Research ArticleCardiologyCell biologyNeuroscience
Open Access |  10.1172/jci.insight.197721
10.1172/jci.insight.197721
D801N in ATP1A3-encoded Na/K-ATPase alpha 3 causes cardiac arrhythmogenesis through sodium-calcium exchanger–mediated calcium overload
Minu-Tshyeto K. Bidzimou,1 Padmapriya Muralidharan,1 Zhushan Zhang,2 Danyal Raza,1 Daniel Needs,3 Bo Sun,2 Robin M. Perelli,1 Mary E. Moya-Mendez,2 P.K. Rakesh Manivannan,1 Arsen S. Hunanyan,4 Abbigail Helfer,3 Christine Q. Simmons,5 Alfred L. George Jr.,5 Donald M. Bers,6 Nenad Bursac,3 Mohamad A. Mikati,4,7 and Andrew P. Landstrom1,2
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Bidzimou, M. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Muralidharan, P. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Zhang, Z. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Raza, D. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Needs, D. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Sun, B. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Perelli, R. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Moya-Mendez, M. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Manivannan, P. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Hunanyan, A. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Helfer, A. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Simmons, C. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by
George, A.
in:
PubMed
|
Google Scholar
|

1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by
Bers, D.
in:
PubMed
|
Google Scholar
|

1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by
Bursac, N.
in:
PubMed
|
Google Scholar
|

1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Mikati, M. in: PubMed | Google Scholar
1Department of Cell Biology and
2Department of Pediatrics, Division of Cardiology, Duke University School of Medicine, Durham, North Carolina, USA.
3Department of Biomedical Engineering and
4Division of Pediatric Neurology and Developmental Medicine, Department of Pediatrics, Duke University, Durham, North Carolina, USA.
5Department of Pharmacology, Northwestern University Feinberg School of Medicine, Chicago, Illinois, USA.
6Department of Pharmacology, University of California Davis, Davis, California, USA.
7Department of Neurobiology, Duke University, Durham, North Carolina, USA.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
Find articles by Landstrom, A. in: PubMed | Google Scholar
Published April 8, 2026 - More info
JCI Insight. 2026;11(7):e197721. https://doi.org/10.1172/jci.insight.197721.
© 2026 Bidzimou et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: July 22, 2025; Accepted: February 10, 2026
-
Abstract
Short QT syndrome is a heritable arrhythmia disorder linked to sudden cardiac death. We recently identified that individuals with alternating hemiplegia of childhood (AHC), a rare neurodevelopmental disorder, can exhibit shortened corrected QT intervals and elevated risk for ventricular fibrillation. This is especially true for patients with AHC heterozygous for the recurrent ATP1A3-D801N variant, though the underlying cardiac mechanism remains unclear. We hypothesized that the D801N missense impairs Na+/K+-ATPase function, causing Ca2+ overload, shortened action potential duration (APD), and arrhythmias. Using in silico modeling and patient-derived induced pluripotent stem cell cardiomyocytes (iPSC-CMsD801N), we observed shorter APD, elevated intracellular and sarcoplasmic reticulum Ca2+ levels, and delayed afterdepolarizations (DADs) compared with WT. Additionally, increased Ca²+ influx via the Na+/Ca2+ exchanger (NCX1) during depolarization was observed in iPSC-CMsD801N. Simulations and in vitro experiments suggest that reduced ATPase function accelerated inactivation of L-type Ca2+ channels. Pharmacologic inhibition of NCX1 with ORM-10103 normalized APD and reduced DADs. These findings support a Ca2+-mediated mechanism for arrhythmogenesis in ATP1A3-D801N carriers and identify NCX1 as a potential therapeutic target.
-
Introduction
Short QT syndrome (SQTS) is a rare heritable disorder characterized by a shortened QTc interval on a cardiac electrocardiogram (ECG) (1). Individuals with SQTS are at a greater risk of ventricular arrhythmias and sudden cardiac death (SCD) (2, 3). More than 90% of patients with SQTS have an unknown genetic cause and uncertain cellular mechanisms, thus limiting treatment, screening, and prevention (4–6). This lack of mechanistic understanding is reflected in the absence of pharmacologic therapies to treat the disease (7–10).
In light of the unclear molecular underpinnings of SQTS, attention has turned toward fundamental ion transport mechanisms, which are critical for maintaining cardiac excitability and may contribute to arrhythmogenic processes. Na+/K+ ATPases (NKA) belong to a class of proteins that pump intracellular Na+ out of the cell and extracellular K+ into the cell, against their respective electrochemical gradients to maintain the ionic gradients needed for generating action potentials and other physiological phenomena (11). This maintenance of myocyte homeostasis promotes the normal function of other ion-transporting proteins, including the Na+/Ca2+ exchanger (NCX1), a membrane protein that facilitates transport of Ca2+ ions out of the cell with the use of the Na+ gradient (11). By limiting intracellular Na+ accumulation, NKA preserves NCX1 efflux during the action potential plateau and repolarization, thereby preventing excessive Ca2+ loading that would otherwise alter CDI of inward current and the balance of repolarizing currents. In this way, NKA indirectly shapes action potential duration by stabilizing both Ca2+ handling and the timing of the electrogenic NCX1 current (11, 12). The activity of NKA also contributes to an appropriate resting membrane potential (RMP) of cardiomyocytes, which is crucial for action potential dynamics (13). One NKA isoform, ATP1A3-encoded alpha 3 (ATP1A3), has an undefined role in the heart. Classically, variants in ATP1A3 are known to cause a variety of neurologic diseases, with alternating hemiplegia of childhood (AHC) being the most prominent. AHC presents in infancy with episodic hemiplegia, autonomic dysfunction, dystonia, seizures, and oculomotor abnormalities (14). AHC is also associated with increased mortality, with an etiology that has been attributed to sudden unexplained death in epilepsy (SUDEP) (15, 16). Recently, pathogenic variants in ATP1A3 have been found to be associated with short QTc on ECG and predisposition to ventricular arrhythmias (17, 18). Specifically, ATP1A3-D801N is the most recurrent missense accounting for 30%–40% of cases among genotype-positive individuals with AHC, and it is associated with shortened QTc (19). Moreover, patients heterozygous for this variant are at an increased risk of life-threatening ventricular arrhythmias (17, 18). This raises the possibility that abnormal repolarization of the cardiac myocytes due to the ATP1A3-D801N variant may represent a novel mechanism of SQTS and SCD.
In this study, we investigated the expression and function of ATP1A3 in the heart and how the D801N variant may lead to abnormal repolarization and proarrhythmia. We found that diminished Na+/K+-ATPase activity associated with ATP1A3-D801N promotes NCX1-mediated Ca2+ influx at depolarized potentials, which shortens APD, increases intracellular Ca2+ levels and sarcoplasmic reticulum (SR) Ca2+ stores, and consequently predisposes to delayed afterdepolarizations (DADs). We also identified NCX modulation as an approach to rectify APD and decrease DAD prevalence.
-
Results
ATP1A3-D801N has lower NKA function. To begin investigating the role of missense variants in ATP1A3 and short QT development, we first mapped the location of the 3 most common variants in ATP1A3 within the protein structure. We have previously found that the D801N, E815K, and the G947R variants in ATP1A3 are associated with short QT on ECG (18). We mapped these 3 variants to the atomic-level structure of human ATP1A3 and categorized them based on their degree of penetrance, with D801N having high penetrance for short QT. We found that all 3 variants localize to the K+ binding pocket of ATP1A3 (Figure 1A). Since previous evidence has shown that the D801N variant lacks significant ATPase activity (20), we next tested the functional effect of this variant in vitro. HEK293T cells natively express ATP1A3 (21), and overexpression of ATP1A — as well as exposure to ouabain, a selective inhibitor of ATP1A3 — are both toxic to these cells (22). While overexpression of WT ATP1A3 caused cell death, we found that overexpression of ATP1A3-D801N had normal survival. In the presence of ouabain, cell survival was rescued in the setting of overexpression of a ATP1A3 mutant that renders it ouabain resistant; however, ouabain resistant ATP1A3-D801N failed to rescue survival (Supplemental Figure 1, more details in the Supplemental Results; supplemental material available online with this article; https://doi.org/10.1172/jci.insight.197721DS1). These findings suggest that D801N strongly impairs ATP1A3 function.
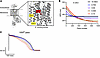 Figure 1
Figure 1ATP1A3-D801N decreases NKA function and is predicted to decrease cardiomyocyte depolarization time. (A) Variant mapping of the 3 most QTc penetrant missenses among patients with AHC. Crystal structure of the human ATP1A3 is shown with the D801, E815, and G947 residues highlighted. Residues in yellow have 10%–20% penetrance of SQTc while residues in red have a penetrance higher than 50% (D801N, 68.57%; E815K, 19.05%; G947R, 12.50%). (B and C) In silico ToR-ORd model of human cardiomyocyte predicting APD as a function of pace number under different NKA functional states.
Using established transcriptional data, we found that ATP1A3 comprises 26.3% of the total ATP1A transcript levels in healthy human myocardium (Supplemental Figures 2 and 3). Conversely, RNA-seq conducted in 3-day-old (P3) and 3-month-old mice found that Atp1a3 transcripts contribute less than 1% of Atp1a in WT murine hearts (Supplemental Figure 4). We found a similar low relative abundance of Atp1a3 among other in vivo experimental models. Atp1a3 transcripts contribute 2% of total Atp1a in rat and sheep hearts (Supplemental Figures 5 and 6), atp1a3 is not expressed in zebrafish heart, and there are no orthologs of ATP1A3 in either Caenorhabditis elegans or drosophila melanogaster (Supplemental Results). These findings suggest that human myocardium, unlike commonly used experimental animal models, expresses substantial ATP1A3 and may be uniquely susceptible to the functional effect of the D801N variant. We next employed an in silico model of human cardiomyocyte electrophysiology, the Tomek-Rodriguez (ToR-ORd) model, to evaluate the effect of decreased NKA function (23). In line with our previous findings suggesting that the D801N variant shortens the QT interval and APD (18), we found that reducing NKA function (by 25%) was associated with shorter APD (Figure 1, B and C). This model also predicted more depolarized RMP as expected for reduced NKA function (Supplemental Figure 7). Overall, these results suggest that loss of proper ATP1A3 function in human myocardium is associated with APD shortening and RMP depolarization.
ATP1A3-D801N is associated with faster cardiac myocyte repolarization and DADs. To determine the mechanism of ATP1A3-D801N–mediated alterations in cardiac repolarization and arrhythmogenesis, we leveraged 2 iPSC lines from D801N+ AHC probands with short QTc (male iPSCD801N and female iPSCD801N-2) (18). Each patient-derived line was compared with a corresponding control: an isogenic WT was used as a control for the male iPSCD801N line and an unrelated healthy male WT line for the female iPSCD801N-2 (iPSCcWT and iPSCWT-2, respectively) (18). Using immunofluorescence, we found that expression of ATP1A3 colocalized with the NCX1 in both iPSC-CMsWT and in iPSC-CMsD801N (Figure 2A and Supplemental Figure 8). Expression of ATP1A1, ATP1A2, ATP1A3, and ATP1B1 in iPSC-CMs were confirmed by Western blot in iPSC-CMsWT and iPSC-CMsD801N (Figure 2B). We recently demonstrated that iPSC-CMsD801N exhibit a shortened APD, depolarized RMP, and DADs by single-cell patch clamp recording (18). We employed a genetically encoded fluorescence voltage sensor (Arclight) to evaluate cellular membrane depolarizations (Figure 2C). Live cell imaging of Arclight transduced iPSC-CMs revealed significantly shorter repolarization time and the presence of DADs in CMs differentiated from both D801N lines compared with respective controls (Figure 2, D–H, and Supplemental Figure 9). Application of ouabain to iPSC-CMWT-2 was associated with shortening of APD50, APD90, and more depolarized maximum diastolic potential (MDP) compared with vehicle treatment, all of which were similar to action potential kinetics of iPSC-CMsD801N (Supplemental Figure 10) (18). Together, these findings demonstrate that D801N does not alter expression or trafficking of ATP1A3 to the membrane, yet it is associated with shortened APD and DAD generation.
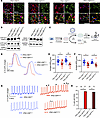 Figure 2
Figure 2iPSC-CMD801N have shortened APD and delayed afterdepolarizations. (A) Representative images of iPSC-CMs immunofluorescence: ATP1A3 (green), NCX1 (red), and DAPI (blue). (B) Western blots of ATP1A1-3, ATP1B1, and GAPDH expression in iPSC-CMsWT and iPSC-CMsD801N. (C) Schematic of experimental approach for iPSC-CMs using whole cell patch clamp or a genetically encoded fluorescent voltage sensor (Arclight), packaged in lentivirus (LV). (D) Representative fluorescence tracing of Arclight. (E and F) Fluorescence change duration at 50% and 90% from baseline (representing APD50 and APD90, respectively). For iPSC-CM^cWT, iPSC-CM^D801N, iPSC-CM^WT-2, iPSC-CM^D801N-2, n = 19, 21, 22, and 13, respectively. (G) Representative tracings of iPSC-CMs spontaneous oscillations in Arclight transduced iPSC-CMs. In spontaneously fluorescing cells, arrows identify irregularly triggered fluorescence indicative of delayed afterdepolarizations (DADs). (H) Bar graph quantifying percent cells with DADs. Live cell experiments were statistically analyzed with a hierarchical approach. Fischer’s exact test was conducted on H. E and F were statistically tested with a hierarchical approach.*P < 0.05, **P < 0.01.
iPSC-CMD801N have Ca2+ miniwaves, greater SR Ca2+ store levels, and higher cytosolic Ca2+ levels. Ion transport coupling between outward NKA and inward NCX1 currents allow the cell to repolarize after each depolarization, and changes in NKA function and [Na+]i can affect NCX1-mediated Ca2+ flux (13). To explore the influence of NKA on Ca2+ dynamics in our cellular model, we conducted live cell imaging of Cal-520–loaded iPSC-CMs. We observed that iPSC-CMsD801N have similar Ca2+ transient amplitudes and similar transient resolution time (tau) when compared with iPSC-CMsWT (Figure 3, A–C, and Supplemental Figure 11). We also found that iPSC-CMsD801N are predisposed to the generation of miniwaves, which manifest as small, truncated Ca2+ transients, and are a known cellular mechanism of triggered arrhythmias (24). Conversely, iPSC-CMsWT demonstrated no miniwaves (Figure 3C and Supplemental Figure 12). We next measured SR Ca2+ store levels with the application of 10 mM caffeine and found higher SR Ca2+ store levels in iPSC-CMsD801N compared with iPSC-CMsWT (Figure 3, D and E). Using Fura-2 with prepacing of cardiomyocytes at 0.5 Hz, we found higher basal levels of [Ca2+]i in iPSC-CMsD801N compared with controls (Figure 3F), which was in close agreement with our in silico model showing that lower NKA function were associated with higher [Ca2+]i and Ca2+ stores (Supplemental Figure 13).
![iPSC-CMD801N have miniwaves, higher SR [Ca2+] store content, and higher [Ca2+]i. iPSC-CMD801N have miniwaves, higher SR [Ca2+] store content, and higher [Ca](//df6sxcketz7bb.cloudfront.net/manuscripts/197000/197721/small/jci.insight.197721.f3.gif) Figure 3
Figure 3iPSC-CMD801N have miniwaves, higher SR [Ca2+] store content, and higher [Ca2+]i. (A) Representative Ca2+ transients of Cal-520 loaded iPSC-CMsWT and iPSC-CMsD801N paced at 0.5 Hz. Black arrows indicate miniwaves. (B) Quantification of transient fluorescence amplitude. (C) Bar graph demonstrating percent cells with miniwaves. Fisher’s exact test. (D) Representative trace of Cal-520 fluorescence upon application of 10 mM caffeine as a measure of sarcoplasmic reticulum (SR) Ca2+ load. (E) SR store fluorescence amplitude. For iPSC-CM^cWT, iPSC-CM^D801N, iPSC-CM^WT-2, iPSC-CM^D801N-2, n = 43, 27, 31, and 29, respectively. (F) Diastolic fura-2 ratio after 0.5Hz pacing. For iPSC-CM^cWT, iPSC-CM^D801N, iPSC-CM^WT-2, iPSC-CM^D801N-2, n = 133, 156, 87, and 53, respectively. Black dots represent experimental means. Live cell experiments were statistically analyzed with a nested approach. Fischer’s exact test was conducted on C. *P < 0.05, **P < 0.01.
To determine whether elevated SR Ca2+ content was associated with greater Ca2+ leak from the SR, we explored RyR2-mediated leak by measuring Ca2+ sparks with confocal live cell imaging. We found that iPSC-CMsD801N had comparable spark frequency compared with controls (Supplemental Figure 14, A and B). Furthermore, while sparks from iPSC-CMsD801N cells had higher amplitude, all other Ca2+ spark parameters, such as spark width and duration, were similar to iPSC-CMsWT (Supplemental Figure 14, C–E). The higher Ca2+ sparks amplitude would tend to promote propagating Ca2+ waves and limit the rise in SR Ca2+ content. When normalized to SR Ca2+ store levels, which is positively associated with SR Ca2+ leak levels, we found that iPSC-CMsD801N had similar normalized Ca2+ leak compared with control cells (Supplemental Figure 14, F and G). This suggests that the prevalence of miniwaves in D801N myocytes is due to increased SR Ca2+ content rather than a fundamental alteration in RyR2 gating. Overall, we found that iPSC-CMsD801N have higher levels of Ca2+ within the cytosol, as well as the SR, and are predisposed to Ca2+ miniwaves when compared with iPSC-CMsWT.
iPSC-CMD801N exhibit greater NCX1 Ca2+ influx at positive membrane potentials. We further investigated the link between ATP1A3-D801N and Ca2+ overload. Given our findings of increased [Ca2+]i (likely driven by a rise in [Na+]i), we hypothesized that NCX1 function is altered, thus loading iPSC-CMsD801N with intracellular Ca2+ in the setting of a reduced [Na+] gradient. In silico simulations show how lower NKA function increases [Na+]i (Figure 4A) and Ca2+ influx (Supplemental Figure 15). We calculated NCX1 Ca2+ efflux using the tau decay of Ca2+ extrusion after maximal SR store Ca2+ release and found that iPSC-CMD801N had slowed Ca2+ efflux than iPSC-CMcWT (Supplemental Figure 16 and 17). That is consistent with the elevated [Na+]i despite the elevated [Ca2+]i release (Figure 3, D and E). SERCA activity during [Ca2+]i decline was higher in iPSC-CMD801N cells compared with control (Supplemental Figure 18), which reflects the higher intracellular and SR Ca2+ levels and an increase in SERCA versus NCX dominance in cytosolic Ca2+ removal SERCA. The iPSC-CMsD801N had comparable NCX1 expression with iPSC-CMsWT by Western blot (Supplemental Figure 19), suggesting that the altered NCX function is due to the altered conditions, rather than NCX1 protein expression level. To further test this finding in vitro, we measured NCX1 current as a function of membrane potential under more controlled [Ca2+]i and [Na+]i conditions. In iPSC-CMD801N-2, inward INCX (Ca2+ efflux) was similar to iPSC-CMsWT when cells were held at negative membrane potentials (18). Conversely, outward INCX at positive membrane potentials was higher in iPSC-CMD801N-2 compared with iPSC-CMWT-2, reflecting greater Ca2+ influx during depolarized membrane potentials (Figure 4, B–D). Although the Ca2+ flux toward new steady state can be contributed by other compensatory mechanisms, we wanted to evaluate the NCX1 specific contribution. To accomplish this, we calculated the driving force of NCX1 flux in D801N and WT cardiomyocytes (25, 26) and found higher Ca influx in D801N (Figure 4, E and F).
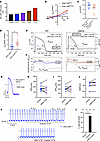 Figure 4
Figure 4iPSC-CMD801N have greater NCX1 Ca2+ influx at positive membrane potential. (A) In silico ToR-ORd human cardiomyocyte model; graph illustrates [Na+]i as a function of NKA functional states after 1,000 paces. (B) IV curve of INCX in iPSC-CMWT-2 and iPSC-CMD801N-2. (C) INCX current of iPSC-CMWT-2 and iPSC-CMD801N-2 at their respective maximum diastolic potential. (D) INCX current of iPSC-CMWT-2 and iPSC-CMD801N-2 at +80mV. (E and F) Graphs illustrating changes in ENa/Ca during the action potential of WT and D801N cardiomyocytes. NCX mediated Ca2+ influx is thermodynamically favored when Em > ENa/Ca and NCX-mediated Ca2+ efflux is favored when Em < ENa/Ca. (G) AP trace of iPSC-CMWT-2 treated with vehicle and with 1 × 10–5 M ORM10103 + 1 × 10–7 M ouabain. (H–J) Action potential duration at 50% repolarization (APD50), 90% repolarization (APD90), and maximum diastolic potential (MDP) of vehicle treated and ouabain + ORM10103–treated iPSC-CMWT-2. (K and L) Ouabain-mediated DAD rescue by cotreatment with ORM-10103. Arrows indicate delayed afterdepolarizations (DAD) events). Results were statistically analyzed with a Mann-Whitney U test or Wilcoxon matched-pairs test. Fischer’s exact test was conducted on K. *P < 0.05, **P < 0.01.
To test whether Ca2+ influx can be modulated to be a net efflux and consequently alter APD, we used ORM-10103, a small molecule that preferentially inhibits NCX1 Ca2+ influx (27). After pretreatment with ouabain to partially inhibit NKA (and raise [Na+]i), treatment of iPSC-CMWT-2 with ORM-10103 normalized the APD and MDP comparable with vehicle-treated iPSC-CMWT-2 parameters (Figure 4, G–J). We also observed that ORM-10103 prevented ouabain-induced DADs in iPSC-CMWT-2 (Figure 4, K and L). Taken together, we conclude that ATP1A3-D801N promotes NCX1-mediated Ca2+ influx at positive membrane potentials, limiting net NCX Ca2+ efflux, and that inhibition of NCX1-mediated Ca2+ influx normalizes APD shortening associated with ATP1A3 inhibition and prevents DAD generation.
Lower NKA function promotes faster L-type Ca2+ channel inactivation. To determine the biophysical mechanism of shortened APD, we hypothesized that phase 2 of the action potential is shorter in iPSC-CMsD801N because of Ca2+-dependent inactivation (CDI) of the L-type Ca2+ channel (LTCC). In silico simulations suggest that lower NKA function was associated with a very slight increase in peak ICa,L, accelerated inactivation and a negative shift in steady state LTCC availability (Figure 5, A and B). Additionally, we found that lower NKA activity was associated with faster IKr inactivation but with lower peak current, suggesting a compensatory response to shortened APD rather than the driving effect (Supplemental Figure 20). In vitro voltage-clamp of iPSC-CMD801N-2 had a trend toward faster inactivation compared with control, with similar peak current amplitudes (Figure 5, C–E). However, this interpretation is complicated by the residual outward currents present in our relatively physiological solutions. Thus, our ATP1A3-D801N cardiomyocytes may have faster ICa,L inactivation.
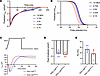 Figure 5
Figure 5Lower NKA functional states lead to faster L-type Ca2+ channel inactivation. (A and B) In silico TOr-ORd human cardiomyocyte model using decreasing NKA functional states after 1,000 paces. Peak LTCC current (A) and calculated LTCC steady state inactivation (B). (C–E) Patch clamp recordings. (C) Current density raw trace of iPSC-CMWT-2 and iPSC-CMD801N-2 using a 1-step activation protocol. (D) Bar graph illustrating current density per cell. (E) Bar graph illustrating tau of decay per cell. A Mann-Whitney U test was conducted, and no statistical difference was found.
Inhibiting NCX1 Ca2+ influx with ORM-10103 rescues APD and prevents DADs in iPSC-CMD801N. We tested the feasibility of rescuing the shortened APD and DAD generation in iPSC-CMsD801N using outward-biased INCX inhibition. Using Arclight-transduced cardiac myocytes, we measured APD50 and APD90 in vehicle-treated and ORM-10103–treated cardiomyocytes. We found that ORM-10103 increased APD50 and APD90 in iPSC-CMsD801N but had no effect on APD in iPSC-CMsWT (Figure 6, A–C). This is consistent with the idea that there is relatively little outward INCX at normal [Na+]i levels (13). We also observed that ORM-10103 suppressed DADs in iPSC-CMsD801N (Figure 6D). In summary, we found that inhibition of NCX1-mediated Ca2+ influx with ORM-10103 rescued APD and prevented DADs in the D801N iPSC-CM models, which suggests a potentially new therapeutic approach to shortened QTc in ATP1A3-D801N (Figure 7).
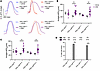 Figure 6
Figure 6NCX1 Ca2+ influx inhibition with ORM-10103 rescues APD and prevents DADs in iPSC-CMsD801N. Rescue of APD and DADs with ORM-10103 on Arclight transduced cardiomyocytes. (A) Representative fluorescence tracing of vehicle and ORM-10103–treated cells. (B and C) Fluorescence change duration at 50% and 90% from baseline representing APD50 and APD90, respectively. (D) Percent cells with DADs when treated with vehicle or 1 × 10–5 M ORM-10103. Black dots represent experimental means. Results were statistically analyzed with a hierarchical test. Fischer’s exact test was conducted on D. Panels B and C were statistically analyzed with a hierarchical test. *P < 0.05, **P < 0.01.
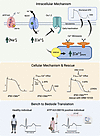 Figure 7
Figure 7Cellular mechanism of ATP1A3-D801N. Expression of ATP1A3-D801N leads to a decrease in NCX1 mediated Ca2+ efflux. This causes a buildup of intracellular [Ca2+]i and SR store, favoring miniwaves and delayed afterdepolarizations (DADs). In the setting of Ca2+ overload, Cav1.2 rapidly inactivates which shortens phase 2 and leads to a shorter action potential (APD). Inhibition of NCX1-mediated Ca2+ influx with ORM-10103 rescues both APD and DADs in iPSC-CMsD801N, suggesting that NCX1 can be modulated in patients with ATP1A3-D801N to increase ventricular repolarization time and decrease arrhythmic risk.
-
Discussion
Our findings presented herein demonstrate that individuals with ATP1A3-D801N have a myocardium that is susceptible to arrhythmia. In our experiments using AHC patient–derived iPSC-CMsD801N and the ToR-ORd model, we found that ATP1A3-D801N leads to elevation of [Na+]i, which limits NCX1-mediated Ca2+ efflux and also increases Ca2+ influx at depolarized potentials. Over time, this leads to Ca2+ overload, both in the cytoplasm and SR. These Ca2+ overloaded cells contribute to the APD and DAD phenotype via outward INCX and miniwaves, respectively.
The modulators of cardiac myocyte repolarization are still emerging. More than 90% of patients with SQTS have an unknown genetic cause and unknown cellular mechanism, thus limiting treatment, screening, and prevention of life-threatening ventricular arrhythmias (5, 6). Challenges in identifying mechanisms of SQTS have slowed the development of models that can recapitulate disease and test potential treatments. Most genotype-positive individuals with heritable SQTS have gain-of-function variants in K+-channel–encoding genes such as KCNH2 and KCNQ1 (28). These variants lead to increased outward current, which shorten phase 3 of the ventricular action potential and consequently shorten its APD and shortened QT intervals on ECG. Conversely, loss-of function variants in CACNA1C encoded LTCC have been associated with SQTS (29). It has been shown in vitro that SQTS-associated CACNA1C variants exhibit smaller inward current that contributes to shorter phase 2 of the action potential (29). To date, there is no mechanism to our knowledge to explain how variants in NKA genes cause faster cardiac repolarization or arrhythmias, although reduced NKA function is expected to reduce the outward Na-pump current and shift INCX in the outward direction, thus shortening APD. Multiple clinical studies have been reported in the treatment of SQTS, and hydroquinidine has been shown to be the most successful in increasing APD and reducing arrhythmic burden (7). However, those studies overly represented K+ channel variants. While targeting NCX1 activity may represent a rational strategy to mitigate Ca2+ overload secondary to ATP1A3-D801N–mediated NKA dysfunction, such an approach is likely context dependent. In SQTS unrelated to NKA variants, or under conditions of limited Ca2+ loading, NCX1 inhibition could impair contractility and alter electrophysiological stability, underscoring the need for careful therapeutic evaluation.
NKA function has historically been a target for pharmacologic intervention of heart failure with cardiac glycosides. Partial inhibition of NKA increases [Na+]i and promotes NCX1-dependent Ca2+ loading of the cardiac myocytes and improved contractility (30). Of note, significant inhibition of ATP1A3 with the cardiac glycoside ouabain has been shown to decrease NKA function and lead to pathologic increases in [Ca2+]i and DAD-triggered arrhythmias (31). We find evidence of both elevated [Ca2+]i and DADs in the setting of the D801N missense variant, as well as higher [Na+]i with lower NKA function in the ToR-ORd model. While we are the first to show this in the cardiac myocyte, previous reports have found that D801N affects K+ binding to NKA, which impairs the pump cycling from the high energy state phosphorylated conformation (E2P) to the unphosphorylated conformation (E2) and decreases NKA pump function. (20). We find that ouabain causes shortened APD, and we observe both shortened APD and Ca2+ overload phenomenon in iPSC-CMsD801N, suggesting that lower ATP1A3 function leads to increased [Na+]i and [Ca2+]i, which promote generation of Ca2+ miniwaves and DADs in the myocardium.
Excess intracellular Ca2+ shortens APD by altering the balance of inward and outward ionic currents through both direct Ca2+-dependent modulation and secondary changes in electrogenic Ca2+ handling. Elevated cytosolic Ca2+ accelerates CDI of LTCCs, reducing inward ICa,L during the plateau phase and thereby favoring earlier repolarization (32). In parallel, Ca2+ overload suppresses IK1 and can enhance repolarizing K+ currents such as IKr, shifting the net current balance toward APD shortening and increased susceptibility to arrhythmogenesis (33). Experimental and computational studies further demonstrate that Na+ overload–driven Ca2+ accumulation via NCX influx augments repolarizing INaK and INaCa, contributing to APD shortening despite increased Ca2+ influx and Ca2+ transient amplitude (34).
Ca2+-activated K+ currents also participate in Ca2+-mediated APD modulation. Small-conductance Ca2+-activated K+ (SK) channels (ISK) can enhance repolarization and shorten APD when activated by low-to-moderate increases in intracellular Ca2+ (35). However, ISK exhibits a biphasic dependence on [Ca2+]i, such that more severe Ca2+ overload induces Ca2+-dependent block, limiting its contribution during the plateau phase (36). Accordingly, in pathological settings characterized by augmented Ca2+ influx such as in ATP1A3-D801N–expressing ventricular myocytes, the contribution of ISK to phase-2 APD shortening is anticipated to be reduced, with APD shortening dominated instead by enhanced CDI of ICa,L and Ca2+-driven changes in other repolarizing and electrogenic transport currents.
Consistent with this integrated framework, intact-heart experiments and human ventricular modeling demonstrate that elevations in cytosolic and SR Ca2+, particularly at higher pacing rates, enhance CDI of ICa,L, modify RyR2 gating, and act synergistically with rate-dependent increases in repolarizing K+ currents (Ito, IKr, IKs) to shorten APD (37). Clinically oriented experimental-computational studies and population-based modeling further support an inverse relationship between Ca2+ loading and APD across mammalian preparations, largely mediated by Ca2+-dependent regulation of ICa,L, and secondary alterations in electrogenic Ca2+ transport, including Na+/Ca2+ exchange and Ca2+-ATPase activity, linking Ca2+ overload to both earlier repolarization and increased risk of Ca2+ mediated triggered activity (38–40).
Multiple ATP1A3 pathogenic variants have been associated with human disease, including AHC (41). AHC presents in infancy with epilepsy, episodic hemiplegia, autonomic dysfunction, dystonia, nystagmus, and sudden unexpected death. Variants in ATP1A3 account for 70% of reports of AHC and the D801N missense-causing variant is the most recurrent, associated with 30%–40% of instances of AHC (19). While this genetic association is known, the cause of sudden death in these patients remains unclear. Individuals with AHC have been viewed as at-risk for SUDEP, which is postulated to have multifactorial etiologies: neurologic, cardiovascular, and respiratory as well as iatrogenic (42). While the contribution of each factor to instances of SUDEP is not well understood, epilepsy can coexist with heritable arrhythmic syndromes (43). In a study of patients with SUDEP, 11% had a postmortem genetic finding of pathogenic/likely pathogenic variants in Na+ or K+ channel encoding genes, and many patients were found to have variants of uncertain significance in cardiac channelopathy-associated genes (42). Thus, genetic variants in genes that are responsible for the depolarization and repolarization of the cardiac myocyte may be common causes of SUDEP. Although there are no FDA-approved NCX1 modulating therapies, multiple ORM1-10103 analogs have been investigated in cardiac arrhythmia models (44), suggesting an important target of cardiac pathophysiology.
There is a shared substrate of risk in patients with ATP1A3-D801N, reflected in both a neurologic and cardiac risk of sudden death (45–47). For example, QTc variability has also been shown to transiently occur in patients with epilepsy during the ictal and post-ictal states (48). Given that ATP1A3-D801N causes seizures as well as dysautonomia in AHC, it raises the possibility of a central exacerbation of underlying myocardial arrhythmia susceptibility. A recent study in Atp1a3D801N mice, which has exceedingly low expression of Atp1a3 in the heart, demonstrated a prodromal arrhythmic period that occurred before the pre-ictal window of terminal seizures, suggestive of an additional susceptibility to arrhythmia from dysautonomia alone (49).
Further studies should explore the autonomic regulation of cardiac arrhythmias in ATP1A3-D801N. In a recent preprint, we have shown that patients with ATP1A3-D801N exhibit paradoxical QT and QTc shortening at lower heart rates (50). We have also demonstrated that Atp1a3D801N mice have increased sinus arrhythmia and sinus pauses in the early pre-ictal states (49). Consistent with these findings, other investigators have also shown that gain-of-function variants in other SQTS genes, such as the anion exchanger SLC4A3, also lead to paradoxical QTc shortening (51) and that augmented vagal effects may trigger ventricular arrhythmias in SQTS (52). We hypothesize that sinus pauses and bradycardia may precipitate ectopic activity and increase the arrhythmic vulnerability of a myocardium expressing ATP1A3-D801N. If validated, this mechanism may be relevant to other forms of SQTS.
Limitations. Our current study is limited by the immaturity of iPSC-CMs, which lack well-developed T-tubules. Subcellular localization, particularly within the T-tubule system, may be key to understanding ATP1A3 function. In murine hearts, Atp1a2 is known to preferentially express at T-tubules, while Atp1a1 shows no specific localization within the sarcolemma (53). To address this, primary adult cardiomyocytes isolated from human tissue, as well as engineered heart tissues and organoids, may be better suited for investigating the subcellular localization of ATP1A3 relation to ATP1A1-2 and NCX1.
ATP1A3 constitutes roughly 26% of total NKA alpha subunit transcripts in cardiomyocytes, but a larger apparent loss of NKA function than observed here in the ATP1A3-D801N cells (e.g., on APD), according to the ToR-ORd simulations, suggests that the D801N missense variant may also impair the activity of the WT allele. In our previous international, multicenter case-control study, we observed that patients with heterozygous loss-of-function variants such as splice variants or ATP1A3 deletions as well as other missense variants that are not D801N did not display QTc shortening (18), supporting the hypothesis that D801N may act through a different mechanism, possibly a dominant-negative mechanism. This interpretation is consistent with experimental evidence from Xenopus laevis oocytes expressing human ATP1A3 transcripts (54). Although our mechanistic work did not directly assess the effect of the missense allele on the function of WT subunits, future studies using antisense oligonucleotides as a potential rescue approach could help address this important gap.
The autonomic nervous system findings in both patients with AHC and Atp1a3D801N mice underscore a critical gap in our understanding of neuro-cardiac interactions. The potential role of autonomic abnormalities in patients with AHC leading to increased risk for arrhythmias in the setting of short QT remains unknown. We hypothesize that impaired autonomic regulation, either peri-ictal or in the setting of AHC-mediated bradycardia, may provide a “second hit,” which exacerbates the arrhythmic predisposition imparted by a shortened QT interval. Further studies are needed to explore this hypothesis. Importantly, animal models such as mouse, rat, sheep, C. elegans, Drosophila, and zebrafish, either express very low levels of Atp1a3 in the heart or lack a cardiac ortholog entirely. This lack of ATP1A3 ortholog in commonly used animal models has limited our ability to address the translations of our findings in an animal model because the animals do not adequately recapitulate the cardiac pathology that manifests in humans harboring ATP1A3-D801N.
Additionally, systemic inhibition of NCX1 may affect exchanger function in noncardiac tissues, making it important to understand potential off-target effects of this therapeutic strategy. Because we lacked a mouse model with endogenous cardiac Atp1a3 expression, we were unable to evaluate the effect of ORM-10103 on other organ systems. Nonetheless, our findings provide proof of concept. Further studies should focus on models that more faithfully recapitulate human ATP1A3 expression in the myocardium, enabling assessment of NCX inhibition beyond the heart. These may include coculture systems using patient-derived iPSC cardiomyocytes and neurons to address autonomic abnormalities found in this disease, as well as transgenic mice engineered to express cardiac Atp1a3 at human-comparable levels.
In conclusion, our findings contribute to the growing evidence linking ATP1A3 to channelopathies, the phenotypic expansion of AHC, and the possible mechanistic link between SQTS with SUDEP.
-
Methods
Sex as a biological variable. This study uses cell lines from 1 female AHC patient and 1 male AHC patient, and similar findings were observed in these lines.
Statistics. For all experiments, a minimum of 3 passages were used for each line per experiment. To determine statistical significance among groups, a paired 2-tailed Student’s t test was performed for data with a normal distribution and 2 groups, and a 1-way ANOVA with multiple comparisons was used for 3 groups. For statistical tests assuming a normal distribution, the Shapiro-Wilk test for normality was used. For nonparametric data, a Mann-Whitney U test was used when comparing 2 groups. Fisher’s exact tests were used for categorical data. Mann-Whitney U tests or Wilcoxon matched-pairs tests were conducted for nonparametric statistical analysis for whole cell patch clamp experiments. Live cell imaging experiments were analyzed using a hierarchical statistical approach, as previously described (55). With nonparametric data, the data were first log10-transformed before applying the R code. Data are presented as the mean ± SEM. P < 0.05 were considered statistically significant.
Study approval. This study was approved by Duke University IRB (Pro00056651 and Pro00094341). Informed consent was obtained from each participant or participant’s parent/legal guardian in accordance with the Declaration of Helsinki. Methods are further detailed in the Supplemental Methods.
Data availability. A detailed description of all experimental methods can be found in the Supplemental Methods. This study reanalyzes previously published RNA-seq data as detailed in the Supplemental Methods. Supporting data values of relevant panels are provided in the Supporting Data Values file. Raw data are available upon reasonable request to the corresponding author.
-
Author contributions
Design and interpretation of results were contributed by MTKB, PM, ZZ, DR, DN, BS, RMP, MEMM, PKRM, ASH, AH, CQS, ALG, DMB, NB, MAM, and APL. Supervisory roles were contributed by ALG, DMB, NB, MAM, and APL. Data acquisition was contributed by MTKB, PM ZZ, DR, DN, BS, MEMM, and PKRM. Writing of manuscript was contributed by MTKB, PM, RMP, ALG, DMB, and APL.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- The Paul Gillette PACES Research Grant (MTKB.)
- National Institute of Health (NIH) T32 GM007171 (MTKB).
- NIH (R01-HL160654, R01-HL166217) (APL).
- Doris Duke Charitable Foundation (CSDA-2020098) (APL).
- John Taylor Babbitt Foundation (APL).
- The Hartwell Foundation (APL).
- Additional Ventures (APL).
- Y.T. and Alice Chen Pediatric Genetics and Genomics Research Center (APL).
- NIH (NS125785) (ALG).
- Alternating Hemiplegia of Childhood Foundation (ALG).
-
Acknowledgments
We acknowledge the use of Biorender.com with assistance in figure design.
Address correspondence to: Andrew P. Landstrom, Duke University Medical Center, Box 2652, Durham, North Carolina, 27710, USA. Phone: 919.684.3028; Email: andrew.landstrom@duke.edu.
-
Footnotes
Conflict of interest: ASH and MAM have intellectual property interest in US Patent No. 11,717,560 for gene therapy for ATP1A3-related diseases.
Copyright: © 2026, Bidzimou et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(7):e197721.https://doi.org/10.1172/jci.insight.197721.
-
References
- Bidzimou MK, Landstrom AP. From diagnostic testing to precision medicine: the evolving role of genomics in cardiac channelopathies and cardiomyopathies in children. Curr Opin Genet Dev. 2022;76:101978.
- Pereira R, et al. Short QT syndrome in pediatrics. Clin Res Cardiol. 2017;106(6):393–400.
- Campuzano O, et al. Recent advances in short QT syndrome. Front Cardiovasc Med. 2018;5:149.
- Chugh SS, et al. Epidemiology of sudden cardiac death: clinical and research implications. Prog Cardiovasc Dis. 2008;51(3):213–228.
- Ackerman MJ, et al. HRS/EHRA expert consensus statement on the state of genetic testing for the channelopathies and cardiomyopathies this document was developed as a partnership between the Heart Rhythm Society (HRS) and the European Heart Rhythm Association (EHRA). Heart Rhythm. 2011;8(8):1308–1339.
- Landstrom AP, et al. Genetic testing for heritable cardiovascular diseases in pediatric patients: a scientific statement from the American Heart Association. Circ Genom Precis Med. 2021;14(5):e000086.
- Mazzanti A, et al. Hydroquinidine prevents life-threatening arrhythmic events in patients with short QT syndrome. J Am Coll Cardiol. 2017;70(24):3010–3015.
- Mizobuchi M, et al. Nifekalant and disopyramide in a patient with short QT syndrome: evaluation of pharmacological effects and electrophysiological properties. Pacing Clin Electrophysiol. 2008;31(9):1229–1232.
- Giustetto C, et al. Long-term follow-up of patients with short QT syndrome. J Am Coll Cardiol. 2011;58(6):587–595.
- Al-Khatib SM, et al. 2017 AHA/ACC/HRS guideline for management of patients with ventricular arrhythmias and the prevention of sudden cardiac death: executive summary: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines and the Heart Rhythm Society. Heart Rhythm. 2018;15(10):e190–e252.
- Shattock MJ, et al. Na+/Ca2+ exchange and Na+/K+-ATPase in the heart. J Physiol. 2015;593(6):1361–1382.
- Contreras RG, et al. Na+/K+-ATPase: more than an electrogenic pump. Int J Mol Sci. 2024;25(11):6122.
- Bers DM. Cardiac excitation-contraction coupling. Nature. 2002;415(6868):198–205.
- Mikati MA, et al. Revision of the diagnostic criteria of alternating hemiplegia of childhood. Eur J Paediatr Neurol. 2021;32:A4–A5.
- Panagiotakaki E, et al. Clinical profile of patients with ATP1A3 mutations in alternating hemiplegia of childhood-a study of 155 patients. Orphanet J Rare Dis. 2015;10:123.
- Uchitel J, et al. Alternating hemiplegia of childhood: evolution over time and mouse model corroboration. Brain Commun. 2021;3(3):fcab128.
- Moya-Mendez ME, et al. ATP1A3-encoded sodium-potassium ATPase subunit alpha 3 D801N variant is associated with shortened QT interval and predisposition to ventricular fibrillation preceded by bradycardia. J Am Heart Assoc. 2021;10(17):e019887.
- Moya-Mendez ME, et al. ATP1A3 variants, variably penetrant short QT intervals, and lethal ventricular arrhythmias. JAMA Pediatr. 2025;179(5):529–539.
- Capuano A, et al. Alternating hemiplegia of childhood: understanding the genotype-phenotype relationship of ATP1A3 variations. Appl Clin Genet. 2020;13:71–81.
- Weigand KM, et al. Alternating hemiplegia of childhood mutations have a differential effect on Na(+),K(+)-ATPase activity and ouabain binding. Biochim Biophys Acta. 2014;1842(7):1010–1016.
- Kockskamper J, et al. Comparison of ouabain-sensitive and -insensitive Na/K pumps in HEK293 cells. Biochim Biophys Acta. 1997;1325(2):197–208.
- Cherniavsky Lev M, et al. Cardiac glycosides induced toxicity in human cells expressing α1-, α2-, or α3-isoforms of Na-K-ATPase. Am J Physiol Cell Physiol. 2015;309(2):C126–C135.
- Tomek J, et al. Development, calibration, and validation of a novel human ventricular myocyte model in health, disease, and drug block. Elife. 2019;8:e48890.
- Tomek J, et al. SparkMaster 2: a new software for automatic analysis of calcium spark data. Circ Res. 2023;133(6):450–462.
- Piacentino V, et al. Cellular basis of abnormal calcium transients of failing human ventricular myocytes. Circ Res. 2003;92(6):651–658.
- Weber CR, et al. Dynamic regulation of sodium/calcium exchange function in human heart failure. Circulation. 2003;108(18):2224–2229.
- Jost N, et al. ORM-10103, a novel specific inhibitor of the Na+/Ca2+ exchanger, decreases early and delayed afterdepolarizations in the canine heart. Br J Pharmacol. 2013;170(4):768–778.
- Patel U, Pavri BB. Short QT syndrome: a review. Cardiol Rev. 2009;17(6):300–303.
- Antzelevitch C, et al. Loss-of-function mutations in the cardiac calcium channel underlie a new clinical entity characterized by ST-segment elevation, short QT intervals, and sudden cardiac death. Circulation. 2007;115(4):442–449.
- Akera T, Brody TM. Inotropic action of digitalis and ion transport. Life Sci. 1976;18(2):135–144.
- Hewett KW, Rosen MR. Alpha and beta adrenergic interactions with ouabain-induced delayed afterdepolarizations. J Pharmacol Exp Ther. 1984;229(1):188–192.
- Bers DM, Morotti S. Ca(2+) current facilitation is CaMKII-dependent and has arrhythmogenic consequences. Front Pharmacol. 2014;5(2+):144.
- Fan X, et al. Increased intracellular calcium concentration causes electrical turbulence in guinea pig ventricular myocytes. Sci China Life Sci. 2011;54(3):240–247.
- Faber GM, Rudy Y. Action potential and contractility changes in [Na(+)](i) overloaded cardiac myocytes: a simulation study. Biophys J. 2000;78(5):2392–2404.
- Herrera NT, et al. Dual effects of the small-conductance Ca2+-activated K+ current on human atrial electrophysiology and Ca2+-driven arrhythmogenesis: an in silico study. Am J Physiol Heart Circ Physiol. 2023;325(4):H896–H908.
- Bronk P, et al. Impact of ISK voltage and Ca2+/Mg2+-dependent rectification on cardiac repolarization. Biophys J. 2020;119(3):690–704.
- Alvarez JAE, et al. Local control model of a human ventricular myocyte: an exploration of frequency-dependent changes and calcium sparks. Biomolecules. 2023;13(8):1259.
- Bartolucci C, et al. Simulation of the effects of extracellular calcium changes leads to a novel computational model of human ventricular action potential with a revised calcium handling. Front Physiol. 2020;11:314.
- Grandi E, et al. Theoretical investigation of action potential duration dependence on extracellular Ca2+ in human cardiomyocytes. J Mol Cell Cardiol. 2009;46(3):332–342.
- Carmeliet E. Intracellular Ca(2+) concentration and rate adaptation of the cardiac action potential. Cell Calcium. 2004;35(6):557–573.
- Heinzen EL, et al. De novo mutations in ATP1A3 cause alternating hemiplegia of childhood. Nat Genet. 2012;44(9):1030–1034.
- Chahal CAA, et al. Systematic review of the genetics of sudden unexpected death in epilepsy: potential overlap with sudden cardiac death and arrhythmia-related genes. J Am Heart Assoc. 2020;9(1):e012264.
- Parisi P, et al. Coexistence of epilepsy and Brugada syndrome in a family with SCN5A mutation. Epilepsy Res. 2013;105(3):415–418.
- Otsomaa L, et al. Discovery and characterization of ORM-11372, a novel inhibitor of the sodium-calcium exchanger with positive inotropic activity. Br J Pharmacol. 2020;177(24):5534–5554.
- Parker LE, et al. Characterization of sedation and anesthesia complications in patients with alternating hemiplegia of childhood. Eur J Paediatr Neurol. 2022;38:47–52.
- Thamby J, et al. Characteristics of non-sleep related apneas in children with alternating hemiplegia of childhood. Eur J Paediatr Neurol. 2024;48:101–108.
- Kansagra S, et al. Polysomnography findings and sleep disorders in children with alternating hemiplegia of childhood. J Clin Sleep Med. 2019;15(1):65–70.
- Dagar S, et al. Evaluation of electrocardiographic parameters in patients with epileptic seizure. Acta Neurol Belg. 2020;120(2):321–327.
- Hunanyan AS, et al. Progressive central cardiorespiratory rate downregulation and intensifying epilepsy lead to sudden unexpected death in epilepsy in mouse model of the most common human ATP1A3 mutation. Epilepsia. 2025;66(3):899–913.
- Srour MK, et al. Children and adolescent patients with variants in the ATP1A3 -encoded sodium-potassium ATPase Alpha-3 subunit demonstrate an impaired QT response to bradycardia and predisposition to sinus node dysfunction [preprint]. https://doi.org/10.1101/2024.08.31.24312446 Posted on medRxiv October 31, 2024.
- Giladi M, et al. A gain-of -function SLC4A3 mutation causes short-QT syndrome: from molecular analysis to novel diagnostic testing. JACC Clin Electrophysiol. 2025;11(7):1583–1594.
- Hamayel H, et al. Cardioneuroablation for the treatment of vagal mediated ventricular fibrillation in short QT syndrome, case report. Pacing Clin Electrophysiol. 2025;48(9):1010–1013.
- Yuen GK, et al. Subcellular localization of Na/K-ATPase isoforms in ventricular myocytes. J Mol Cell Cardiol. 2017;108:158–169.
- Li M, et al. A functional correlate of severity in alternating hemiplegia of childhood. Neurobiol Dis. 2015;77:88–93.
- Sikkel MB, et al. Hierarchical statistical techniques are necessary to draw reliable conclusions from analysis of isolated cardiomyocyte studies. Cardiovasc Res. 2017;113(14):1743–1752.
-
Version history
- Version 1 (April 8, 2026): Electronic publication



![iPSC-CMD801N have miniwaves, higher SR [Ca2+] store content, and higher [Ca2+]i. iPSC-CMD801N have miniwaves, higher SR [Ca2+] store content, and higher [Ca](http://df6sxcketz7bb.cloudfront.net/manuscripts/197000/197721/small/jci.insight.197721.f3.gif)






