblast transition via Setdb2. We identified that TNF-α increases Setdb2 in fibroblasts via a JAK1,3/STAT3 signaling pathway, where pharmacologic or genetic manipulation of this pathway altered Setdb2 in fibroblasts. We also found that fibroblasts treated with pro-inflammatory macrophage supernatants displayed increased Setdb2 and downregulated myofibroblast genes; inhibition of the TNF-α receptor reduced the upregulation of Setdb2. In diabetes, we showed that TNF-α signaling was increased in wound fibroblasts, which functions to increase Setdb2 expression and represses fibroblast to myofibroblast transition. Fibroblast-specific knockdown of SETDB2 and therapeutic inhibition of JAK1,3/STAT3 improved diabetic wound repair, where wound fibroblasts expressed increased myofibroblast genes. This study is the first to our knowledge to identify an epigenetic mechanism for reduced fibroblast to myofibroblast transition in diabetic wounds. Therapeutic targeting of the TNF-α/STAT3/SETDB2 axis in wound fibroblasts may improve diabetic wound healing.
Authors
Tyler M. Bauer, Kevin D. Mangum, Samuel D. Buckley, James Shadiow, Amrita D. Joshi, Christopher O. Audu, Jadie Y. Moon, Lindsey D. Hughes, Rachel Bogel, Lam C. Tsoi, Qinmennge Li, He Zhang, Steven Kunkel, Johann E. Gudjonsson, Frank M. Davis, Katherine A. Gallagher
Citation Information: JCI Insight. 2025. https://doi.org/10.1172/jci.insight.198074.
Abstract
Juvenile idiopathic arthritis (JIA) is the most prevalent chronic inflammatory arthritis of childhood, yet the spatial organization in the synovium remains poorly understood. Here, we perform subcellular-resolution spatial transcriptomic profiling of synovial tissue from patients with active JIA. We identify diverse immune and stromal cell populations and reconstruct spatially defined cellular niches. Applying a newly developed spatial colocalization analysis pipeline, we uncover microanatomical structures, including endothelial–fibroblast interactions mediated by NOTCH signalling, and a CXCL9-CXCR3 signaling axis between inflammatory macrophages and CD8+ T cells, alongside the characterization of other resident macrophage subsets. We also detect and characterize tertiary lymphoid structures marked by CXCL13-CXCR5 and CCL19-mediated signaling from Tph cells and immunoregulatory dendritic cells, analogous to those observed in other autoimmune diseases. Finally, comparative analysis with rheumatoid arthritis reveals JIA-enriched cell states, including NOTCH3+ and CXCL12+ sublining fibroblasts, suggesting potentially differential inflammatory programs in pediatric versus adult arthritis. These findings provide a spatially resolved molecular framework of JIA synovitis and introduce a generalizable computational pipeline for spatial colocalization analysis in tissue inflammation.
Authors
Jun Inamo, Roselyn Fierkens, Michael R. Clay, Anna Helena Jonsson, Clara Lin, Kari Hayes, Nathan D. Rogers, Heather Leach, Kentaro Yomogida
Citation Information: JCI Insight. 2025. https://doi.org/10.1172/jci.insight.194642.
Abstract
Authors
Zhehao Tan, Gio Wu, Daniela Salgado Figueroa, Paramita Dutta, Zachary Jaeger, Marissa Mazurie, David Schairer, Dawn Eichenfield, Wynnis L. Tom, Lauren Galli, Lawrence Eichenfield, Bob Geng, Brian Hinds, Hal M. Hoffman, Lori Broderick, Ben Croker, Ferhat Ay, Reid Oldenburg
Citation Information: JCI Insight. 2025. https://doi.org/10.1172/jci.insight.198590.
Abstract
BACKGROUND. Cannabidiol (CBD) is increasingly used for pain management, including in transplant recipients with limited analgesic options. Its immunomodulatory effects in humans are not well defined at a single cell level at CBD steady state with concomitant tacrolimus treatment. METHODS. In a Phase 1 ex vivo study, peripheral blood mononuclear cells from 23 participants who received oral CBD (Epidiolex®) up to 5 mg/kg twice daily for 11 days were collected before CBD (pre-CBD) and at steady state (post-CBD). Lymphocytes were isolated and stimulated with anti-CD3/CD28 antibodies, with or without tacrolimus (5 ng/mL). Pharmacodynamic responses were assessed using CellTiter-Glo® proliferation, single-cell/nucleus RNA sequencing, cytokine assays, and flow cytometry. Steady-state plasma concentrations of CBD were quantified via tandem mass spectrometry. RESULTS. We identified an increased proportion of T effector memory (TEM) cells post-cannabidiol (22% increase), which correlated with CBD plasma concentrations (R = 0.77, P-value = 0.01). Cannabidiol reduced proliferation of T (37% decrease) and CD70hi B (17% decrease) lymphocytes with additive immunosuppressive effects to tacrolimus. Single-cell RNA sequencing revealed reduced IL2 and TNF signaling and altered receptor–ligand networks in TEM cells. Post-cannabidiol cytokine assays revealed elevated proinflammatory IL-6 protein levels and anti-inflammatory IL-10 levels, with reduced TNF-α, LTA, and IL-2. In flow cytometry, the proportion of TEM and TEMRA increased post-cannabidiol with tacrolimus. CONCLUSION. Cannabidiol exhibits mixed immunomodulatory effects with pro- and anti-inflammatory signals. Understanding the clinical safety of cannabidiol use is important given the paucity of pain control options available for immunocompromised transplant populations.
Authors
Debora L. Gisch, Sachiko Koyama, Jumar Etkins, Gerald C. So, Daniel J. Fehrenbach, Jessica Bo Li Lu, Ying-Hua Cheng, Ricardo Melo Ferreira, Evan Rajadhyaksha, Kelsey McClara, Mahla Asghari, Asif A. Sharfuddin, Pierre C. Dagher, Laura M. Snell, Meena S. Madhur, Rafael B. Polidoro, Zeruesenay Desta, Michael T. Eadon
No posts were found with this tag.