Research ArticleHematologyImmunology
Open Access |  10.1172/jci.insight.193686
10.1172/jci.insight.193686
Rare epigenetic alterations are conserved across hematopoietic differentiation stages after mycobacterial infection
Brandon T. Tran,1,2,3 Pamela N. Luna,4 Ruoqiong Cao,2,3,5 Duy T. Le,2,3,5 Apoorva Thatavarty,2,3,6 Laure Maneix,3,7,8 Bailee N. Kain,9 Scott Koh,2 Andre Catic,3,7,9,10 and Katherine Y. King2,3,5
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Tran, B. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Luna, P. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Cao, R. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Le, D. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Thatavarty, A. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Maneix, L. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Kain, B. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Koh, S. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by Catic, A. in: PubMed | Google Scholar
1Graduate Program in Cancer and Cell Biology,
2Department of Pediatrics, Division of Infectious Disease, Texas Children’s Hospital and Baylor College of Medicine,
3Stem Cells and Regenerative Medicine Center,
4Department of Molecular and Human Genetics,
5Graduate Program in Immunology and Microbiology,
6Graduate Program in Molecular and Human Genetics,
7Huffington Center of Aging, and
8Dan L. Duncan Comprehensive Cancer Center, Baylor College of Medicine, Houston, Texas, USA.
9Division of Experimental Hematology, Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
10Department of Molecular and Cellular Biology, Baylor College of Medicine, Houston, Texas, USA.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
Authorship note: BTT and PNL contributed equally to this work.
Find articles by King, K. in: PubMed | Google Scholar
Authorship note: BTT and PNL contributed equally to this work.
Published December 9, 2025 - More info
JCI Insight. 2026;11(2):e193686. https://doi.org/10.1172/jci.insight.193686.
© 2025 Tran et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: March 20, 2025; Accepted: December 3, 2025
-
Abstract
Infection leads to durable cell-autonomous changes in hematopoietic stem and progenitor cells (HSPCs), resulting in production of innate immune cells with heightened immunity. The mechanisms underlying this phenomenon, termed central trained immunity, remain poorly understood. We hypothesized that infection induces histone modifications leading to changes in chromatin accessibility that are conserved during differentiation from HSPCs to myeloid progenitors and monocytes. We conducted genome-wide surveillance of histone marks H3K27ac and H3K4me3 and chromatin accessibility in hematopoietic stem cells, multipotent progenitor 3, granulocyte-monocyte progenitors, and monocytes and macrophages of naive and Mycobacterium avium–infected mice. IFN signaling pathways and related transcription factor binding motifs including IRFs, NF-κB, and CEBP showed increased activating histone marks and chromatin accessibility across cell types. However, histone marks and increased chromatin accessibility were conserved at only a few loci, notably Irf1 and Gbp6. Knock out of IRF1 disrupted enhanced mitochondrial respiration and bacterial killing in human monocyte cell lines, while GBP6-KO monocyte cell lines showed dysregulated mitochondrial respiration. In summary, this study identifies IRF1 and GBP6 as 2 key loci at which infection-induced systemic inflammation leads to epigenetic changes that are conserved from HSPCs to downstream monocytes, providing a mechanistic avenue for central trained immunity.
-
Introduction
The adaptive and innate immune systems work in concert to prevent and fight infections. While the adaptive immune system traditionally has been considered the seat of immunological memory through the generation of memory lymphoid cells, emerging evidence indicates that innate immune cells, such as macrophages and neutrophils, contribute to immune memory (1, 2). Indeed, innate immune cells, when challenged by pathogens or pathogen-associated molecular patterns (PAMPs), undergo epigenetic reprogramming to elicit heightened secondary immune responses (1–3). Hallmarks of the innate immune memory immunophenotype include enhanced oxidative phosphorylation and lipid metabolism, proinflammatory cytokine production, phagocytosis, and pathogen clearance (4–7). Epidemiological studies and experimental vaccination trials in humans have further demonstrated that vaccination can confer improved nonspecific immune responses by innate effector cells (8–10).
Innate memory immune responses have been recorded for as long as 1 year after the initiating event in human studies, beyond the lifetime of most circulating innate immune cells (9, 11). This observation led to the speculation that long lived hematopoietic stem and progenitor cells (HSPCs) play a crucial role in the development and maintenance of innate immune memory. HSPCs, which consist of hematopoietic stem cells (HSCs) and 4 types of multipotent progenitors (MPPs 1–4), are responsible for lifelong blood production. HSPCs respond to environmental cues such as inflammatory cytokines, infection, diet, and aging (12–16). Furthermore, HSCs have been shown to harbor sustained epigenetic changes after infection (17–20). Several studies using functional assays and next-generation sequencing techniques found that HSPCs can be epigenetically altered to produce downstream immune cells with enhanced metabolism, cytokine production, and bacterial clearance (18, 21, 22). Even after serial transplantation into immunologically naive recipient mice, HSPCs continued to produce downstream macrophages with enhanced cross-protective functional capacity (18, 23, 24). Thus, inflammation-responsive HSPCs may be reprogrammed to promote increased myeloid differentiation and produce trained innate immune cells, a concept termed central trained immunity (CTI). Fully elucidating the specific mechanisms by which HSPCs confer CTI has, therefore, become a critical avenue by which to improve host immunity.
CTI is thought to be generated by inflammation-induced epigenetic reprogramming of histone modifications and other chromatin-modifying programs that poise the chromatin for faster and stronger immune responses. Studies have found that HSPCs and innate immune cells acquire activating histone modifications — including H3K4 mono- and trimethylation (H3K4me1, H3K4me3), and H3K27 acetylation (H3K27ac) — to promote myeloid differentiation and innate immune responses (18, 19, 21). However, how changes at the stem and progenitor level lead to changes in downstream cells remains unclear (3, 18, 25, 26). One potential mechanism is direct conservation of epigenetic modifications acquired by HSPCs (1, 2). Indeed, histone modifications can be passed down during DNA replication and are critical in cell function (27–30). Therefore, histone modifications may play a role in CTI generation, but few studies have provided direct evidence supporting this.
Here, we assessed whether sites of H3K4me3 and H3K27ac, modifications frequently enriched in CTI and reported to be conserved through cell division and DNA replication (29), and chromatin accessibility are conserved between HSPCs and innate immune cells after inflammatory stress. Utilizing a Mycobacterium avium infection model of CTI, we found that increased IFN-β cytokine signaling and associated transcription motifs were conserved across all cell types, underscoring the central role of cytokine signaling as a mediator of CTI. Activating histone modifications and chromatin accessibility changes were only conserved at a small number of genetic loci, suggesting that these loci are relevant in the establishment of CTI.
-
Results
M. avium infection induces long lasting central HSPC trained immunity to promote altered macrophage immune and metabolic function. In prior work, we found that M. avium–stimulated HSPCs generate bone marrow–derived macrophages (BMDMs) with heightened immunophenotypes, including increased cytokine production, bacterial clearance, and metabolic rewiring (22) (Figure 1A). Here, we confirmed those findings by showing that BMDMs derived from M. avium–trained HSPCs have increased maximal oxygen consumption rate (OCR) (P = 0.0355) and proton leak (P = 0.0207) (Figure 1, B–D). Increased OCR is linked to higher utilization of oxidative phosphorylation, while increased mitochondrial proton leak is associated with damaged mitochondria and ROS production (31, 32). We also found that M. avium–trained BMDMs produced significantly increased TNF-α (P = 0.002) and IL-6 (P = 0.0112), proinflammatory cytokines that can promote HSPC differentiation (33, 34) and commonly known to be secreted by trained macrophages (6, 24) (Figure 1, E and F).
 Figure 1
Figure 1Mycobacterium avium infection promotes HPSC central trained immunity with enhanced macrophage immune and metabolic function. (A) Experimental design for generating bone marrow–derived macrophages (BMDMs) from hematopoietic stem and progenitor cells (HSPCs). Cells were sorted from pooled mouse bone marrow samples (n = 5–7 per group), cKit-enriched, and sorted for lineage– cells. BMDMs generated from sorted cells were counted and seeded for respective assays after culture. (B) Seahorse Mitostress test assay in M. avium–trained BMDMs (red) and naive/untrained BMDMs (blue). Experiment was repeated twice. (C and D) Calculated maximum oxygen consumption rate (C) and proton leak (D) of M. avium–trained and naive BMDMs. (E and F) TNF-α (E) and IL-6 (F) secreted by M. avium–trained and naive BMDMs. Cytokine release assay performed once. (G) Experimental schematic for trained immunity chimeric transplant and challenge model. In total, 200,000 CD45.2 cKit+ cells from naive or M. avium–trained mice were transplanted with 200,000 CD45.1 rescue marrow into lethally irradiated CD45.1 recipients, which were challenged with M. avium 12 weeks after transplant. (H) Splenic M. avium colony forming units counts from M. avium–trained and naive challenged transplant recipients. n = 4–6 per experiment, representative findings from 2 independent experiments. Mean ± SEM shown per experiment. For statistical analysis (C, D, and H), 2-tailed Student’s unpaired t test. E and F used ordinary 1-way ANOVA. *P < 0.05, **P < 0.01.
To better understand how M. avium training affects host protection in vivo, we performed competitive transplants of HSPCs that were untrained or trained with M. avium infection into naive recipients (Figure 1G). Both groups produced trilineage hematopoiesis 3 months later, indicating engraftment in the transplant recipients (Supplemental Figure 1, A–C; supplemental material available online with this article; https://doi.org/10.1172/jci.insight.193686DS1). Recipients were then challenged with M. avium for 4 weeks, and splenic bacterial burden was measured as an indicator of host protection. While we did not detect any difference in mortality between the groups, we found significantly lower bacterial burden within spleens of M. avium–trained HSPC recipients compared with untrained controls (P = 0.006) (Figure 1H), validating prior findings (8, 11, 22, 35). Collectively, our work suggests that infection and inflammation reprogram HSPCs to produce trained innate immune cells with improved host protection capacity (9, 11, 21, 23, 24).
M. avium–infected HSPCs and monocytes/macrophages exhibit restricted overlap of H3K27ac acquisition. We hypothesized that changes in histone modifications are conserved through differentiation from early hematopoietic progenitors through myeloid progenitors and macrophages to mediate trained immunity. To determine if cells across 4 stages of differentiation share inflammation-induced histone modification changes, we performed cleavage under targets and release using nuclease sequencing (CUT&RUN-Seq) on cell populations from M. avium–infected and naive mice (Figure 2, A and B). We targeted HSCs, myeloid biased multipotent progenitor 3s (MPP3s), granulocyte-monocyte progenitors (GMPs), and monocytes/macrophages (Mo/Macs). H3K27ac CUT&RUN-Seq datasets we previously generated from M. avium–infected and naive HSCs were used for differential analysis (25).
 Figure 2
Figure 2M. avium–infected HSPCs and MPP3s exhibit significant increases in H3K27ac at genomic loci associated with antigen presentation and differentiation. (A) Workflow of acquiring cells for cleavage under targets and release using nuclease sequencing (CUT&RUN-Seq). Experiment was performed twice. (B) Bioinformatic pipeline performed on CUT&RUN-generated libraries. (C and D) Gene ontology of biological processes associated with gene loci with significantly increased H3K27ac presence in HSCs (C) or MPP3s (D) from M. avium–infected mice. (E–G) Genomic tracks of H3K27ac presence at Dusp2 (E), H2-Eb1 (F), or Irf1 (G) in HSPCs and Mo/Macs in both M. avium and naive conditions. (H) Histone H3K27ac enrichment heatmap of selected gene loci across M. avium–infected and naive HSPC subpopulations and macrophages, representative of 2 independent biological replicates. Associated genes in red were significant in at least 3 cell types in respective differential analyses. Bioconductor package Diffbind was used for statistical comparison of M. avium versus naive state. Cutoff FDR < 0.05 is considered significant. *P < 0.05, **P < 0.01, ***P < 0.001.
Profiling of cell populations through PCA plotting and Pearson correlation (Supplemental Figure 2, A and B) showed that our cell populations clustered as expected based on cell type. Cell type was a stronger driver of cell clustering than infection, as cells from naive and infected animals were interspersed in the PCA. MPP3s and GMPs were similar, with Pearson correlation blending both populations (Supplemental Figure 2B), consistent with their shared role as myeloid-biased progenitors.
Peakcalling of the samples in both M. avium–infected and naive states revealed that all cell types had high levels of H3K27ac histone modifications across the chromatin landscape for general biological processes important for maintaining cellular function. For example, all HSPCs contained H3K27ac modifications at gene loci associated with stemness, including Adipor2, Irf2bp2, and Spi1 (Supplemental Figure 2, C–E). Overall, H3K27ac datasets captured loci expected to be essential for hematopoietic cellular functions.
We next performed differential analysis, comparing each M. avium–infected subpopulation to their naive counterpart. These comparisons were performed to determine whether any changes seen at 1 subpopulation were consistent through differentiation. HSCs displayed a significant increase in H3K27ac modifications at regions associated with immune system processes (FDR = 7.05 × 10–3), definitive hemopoiesis (FDR = 6.72 × 10–3), and regulation of cell cycle (FDR = 4.07 × 10–3) (Figure 2C and Supplemental Tables 1 and 2). These included Dusp2 (FDR = 2.3 × 10–2, FC = 2.29), Irf1 (FDR = 4.2 × 10–3, FC = 2.62), and Hoxb4 (FDR = 2.9 × 10–2, FC = 1.95) (Figure 2, E and G, Supplemental Figure 2F, and Supplemental Tables 1 and 2). Additionally, H3K27ac was increased at MHC-II antigen processing and antigen presentation genes (FDR = 1.23 × 10–5) like H2-Aa (FDR = 7.95 × 10–4, FC = 3.71), H2-Eb1 (FDR = 4.28 × 10–4, FC = 4.45), and H2-Ab1 (FDR = 2.5 × 10–2, FC = 2.01) (Figure 2F; Supplemental Figure 2, G and H; and Supplemental Tables 1 and 2). These findings are consistent with reports that HSCs harbor antigen processing and presentation capacity (36).
These changes in H3K27Ac were mirrored in MPP3s (Figure 2D). Like HSCs, MPP3s exhibited significant increases of H3K27ac at H2-Aa (FDR = 7.26 × 10–3, FC = 1.77), H2-Eb1 (2.75e-04, FC = 2.11), and Irf1 (FDR = 1.48 × 10–3, FC = 1.82) (Figure 2, F and G, Supplemental Figure 2, G–J, and Supplemental Tables 3 and 4). MPP3s uniquely displayed increases of H3K27ac at Cebpb (FDR = 1.14 × 10–2, FC = 1.58) (Supplemental Figure 2K), a pioneer factor that is critical in LPS-driven HSPC-trained immunity (18).
By comparison, GMPs showed minimal increases in H3K27ac, though many loci had high baseline acetylation (Supplemental Tables 5 and 6). Notably, Mo/Macs from M. avium–infected mice exhibited a significant increase in H3K27ac compared with naive Macs at Irf1 (FDR = 5.35 × 10–4, FC = 3.73) (Figure 2G and Supplemental Table 7), a critical mediator of IFN responses in Macs (17, 37).
Collectively, we did not find any loci with significantly increased H3K27ac across all 4 stages of myeloid differentiation, indicating that H3K27ac alone cannot account for CTI conservation in myeloid progenitors (Figure 2H). However, increased H3K27ac was conserved in M. avium–infected HSCs, MPP3s, and Mo/Macs at Irf1, one of the few sites with increased H3K27ac across several stages of differentiation.
HSPCs and Mo/Macs share H3K4me3 modifications at loci associated with IFN responses and GTPase function. We next performed CUT&RUN-Seq analysis for H3K4me3 (Supplemental Figure 3, A and B). Cells clustered similarly to our H3K27ac datasets, with MPP3s and GMPs blending based on Pearson correlation (Supplemental Figure 3B).
As expected, H3K4me3 marks were found at loci associated with basic cellular function in all cell populations in both naive and M. avium–infected states, including known HSPC regulators like Spi1/Pu.1 and Runx1 (Supplemental Figure 3, C and D). Interestingly, many inflammatory response genes, specifically IFN signaling pathway genes Ifnar1, Stat1, and Stat2, and Tet2 had steady H3K4me3 signal in all cell types and states (Supplemental Figure 3, E–I). These findings reinforce the concept that tonic levels of IFN signaling within the bone marrow contribute to steady state hematopoiesis (38–40).
In HSCs, there were significant increases in H3K4me3 at biological processes associated with responses to IFN-β (FDR = 9.69 × 10–6) (Figure 3A and Supplemental Tables 8 and 9). Many of these enriched loci were closely tied to GTPase activity (FDR = 8.89 × 10–3) and GTP binding (FDR = 1.59 × 10–2) (Supplemental Tables 8 and 9), including Gm12250 (FDR = 2.3 × 10–2, FC = 1.82), Gm4951 (FDR = 5.64 × 10–3, FC = 2.91), Iigp1 (FDR = 1.88 × 10–3, FC = 3.83), and guanylate binding protein 6 (Gbp6) (FDR = 2.32 × 10–2, FC = 2.72) (Figure 3, D–G, Supplemental Figure 3J, and Supplemental Tables 8 and 9). While several of these GTPases are mouse specific, GBP6 and GBP family proteins have human homologs that are reported to be crucial in mycobacterial immunity (41, 42).
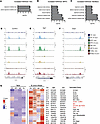 Figure 3
Figure 3M. avium–infected HSPCs and macrophages exhibit significant increases in H3K4me3 at genomic loci associated with responses to IFN-β. (A–C) Gene ontology of biological processes associated with gene loci with significantly increased H3K4me3 presence in HSCs (A), MPP3s (B), or Mo/Macs (C) from M. avium–infected mice. (D–F) Genomic tracks of H3K4me3 presence at Gm4951 (D), Iigp1 (E), or Gbp6 (F) in HSPCS and Mo/Macs in both M. avium–infected and naive conditions. (G) Histone H3K4me3 enrichment heatmap of selected gene loci across M. avium–infected and naive HSPC subpopulations and macrophages, representative of 2 biological replicates. Associated genes in red were significant in at least 3 cell types in respective differential analyses. Data compiled from 2 independent experiments. Bioconductor package Diffbind was used for statistical comparison of M. avium versus naive state. Cutoff FDR < 0.05 is considered significant. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Much like HSCs, MPP3s from M. avium–infected mice had significant increases in H3K4me3 at the same set of GTPases (Figure 3, D–G; Supplemental Figure 3D; and Supplemental Tables 10 and 11) and immune processes including responses to IFN-β (FDR = 1.82 × 10–11), type II IFN (FDR = 3.36 × 10–6), and innate immune responses (FDR = 3.71 × 10–3) (Figure 3B and Supplemental Tables 10 and 11). Immune response genes such as Ifitm1 (FDR = 2.36 × 10–6, FC = 2.44), Batf2 (FDR = 4.33 × 10–3, FC = 2.55), Serpina3f (FDR = 3.24 × 10–2, FC = 1.44), and Serpina3g (FDR = 6.27 × 10–8, FC = 2.25) were also significantly increased upon M. avium infection (Figure 3G; Supplemental Figure 3, K–M; and Supplemental Tables 10 and 11). GMPs from M. avium–infected mice showed increased H3K4me3 modification at Gm4951 (FDR = 9.71 × 10–8, FC = 3.99), Serpina3f (FDR = 1.06 × 10–5, FC = 3.78), and Serpina3g (FDR = 7.63 × 10–4, FC = 1.75), while H3K4me3 modification was unchanged at Gbp6 and Iigp1 (Figure 3, D and G; Supplemental Figure 3M; and Supplemental Tables 12 and 13). Like H3K27ac, the H3K4me3 mark was present at baseline in GMPs at many of the sites at which we found increases in other cell populations, reducing the sensitivity of the differential analysis in this cell type (Supplemental Tables 12 and 14).
Mo/Macs from M. avium–infected mice also showed consistent increases in H3K4me3 associated with responses to IFN-β (FDR = 2.1 × 10–8) (Figure 3C and Supplemental Tables 15 and 16) and at GTPases like Gm4951 (FDR = 4.62 × 10–5, FC = 2.98), Iigp1 (FDR = 6.07 × 10–5, FC = 2.99), and Gbp6 (FDR = 1.56 × 10–4, FC = 2.63) (Figure 3, D–G). Ciita (FDR = 3.83 × 10–3, FC = 1.58), a master transcription factor (TF) of MHC-II antigen presentation, showed increased H3K4me3 in this analysis (43) (Supplemental Figure 3N and Supplemental Tables 15 and 16). Interestingly, H3K4me3 marks were decreased in Mo/Macs upon infection at some gene loci associated with metabolism, DNA repair, and macromolecule processing (Supplemental Tables 15 and 17), while other regions such as IL-27 (Il-27), a multifunction cytokine that can potentially elicit Th1 responses, were unchanged (44) (Figure 3G and Supplemental Figure 3O).
Overall, we discovered a limited number of loci with increased H3K4me3 conserved across all 4 stages of myeloid differentiation upon M. avium infection, including GTPases Gm4951 and Gbp6 that mediate IFN-β responses (Figure 3G). To determine whether these findings were specific to M. avium infection, we tested whether H3K4me3 marks were induced at Gbp6 after BCG exposure. CUT&RUN qPCR of Gbp6 showed similar acquisition of H3K4me3 after either M. avium or BCG infection, indicating a common response to both training agents (Supplemental Figure 3P).
Loci associated with IFN responses and GTPase function showed increased chromatin accessibility in HSPCs and Mo/Macs upon M. avium infection. To understand how the overall chromatin accessibility landscape is changed in HSPCs upon M. avium infection, we performed single-cell assay for transposase accessibility sequencing (scATAC-Seq) on targeted HSPCs and downstream terminally differentiated cells, including Mo/Macs, neutrophils, and B cells, from pooled mouse samples (Figure 4A and Supplemental Figure 4A). To mirror our prior single-cell transcriptomic study using the same infection model (22), we specifically isolated long-term HSCs (LT-HSCs) and myeloid-biased subpopulations, including CD41+ HSCs, MPP3s, and GMPs, to acquire sufficient cell numbers for differential analyses and enhance representation of rare cell types. We found similar clustering in both M. avium–infected and naive states, with HSCs near MPPs and GMPs clustering near Mo/Macs subpopulations (Figure 4B). Neutrophils were comparatively distant from the HSPC clusters, indicating a more distinct chromatin landscape. Like our CUT&RUN-Seq datasets, GMPs and MPP3s blended together in the predicted UMAPs. Interestingly, we saw a significant separation between common lymphoid progenitors (CLPs) and B cells, suggesting that the chromatin landscape of lymphoid cells changes extensively during terminal differentiation.
 Figure 4
Figure 4HSPCs and monocytes exhibit distinct chromatin accessibility changes upon M. avium infection. (A) Workflow schematic for acquiring targeted cell populations for scATAC-Seq. (B) Predicted UMAP plotting of targeted HSPC and lineage subpopulations in both naive and M. avium–infected states. (C) Heatmap accessibility of peaks found to be significantly changed in at least 1 cell type, with asterisks representing the level of significance at each gene locus in a specific cell population. (D and E) Gene ontology enrichment analysis in GMPs (D) and monocytes (E). (F) Genomic coverage plot of Gbp6. Cell populations with significant increases were marked with an asterisk. Peaks that are significantly differentially accessible (Padj < 0.05) between M. avium–infected and naive states in at least 1 cell type are highlighted with an asterisk.
Peakcalling revealed chromatin regions that were consistently accessible across progenitors and differentiated myeloid cells independent of M. avium infection. These included regions such as Jun and Irf1, both TFs we have previously reported in HSPC transcriptional responses to M. avium infection (17) (Supplemental Figure 4, B and C). These data indicate that the chromatin is already open at these loci, with induction of transcription depending more on additional factors such as TFs or acquisition of active histone modifications (17, 18).
We next performed differential analysis to identify chromatin accessibility changes upon M. avium infection. Gene ontogeny analysis revealed that no biological processes were significantly more accessible in M. avium–infected LT-HSCs (Supplemental Table 18). However, the antigen presentation gene Cd74 had increased accessibility in LT-HSCs upon M. avium infection (Padj = 2.04 × 10–9, log2FC ~2.17) (Supplemental Figure 4D and Supplemental Table 18). M. avium–infected MPP3s exhibited changes in chromatin accessibility at genomic loci identified in our CUT&RUN-Seq analysis (Supplemental Table 19). These included Iigp1 (Padj = 3.24 × 10–7, log2FC ~4.23), Serpina3g (Padj = 1.18 × 10–2, log2FC ~0.88), Gm4951 (Padj = 1.16 × 10–2, log2FC ~4.50), and Gm4841 (Padj = 4.66 × 10–2, log2FC ~3.59) (Figure 4C, Supplemental Figure 4, E–G, and Supplemental Table 19).
Comparatively, we found many more chromatin accessibility changes in GMPs and Mo/Macs upon M. avium infection. IFN-associated biological responses such as response to protozoan and bacterium and response to cytokines were increased in GMPs upon infection (Figure 4D and Supplemental Table 20). GTPases that we previously noted including GBPs; Gbp6 (Padj = 3.24 × 10–9, log2FC = 3.41), Gbp8 (Padj = 6.47 × 10–6, log2FC = 4.55), Iigp1 (Padj = 4.26 × 10–5, log2FC = 9.26), Irgm2 (Padj = 2.20 × 10–4, log2FC ~1.59), Serpina3g (Padj = 6.92 × 10–6, log2FC = 1.41), and Serpina3f (Padj = 5.03 × 10–3, log2FC = 1.12) had significantly increased chromatin accessibility in GMPs from M. avium–infected mice (Figures 4, C and F, Supplemental Figure 4F, and Supplemental Table 20). Many of these loci demonstrated histone modifications at the MPP3 level in the CUT&RUN-Seq, leaving open the possibility that epigenetic changes occurring at those loci may have contributed to chromatin openness at the downstream GMP level.
Significant changes in chromatin accessibility were seen in Mo/Macs from M. avium–infected mice at loci similar to GMPs, including Gbp6 (Padj = 5.57 × 10–10, log2FC = 1.95), Gbp8 (Padj = 2.95 × 10–10, log2FC = 2.46), Serpina3g (Padj = 6.15 × 10–14, log2FC = 1.95), and Serpina3f (Padj = 2.00 × 10–8, log2FC ~1.37) (Figure 4, C and F, Supplemental Figure 4G, and Supplemental Table 21). Additional GBPs — Gbp7 and Gbp2 (Padj = 2.62 × 10–3, log2FC ~0.966), Gbp9 and Gbp4 (Padj = 1.17 × 10–2, log2FC = 0.759), and secreted factors (Il12a (Padj = 3.69 × 10–4, log2FC ~0.324), Il12b (Padj = 1.299 × 10–6, log2FC = 1.08), Il9 (Padj = 4.43 × 10–3, log2FC = 1.09) — were also increased (Supplemental Table 21). We also found Ccl5, a chemokine found associated with COVID-19, HIV, and HSC IFN responses (25, 26, 45), had increased in accessibility in Mo/Macs (Padj = 3.72 × 10–6, log2FC = 0.557) (Supplemental Table 21). Gene set enrichment analysis indicated that biological processes associated with immune responses, including response to cytokine and responses to IFN-γ were significantly more accessible (Figure 4E and Supplemental Table 21). The inflammatory gene set “response to lipopolysaccharide,” including loci such as at Cebpe (Padj = 8.39 × 10–4, log2FC = 0.764), Icam1 (Padj = 9.53 × 10–3, log2FC = 0.502), Nfkbiz (Padj = 2.91 × 10–9, log2FC ~1.58), Tnfaip3 (Padj = 1.96 × 10–3, log2FC = 1.01), and Tnfrsf1b (Padj = 2.95 × 10–3, log2FC = 0.342) was also enriched (Figure 4E and Supplemental Table 21). Antigen processing genes Cd74 (Padj = 5.89 × 10–8, log2FC = 0.586) and H2-Aa (Padj = 5.36 × 10–3, log2FC = 0.807) were also enriched in Mo/Macs (Supplemental Figure 4, D and G, and Supplemental Table 21). Overall, these findings affirm that Mo/Macs undergo changes in the chromatin landscape to elicit inflammation-mediated immune responses.
Collectively, these findings reveal only a few sites with consistently open or increased chromatin accessibility across 4 stages of myeloid differentiation. Notably, IFN-stimulated TFs like Jun and Irf1 had consistent accessibility, while GTPases like Gm4951, which was significant in MPP3s and Mo/Macs, had partial increases in all HSCs and GMPs without reaching statistical significance (Supplemental Figure 4E). Gbp6, a locus that demonstrated increased H3K4me3 across several cell stages, gained chromatin accessibility in HSPCs, GMPs, and Mo/Macs (Figure 4F). Consistent with the CUT&RUN data, these areas of conserved increased chromatin accessibility lay in IFN and immune response genes.
HSPC subpopulations and Mo/Macs from M. avium–infected mice have conserved enrichment of motifs associated with IFN and NF-κB signaling. TFs are DNA-binding proteins known to participate in coactivator/corepressor recruitment, transcription complex formation initiation, DNA looping, polymerase II recruitment, and — more recently — pioneer factors that initiate and maintain chromatin openness (46–48). Studies have shown that TFs may play a pivotal role in the epigenetic memory of various long-living cell populations, including skin and blood stem cells (18, 49, 50). Notably, CEBPβ has been reported as a pioneer factor crucial for promoting transcription and chromatin accessibility of genes associated with LPS-stimulated HSPC trained immunity (18). Given these findings, we evaluated our scATAC-Seq dataset to determine how M. avium infection affects TF function across stages of myeloid differentiation.
We found a significant increase of inflammatory TF motifs across all cell populations upon M. avium infection. Among HSCs, we expected to find IRF and STAT signaling motifs, as mycobacterial infections strongly induce IFN signaling in HSPCs (17, 21, 24, 51, 52). Indeed, HSCs from M. avium–infected mice were enriched in inflammatory motif signatures associated with IFN signaling — STAT1::STAT2 (Padj = 1.45 × 10–3, FC = 2.46), IRF1 (Padj = 8.22 × 10–50, FC = 2.07), STAT2 (Padj = 8.09 × 10–39, FC = 2.14), IRF9 (Padj = 3.90 × 10–36, FC = 3.09), IRF8 (Padj = 1.60 × 10–30, FC = 3.34). IRF9 and STAT1/2 factors are known to combine to create the stimulated gene factor 3 (ISGF3) complex for canonical type I IFN signaling, and this paradigm is supported by our H3K4me3 CUT&RUN differential analysis. Additional binding motifs associated with NF-κB signaling — RELA (Padj = 3.03 × 10–29, FC = 3.50), NFKB1 (Padj = 2.43 × 10–17, FC = 3.78), NFKB2 (Padj = 3.03 × 10–14, FC = 3.34), CEBP signaling (CEBPG (var.2) (Padj =6.17 × 10–38, FC = 4.07), and CEBPD (Padj = 1.63 × 10–37, FC = 3.89) — and master regulator Runx1 (Padj = 8.61 × 10–34, FC = 2.31) were all enriched in HSCs from infected mice (Figure 5A and Supplemental Table 22). These findings were not unexpected, given that NF-κB and CEBP can be activated upon IFN signaling via PI3K/Akt signaling and MAPK signaling, respectively (53–55). Independently, NF-κB signaling via TNF-α has been recently reported to activate IRF1 and IRF3 (56), further highlighting an interconnection between IRFs and NF-κB activity.
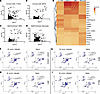 Figure 5
Figure 5M. avium–infected HSPC subpopulations and monocytes have a conserved, increased enrichment of motifs associated with IFN and NF-κB signaling. (A–D) Volcano plot of motifs significantly enriched in M. avium–infected LT-HSCs (A), MPP3s (B), GMPs (C), and monocytes (D) versus naive controls. Motifs identified based on scATAC-Seq datasets. (E) Heatmap showing the highest average log fold enriched motifs across M. avium–infected LT-HSCs, MPP3s, GMPs, and macrophages. Cutoffs for filtering included P < 1 × 10–20 and LFC > 2 observed in any cell type. (F–I) Motif enrichment UMAP plots of STAT1::STAT2 (F), IRF9 (G), IRF1 (H), and RELA (I) in infected and naive states of HSPC subpopulations. Volcano plots (A–D) were generated for significantly enriched motifs with Padj < 0.05 and FC > 1.
These sets of TF family motifs were strongly enriched in myeloid progenitors and Mo/Macs from M. avium–infected mice. MPP3s and GMPs exhibited significant enrichment of IRFs, NF-κBs, and CEBP families upon infection (Figure 5, B and C, and Supplemental Tables 23 and 24). Among Mo/Macs, the strongest enrichment occurred in NF-κB motifs, including NFKB1 (Padj = 3.91 × 10–28, FC = 4.52), NFKB2 (Padj = 4.32 × 10–22, FC = 3.90), RELA (Padj = 2.25 × 10–51, FC = 4.35), and REL (Padj = 6.90 × 10–44, FC = 4.26), though IRF and CEBP families were also enriched (Figure 5, D and E, and Supplemental Table 25). Collectively, IRF, CEBP, and NF-κB TF families were some of the most enriched motifs across HSPCs, GMPs, and Mo/Macs, suggesting that the activity of these TFs could play a prominent role in CTI (Figure 5, E–I, and Supplemental Tables 22–25). This conserved enrichment could only be observed through Mo/Mac differentiation, whereas neutrophils showed little enrichment of these motifs upon M. avium infection. Separately, these motifs were constitutively present in lymphoid cell populations (Figure 5, F–H). Thus, TF motif analysis affirms strong conservation of IFN, CEBP, and NF-κB signaling among HSPCs, GMPs, and Mo/Macs.
H3K4me3 modifications acquired upon M. avium training are transient. Given that M. avium–trained HSPCs, myeloid progenitors, and Mo/Macs acquired highly conserved H3K4me3 modifications and increased chromatin accessibility at loci associated with IFN-β, we asked whether this histone mark remained stable in rested, M. avium–trained HSPCs. To answer this question, we transplanted M. avium–trained or untrained HSPCs into lethally irradiated CD45.1 mice and waited 12 weeks to assess the H3K4me3 modification landscape (Figure 6A). This transplant method removes the HSPCs and Mo/Macs from the infected environment and provides sufficient time for them to return to steady state. As shown in Figure 1, donor BMDMs from similarly transplanted recipients retain persistent functional changes and support improved bacterial clearance, indicating CTI. Upon CUT&RUN-Seq, we found that H3K4me3 did not persist in progenitor or downstream Mo/Mac populations (Figure 6A). IFN-β associated loci including Iigp1, Gm4591, and Gbp6 showed little to no H3K4me3 presence in both untrained and M. avium–trained, rested states (Figure 6, B–D). Thus, while our findings point toward IFN-β–associated loci as key mechanistic components for trained immunity phenotypes, absence of H3K4me3 in the resting state indicates that this is not the relevant epigenetic mark to encode this memory.
 Figure 6
Figure 6Histone modification H3K4me3 acquired in M. avium training do not persist in HSPCs that are transplanted and rested. (A) Workflow of performing CUT&RUN on H3K4me3 in HSPCs and Mo/Macs that had been rested and transplanted into lethally irradiated recipients. In total, 200,000 CD45.2 M. avium–trained or untrained HSPCs (cKit+) were transplanted with 200,000 CD45.1 rescue marrow into lethally irradiated recipients. (B–D) Genomic tracks of Iigp1 (B), Gm4951 (C), and Gbp6 (D) in either untrained, rested or M. avium–trained, rested HSPCs, and Mo/Macs acquired from transplanted recipients. Data compiled from 2 independent experiments. MACS2 and Diffbind were utilized for analysis.
Loss of GBP6 affects metabolic functions of PMA-treated THP-1 Macs, while loss of IRF1 negatively affects both metabolic and immune function. Our findings highlight that modifications or chromatin openness at loci related to IFN signaling, specifically at Irf1 and Gbp6, are conserved across myeloid differentiation and may, thus, contribute directly to CTI. To assess the functional role of these 2 proteins in human Mo function, we utilized CRISPR-Cas9 editing to excise catalytic domains of GBP6 and IRF1, respectively, from THP-1 cells, a human monocytic cell line (Figure 7A). Deletion of GBP6 or IRF1 was confirmed by PCR in single-cell–derived colonies (Figures 7, B and C), and KO cell lines were expanded for phenotypic testing (57). Both GBP6-KO and IRF1-KO cells successfully differentiated, with consistent total cell numbers, viability, and acquisition of CD14, a marker of mature human Macs (57) (Figure 7D and Supplemental Figure 5, A and B). These findings indicate that KO of IRF1 or GBP6 does not affect Mo-to-Mac maturation or overall viability.
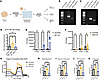 Figure 7
Figure 7Loss of GBP6 affects cellular respiration in THP-1–KO macrophages independent of innate immune function, while loss of IRF1 negatively affects many macrophage functions. (A) Workflow of knocking out GBP6 and IRF1 in THP-1 cells, followed by their maturation into macrophages for immunophenotyping. KO colonies were derived from single CRISPR-edited cells for functional assessment. (B and C) Representative gel image of exon 2 of GBP6 (B) and exon 4 of IRF1 (C). (D) Fold change of CD14 mean fluorescence intensity upon 3-day PMA-treatment (20 ng/mL), compared with uninduced. n = 4 independent experiments. (E and F) Bacterial CFU from M. avium–challenged THP-1 cells (100,000 THP-1 cells per well, M. avium MOI 1) for 4 hours, 3 days, or 5 days. IRF1 KO versus scramble control (E) and GBP6-KO versus scramble control (F). n = 3–6 per group at each time point. (G) Seahorse mitostress assay of Scramble PMA-treated scramble (black), GBP6-KO (gold), and IRF1-KO (blue) THP-1 cells. (H–K) Calculated basal respiration (H), maximum respiration (I), ATP production (J), and proton leak (K) of PMA-treated THP-1 cells. All experiments were performed twice independently. Two-way ANOVA with Šídák’s multiple comparisons was used for D and E, while 1-way ANOVA was used for H–K. ****P < 0.0001.
Next, we challenged KO cell lines with M. avium to determine whether loss of GBP6 or IRF1 would affect bacterial killing function. Compared with controls, M. avium growth was significantly increased in challenged IRF1-KO wells by the day 5 time point (Figure 7E), suggesting an impaired bacterial killing capacity. In contrast, we found no difference in bacterial growth in challenged GBP6-KO cells compared with scramble controls at all time points (Figure 7F).
We then asked whether the loss of GBP6 and IRF1 affected mitochondrial respiration, a key factor in innate immune responses and trained immunity (4, 58). Utilizing the seahorse mitostress test on PMA-treated THP-1 cells (Figure 7F), we found that baseline and maximum respiration of IRF1-KO cells were significantly decreased compared with controls (Figure 7, G and H), indicating impaired oxidative phosphorylation. Furthermore, IRF1-KO cells also demonstrated a significant decrease in ATP production (Figure 7I). Notably, IRF1-KO cells had a subtle but significant decrease in proton leak compared with controls, suggesting that the decreased respiration is not due to damaged mitochondria (Figure 7J). In contrast, GBP6-KO cells had significantly increased respiration across all parameters measured (Figure 7, H–J). However, proton leak was also significantly increased in GBP6-KO cells, indicating mitochondrial damage that may be causing dysregulated metabolic function (Figure 7K). Together, these KO studies highlight that IRF1 has an essential role in Mac killing capacity and metabolism, while GBP6 plays a regulatory role in mitochondrial respiration.
-
Discussion
CTI is a recently described phenomenon in which durable cell autonomous changes in HSPC lead to the generation of downstream innate immune cells with increased metabolism, cytokine production, and pathogen killing capacity. While many studies indicate that CTI enhances host immunity (18, 21, 22, 59), it also can be maladaptive, driving autoinflammation in conditions like long COVID (26, 60–63). The cell-autonomous nature of this phenomenon has been demonstrated through transplant studies and human epidemiologic studies (10, 11, 18); however, the mechanisms by which CTI is encoded within HSPCs remain unclear. Improving our understanding of how CTI develops is a critical advance to determine whether it can be manipulated to either promote host immunity or stop its maladaptive consequences. Here we evaluated whether chromatin modifications, such as histone modifications, chromatin accessibility, and TF motif enrichment, are conserved through stages of innate immune cell differentiation after infection, thereby pinpointing key mechanistic mediators of innate immune memory.
Using an established model of M. avium infection, we validated that immune training confers cell autonomous changes on HSPCs, leading to the generation of BMDMs with heightened inflammatory responses. Our findings are consistent with prior work showing that mycobacterial training produces HSPCs that provide protection against subsequent infectious challenges and can be cross-protective against antigenically distinct infections, arguing against an adaptive immune mechanism (8, 18, 22, 59).
Both CUT&RUN analysis of H3K4me3 and single-cell ATAC-Seq analysis demonstrated that IFN signaling was significantly increased in M. avium–trained HSPCs, MPP3s, GMPs, and Mo/Macs. Notably, the strongest responses were found in the myeloid progenitors, similar to what has been demonstrated after BCG training in humans (19). Furthermore, motif enrichment analysis revealed IRFs to be some of the highest ranked enriched motifs across the cell types. Inflammatory motifs representing NF-κB signaling and CEBP signaling, which can be noncanonically activated by IFNs (18, 54, 55, 64, 65), were also highly enriched. Notably, NF-κB, IRF1, and ISGF3 form a network of TFs that has been reported in LPS and IFN-γ training of BMDMs to contribute to the development of H3K4me1 de novo superenhancers (56). This motif network is found not only in mouse models but also in patients recovered from SARS-CoV-2, which also show a consistent increase in IRF and NF-κB binding motifs (26). Additionally, CEBP and IRF TFs are highly enriched in unique myeloproliferative neoplasm subpopulations that emerge upon type I IFN therapeutics (66). Our findings, summarized collectively in Table 1, further support a critical role for this conserved signaling network in CTI.
With CUT&RUN-Seq, we determined the histone modification landscape of H3K27ac and H3K4me3 in HSPCs and Mo/Macs in both naive and M. avium–infected states. Recent studies have suggested that H3K27ac and H3K4me3 marks critically prime key CTI loci at the progenitor level (18, 67) and that these marks are passed to downstream effector Mo/Macs to produce trained immune memory. However, our data do not strongly support this hypothesis, as we find that most H3K4me3 marks do not persist in the resting stage after infection, and H3K27ac marks are known to be very short-lived (68). Other forms of epigenetic modification or stable TF networks may support CTI. For example, a study reported the role of lactylation as a durable, novel modification critical for the generation and persistence of CTI in BCG-vaccinated reprogramming (67). Assessment of this and other durable marks at sites we identified through scATAC-Seq analysis will likely improve our understanding of the structural basis of CTI.
Increased H3K27ac was conserved at Irf1 across HSCs, MPP3s, and Mo/Macs upon M. avium infection, and the IRF1 binding motif was also highly enriched across all 4 cell types upon infection. While H3K27ac tends to be a transient marker that persists for less than a day (68), the active production of IRF1 during M. avium infection (17) and its high level of motif availability indicates IRF1 as a central mediator of inflammatory responses relevant to CTI. Prior studies have demonstrated that TFs including CEBPβ (18) in HSPCs and AP1 TFs in epithelial skin stem cells and fibroblasts (69, 70) can serve as critical pioneer factors in trained immunity. IRF1 is essential for both HSPC inflammatory–induced differentiation responses (17, 26, 71, 72) and Mac responses involved in IFN responses and innate immune function responses to viral and bacterial responses (37, 71, 73–76). Much as CEBPβ plays a multifunctional role in processing and relaying inflammatory signaling, we believe that IRF1 also serves as a central hub that relays many inflammatory signals to elicit the innate immune response. Indeed, we found that CRISPR KO of IRF1 severely impairs human THP1 Mo and Mac functions, consistent with prior reports of inborn errors of immunity (IEI) (37, 74). Given IRF1’s critical role in promoting gene transcription of metabolic and inflammatory pathways, including production of TNF-α, we speculate that loss of IRF1 function severely dampens the inflammatory reprogramming of HSPCs necessary for CTI (77). Notably, Moorlag et al. reported increased chromatin accessibility at other IRF family members, notably IRF5 and IRF8, in human Mos after BCG vaccination (78), highlighting the relevance of this family.
Consistent with the IFN theme, we found conserved acquisition of H3K4me3 in genes associated with biological responses to IFN-β. GTPases such as Gm4951, Iigp1, Gm4841, and Gbp6 were prominent among these genes. While many of these GTPases are found only in mice, guanylate binding proteins, including GBP6, are reported to be essential for immunity in both mice and humans. For example, silencing of GBP6/GBP10 decreases immune responses in Macs challenged with Mycobacterium bovis (41, 42, 79–81). Notably, GBP4 and GBP5 confer transcriptional memory in HeLa cells responding to IFN-γ, suggesting that GBP family proteins play a role in inflammation-induced immune memory (82, 83). In our scATAC-Seq analysis, Gbp6 gained increased chromatin accessibility across HSPCs and Mo/Mac cell stages. GBP6-KO studies revealed that loss of GBP6 affected metabolic respiration but not bacterial killing, in contrast to prior studies (42, 79). Specifically, we found an overall increase in mitochondrial respiration in GBP6-KO THP1 like Macs, possibly due to higher demands on oxidative phosphorylation at the expense of mitochondrial integrity. We speculate that this overall increase in mitochondrial respiration may be a compensatory mechanism for the damaged mitochondria in GBP6-KO cells. Our findings indicate a possible regulatory role for GBP6 in mitochondrial respiration independent of intracellular pathogen responses. These findings set the stage for future studies on the role of GBP6 in Mac function and CTI (84). Interestingly, GBP6 has been reported to be transcriptionally regulated by IRF1, linking regulation of these 2 key genes (74).
Limitations of the study In this study, we focused our CUT&RUN-Seq on 2 common histone modifications, H3K27ac and H3K4me3, which have been speculated to contribute to CTI in prior studies. However, many other chromatin modifications, both silencing and activating, may affect gene transcription. Careful examination of other marks, guided by the targets identified in this study, will be important in future studies. In scATAC-Seq, it should be noted that we sorted various cell populations to ensure adequate representation; therefore, frequencies of cells observed in that experiment do not reflect their frequencies in the bone marrow. While our model of M. avium–induced trained immunity bears much in common with other models of trained immunity such as those using BCG vaccination or β-glucan (19, 20, 78), differences from model to model undoubtedly exist. While we focused on inflammatory pathways and effectors that are commonly induced in multiple murine models and human studies, more work is necessary to validate these findings in humans (19, 20, 78).
Overall, we provide one of the first studies to our knowledge to determine whether chromatin modifications induced in HSPCs are conserved through stages of myeloid differentiation. We identify 2 key gene targets that demonstrate epigenetic changes in common from the progenitor stage to fully differentiated Macs that may contribute directly to CTI.
-
Methods
Sex as a biological variable
Our previous work in M. avium infection found no significant differences in the mouse pathological responses to M. avium. Therefore, we used both sexes of mice for these experiments.
Mice
For the trained immunity and sequencing experiments, both male and female C57BL/6 (CD45.2) and C57BL/6.SJL (CD45.1) mice were used, with equal number of males and females used per group per experiment. CD45.2 mice from ages 8–16 weeks were used for infection experiments, whereas CD45.1 from 6–9 weeks were used for initial competitive bone marrow transplants.
THP-1 cell culture
THP-1 cells (ATCC TIB-2020) Cells were cultured in RPMI-1640 (Gibco, 11875093) supplemented with 10% FBS (Gibco, 10437-028) and 1% Penicillin (10,000 units/mL) and streptomycin (10,000 μg/mL) (Invitrogen, 15140122). THP-1 cells were passaged up to 20 times before stock was refreshed.
Cell electroporation and CRISPR/Cas9
CRISPR/Cas9 electroporation was completed with the ThermoFisher neon transfection system. Cells were collected and washed 3 times with PBS and counted for viability. Scramble control was purchased from Synthego. GBP6 sgRNAs were generated through NCBI browser, while IRF1 sgRNAs were based on previous reported sgRNAs (85). Specific 5′–3′ sequences are listed in Supplemental Table 26. Generated single guide RNAs (sgRNAs) were used at 1 μg/μL, pooled together per group types and heated to 95°C for 2 minutes to open sgRNA complexes. Afterward, samples were moved onto ice for 2 minutes, followed by the addition Cas9 (Synthego 1 μg/μL) and incubated for 15 minutes at room temperature. Concurrently, 500,000 THP-1 cells were collected per sample, spun down, and resuspended in T buffer from the Neon transfection kit (ThermoFisher, MPK1025). Cells were then added to the Cas9/sgRNA mixture post-incubation, and electroporation was carried out following the Neon transfection system protocol. Electroporation conditions were 1600V with 10 ms spaces and 3 pulses.
M. avium microbial infections
Mice were infected i.v. via retro-orbital injection with M. avium (2 × 106 colony forming units [CFU]) as previously described. M. avium CFUs were quantified plating homogenates onto 7H10 Middlebrook agar plates (BD 262710) supplemented with OADC enrichment supplement (BD 212351).
Bone marrow transplant and trained immunity challenge experiments
For competitive transplants, bone marrow from CD45.2 mice was isolated from harvested tibias, femurs, and pelvic bones and pooled per group. Cells are RBC lysed (BioLegend 420301) and cKit enriched via magnetic bead enrichment (Miltenyi 130-097-146). Cells were then sorted for lineage– cells on a Sony Sorter SH800 with a 100 μM microchip. Live 20,000 CD45.2 Lin– cKit enriched cells and 200,000 CD45.1 rescue marrow were transplanted into CD45.1 mice that had been lethally irradiated CD45.1 recipients (split dose 10 Gy from a Cesium Irradiator). CD45.2 donor cell engraftment is checked with flow cytometry on recipients’ peripheral blood every 4 weeks on a BD Fortessa.
For trained immunity challenge experiments, recipients were infected with 2.0 × 106 CFU M. avium. After 4 weeks, spleens were harvested and plated as described below.
M. avium bacterial plating
Spleens harvested from transplant recipients are weighed and checked for size. Half spleens were then weighed and crushed in media (HBSS, 2% FBS, 1% HEPES) and filtered into tubes at 10 mL resuspension. Volumes were measured by mass, with the assumption of density being 1 g/mL, and crushed spleen samples were serially diluted to 1 × 10–3 to 1 × 106. In total 100 μL of serially diluted homogenate is plated in duplicate per dilution onto 7H10 agar plates supplemented with OADC enrichment (BD 262710, BD 212351). After plates are incubated for one week, plates are colony counted for bacterial colony forming unit (CFU) calculations.
BMDM generation
Lin–, cKit-enriched hematopoietic cells were isolated from whole bone marrow via flow activated cell sorting (FACS). Cells were then plated and expanded in DMEM/F12/Glutamax (Gibco, 10565042) supplemented with 1% penicillin (10,000 units/mL) and streptomycin (10,000 μg/mL) (Invitrogen, 15140122), 10% FBS (Gibco, 10437-028), and 1 mL 1M HEPES (Gibco, 15630080). Cells were treated with both recombinant murine IL-3 (rm-IL3, Peprotech, 50ng/mL) and recombinant murine Mac CSF (rm-M-CSF, Peprotech, 50 ng/mL) to promote myeloid progenitor expansion and Mac expansion. Cells were maintained in a CO2 incubator at 37°C. Cells were transferred to a larger container to eventually be adherent in T75 flasks. Fresh media was added to the existing media per transfer to avoid overstressing cells in culture. Fresh cytokines were added at the same concentrations after each expansion, which happened every other day. After 1 week, cells were treated with only rm-M-CSF.
Seahorse assay
Mature BMDMs were harvested and spun down at 1,200 rpm for 5 minutes at 4°C and resuspended in DMEM culture media for cell counting (Cellometer Auto 2000, Nexcelom, Lawrence, MA). BMDMs were then seeded in 8 well miniplates (Agilent, 103022-100) at a concentration of 7 × 104 cells/well in 80 μL DMEM media and placed in a CO2 incubator at 37°C overnight to allow adherence. One day before the seahorse assay. Sensor cartridges (Agilent, 103022-100) were hydrated with 200 μL ultrapure water and placed in a non-CO2 incubator at 37°C overnight. On the day of the procedures, DMEM was removed from cell plates and washed with 200 μL basal media (Agilent Technologies, 103565-100) supplemented with 25 mM glucose (Agilent, 103578-100), 2 mM Na-Pyruvate (Agilent, 103577-100), and 2 mM L-glutamine (Agilent, 103579-100) for 3 times and replaced by supplemented basal media at a final volume of 175 μL. Sensor cartridge hydrated buffer was replaced by 200 μL Agilent calibration buffer. Various drugs from the Seahorse mitostress test kit (Agilent, 103010-100) were then dissolved in supplemented basal media, with final working concentrations being 1.5 μM Oligomycin, 0.75 μM FCCP, and 0.5 μM AA/Rotenone. Mitochondrial function was assessed on a Seahorse XFp; 8 well analyzer in real time following the injection of oligomycin, FCCP, and AA/Rotenone following the manufacturer’s instructions. Specific mitochondrial parameters were calculated according to the manufacturer’s protocol.
Cytokine analysis
BMDMs were counted and seeded at 200,000 cells per well media (DMEM/F12/Glutamax, 10% FBS, HEPES) for 24 hours before the challenge. M. avium at a multiplicity of infection of 1 (MOI 1) was used to challenge the cells, and supernatant containing cytokines was collected 4 hours after challenge. The supernatant was sent to Eve Technologies (Calgary, AB, Canada) for targeted cytokine analysis with a Luminex 200 platform 10-Plex Discovery Assay via ELISA.
CUT&RUN-Seq
CUT&RUN-Seq protocol was performed with based on published work from the lab and others (25, 86). Bone marrow from M. avium–infected or naive mice was isolated from tibias, femurs, and pelvic bones and pooled per group. After RBC lysis, cells were separated by cKit-enrichment and stained for specific cell surface markers. From cKit+ fractions, 30,000 HSCs (Lin–, cKit-enriched, CD150+, CD48–), 200,000 MPP3s (Lin–, cKit-enriched, CD150–, CD48+, Flk2–, CD34+), and 200,000 GMPs (Lin–, cKit-enriched, CD34+, CD16/32+) were sorted per sample. In total, 200,000 Macs (CD3–, B220–, NK1.1–, CD11b+, Ly6g–, Ly6c+) were sorted per sample from the cKit– fractions. Samples were counted and washed with wash buffer — 50 mL H2O with 20 mM HEPES (Gibco, 15630080), 150 mM NaCl, 0.5 mM Spermidine, and 1 Roche Complete Protease inhibitor EDTA-free tablet (Sigma, 04693132001) — 3 times. Cells were then incubated with activated Concanavalin A beads (Bang Laboratories L2007231C) for 15 minutes at room temperature. Sample slurries were then washed and resuspended in wash buffer supplemented with 0.5% Digitonin (Dig-wash buffer). Dig-wash buffer was used for the remainder of the experimental method, with samples and buffers were kept on ice or at 4°C. Samples were incubated with specific antibodies overnight at 4°C on a rotator. Antibodies used were targeted for Rabbit anti-H3K27ac (Cell Signaling Technologies 8173, 1:50 concentration), Rabbit anti-H3K427me3 (Cell Signaling Technologies 9733, 1:50 concentration), and Rabbit anti-mouse IgG (Jackson ImmunoResearch 315-005-003, 1:50 concentration). After incubation, samples were washed 3 times and incubated with pAG-MNase (Epicypher SKU:15-016) on a rotator for 1 hour at 4°C. Samples were washed 3 times and then resuspended in 150 μL Dig-wash buffer. 2 μL of 100 mM CaCl2 was added to catalyze MNase activity and cutting, and reaction was kept at 4°C for 40 minutes. The reaction was quenched with 2X stopping buffer (5 mL total, H2O, NaCl 340 mM, EDTA 20 mM, EGTA, 4 mM, Digitonin 0.05%, RNase A 100 μg/mL, Glycogen 50 μg/mL, and Spike-in DNA 1 ng/mL; EpiCypher, 18-1401), and samples were heated to 37°C for 20 minutes for MNase/Histone/DNA complex release. Samples were placed on magnetic stands to separate the DNA supernatant from the bead slurry, and DNA purification was completed via phenol chloroform and ethanol precipitation extraction.
For CUT&RUN qPCR, DNA was detected by preamplifation using Invitrogen Platinum SuperFi II polymerase followed by qPCR using SybrGreen Supermix and GBP6-specific primers on a Quantstudio System. Data were normalized using a GAPDH control. Primers were: GBP6 forward primer (5′ TCGAGAGTTCCATCTTGCAGGC 3′), GBP6 reverse primer (5′ AGCAGCTCTTGCTCCTTCTCTG 3′), GAPDH forward primer (5′ CAGGAGAGTGTTTCCTCGTCC 3′), GAPDH reverse primer (5′ TTCCCATTCTCGGCCTTGAC 3′).
For CUT&RUN-Seq, libraries were generated with the services of AdmeraHealth. Quantified DNA by Qubit 2.0 HS DNA (ThermoFisher, Q32851) was assessed by Tapestation High on a D1000 High sensitivity DNA Assay (Agilent Technologies). Library preparation was completed using the KAPA HyperPrep kit (Roche) per the manufacturer’s instructions. Library quality was assessed with Qubit 2.0 (ThermoFisher), Tapestation (Agilent Technologies), and QuantStudio 5 System (Applied Biosystems). Illumina 8 nt dual indices were used. Libraries were pooled equimolarly based on quality and sequenced on an Illumina NovaSeq (Illumina) with a 150 paired-end (PE) read length.
Samples were prepared in biological duplicates, while one negative IgG control was generated per cell type and cell condition.
scATAC-Seq library generation and sequencing
CD45.2 mice were infected with 2 × 106 CFU M. avium for 4 weeks prior to bone marrow isolation. Isolated bone marrow from M. avium–infected and naive mice were collected, pooled by sample group, RBC-lysed, and enriched for cKit+ cells. Specific HSPCs, including LT-HSCs (Lin–, cKit+, CD48–, CD150+, Flk2–, CD34–), CD41+ HSCs (Lin–, cKit+, CD150+, CD41+), MPP3s (Lin–, cKit+, CD48+, CD150–, Flk2–, CD34+), and GMPs (Lin–, cKit+, CD41–, CD16/32+), and lineage cells, including Macs (B220–, CD3–, CD11c–,Gr1+, Mac-1+, Ly6g–, Ly6c+), neutrophils (B220–,CD3–, CD11c–, Gr1+, Ly6g+, Ly6c+), and B cells (B220+, CD3–, Gr1–, Mac-1+), were stained and sorted for with FACS on BD Aria IIs. Cells were counted and combined in equal ratios per group based on live/dead counts after sort at roughly 50,000 cells per group. Nuclei were isolated and loaded onto the Chromium controller system to generate 10X scATAC-Seq libraries. Library preparation and generation was handled by the Baylor Single Cell and Genetics core, and quality control of libraries was completed using a Bioanalyzer and Nanodrop system. Nuclei were isolated prior to analysis on the 10x genomics platform.
Statistical analysis and bioinformatics
Statistics. All statistical methods and handling of data was performed in GraphPad Prism v10.0. Mean with standard deviations for all data figures was used in figures. Individual data points per experiment were assessed for outliers using the ROUT method (Q = 5%). Collective data was then checked for Gaussian Distribution with the Shapiro-Wilk test. Two-tailed Student’s t test, ordinary 1-way ANOVA with Tukey’s multiple comparison, or 2-way ANOVA were used per experiment. Experiments were run at least twice unless stated, and the number of replicates and further statistical details are provided in the figure legends.
CUT&RUN-Seq. FASTQ files were aligned to murine genome mm10 using the BWA aligner in Illumina Basespace, to which files are aligned with bwa and converted into aligned BAM files via samtools. BAM files were used for downstream analysis. MACS2 (v.2.2.7.1) (87) was used for peakcalling using the –broad –q 0.01 functions in addition to standard parameters, with test samples compared with background IgG controls. Bedgraph files produced by MACS2 were used for genomic track visualization using EaSeq (88) and confirmed with integrative genomic viewer (IGV).
Significant peaks were then identified and used for differential analysis using the Diffbind R package (v3.14) (89). Samples were filtered based gene blacklists and IgG greylists, and count matrixes were then generated and normalized by default Diffbind parameters (library size). Differential analysis was performed with cutoffs log2FC > 1 and FDR cutoff < 0.05. Chromosomal regions with significant peaks were then filtered based on fold changes that were positive and negative, to which cis-associated gene identification and gene ontogeny were performed GREAT (v4.04) (90) and rGREAT(v2.6) (91) Gene ontogeny were considered significant if the calculated BinomialFDR < 0.05.
Normalized count matrixes produced by Diffbind were extracted and used to produce clustering heatmap plots based on z scores using DESeq2 (v1.44.0) (92).
scATAC-Seq. Quality control was performed prior to statistical analyses to remove low quality cells. Retained cells required between 500 and 30,000 peak region fragments, at least 25% reads in peaks, a ratio of genomic reads in blacklist regions less than 0.1, a ratio of mononucleosomal to nucleosome-free fragments less than 2.5, and a TSS enrichment score greater than 1.5. The scATAC-Seq datasets from M. avium–infected and naive mice were then integrated to maintain a consistent set of peaks across the 2 datasets. A common peak set was defined using the GRanges reduce function and quantified for each dataset using the FeatureMatrix Seurat function. The datasets were then embedded into a shared low-dimensional space using a reciprocal Latent Semantic Indexing projection.
Cell type labels were predicted by mapping the gene activity computed by the Signac R package to previously published scRNA-Seq data. The cell type annotation for the scRNA-Seq dataset used the Haemopedia Mouse RNA-Seq Atlas (v1.5) (93) and Immgen RNA-Seq profiles (v1.2) (94) as reference datasets to predict cell type labels using the MapQuery Seurat function. To refine the peaks analyzed in the dataset, we performed peak calling using MACS2 via the CallPeaks Signac function with cell type as a grouping variable. These new peaks were combined into a common peak set and quantified across all cells.
Differential accessibility (DA) analysis comparing chromatin openness in M. avium–infected mice relative to naive mice was performed for each cell type individually, as well as overall. Both positive and negative markers were found using the FindMarkers Seurat function utilizing the logistic regression framework with the total number of fragments as a latent variable. Peaks that were present in at least 5% of the cells in either population were included in the DA analysis. Motif enrichment analysis with background peaks matched for overall GC content was performed to identify DNA motifs overrepresented in peaks that were significantly more accessible (P < 0.005) in M. avium–infected mice and naive mice.
All analyses were performed in R (v4.4.1) using the Seurat (v5.1.0) (95) and Signac (v1.14.0) (96) R packages.
Study approval
The handling and maintenance of mice complied with the protocols and guidelines approved by the IACUC at Baylor College of Medicine.
Data availability
All biological materials, reagents, sgRNAs, kits, and bioinformatic tools utilized for these experiments are listed in Supplemental Table 26. Epigenetic datasets are available in GEO, and accession nos. are provided in Supplemental Table 26. Values for all data points in graphs are reported in the Supporting Data Values file.
-
Author contributions
BTT and KYK designed the study, conducted the experiments, and wrote the paper. PNL performed bioinformatic analyses, provided the relevant interpretations, and wrote the paper. RC and SK performed the BMDM functional experiments and analysis. DTL contributed to the CUT&RUN-Seq experiments. BNK designed the scATAC-Seq experiments. AT designed optimized flow cytometry panels for FACS and performed data analysis. LM and AC provided assistance with Seahorse Assays. Coauthor order was agreed by the 2 coauthors prior to manuscript writing.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- Baylor College of Medicine Single Cell Genomics
- NIH 1S10OD018033
- Cytometry and Cell Sorting Core at Baylor College of Medicine
- CPRIT Core Facility Support Award (CPRIT-RP180672)
- NIH (P30 CA125123 and S10 RR024574)
- Cancer Prevention and Research Institute of Texas (CPRIT-RR140038; to AC)
- American Society of Hematology (Bridge grant 2023, round 20)
- Cell and Gene Therapy Training Grant T32HL2332 (BTT)
- F31HL164287 (BTT)
- Hematology Training Grant 5T32DK06445 (AT)
- F30HL172468 (AT)
- P01CA265748 (KYK)
- R35 HL155672 (KYK).
-
Acknowledgments
The authors would like to thank members of the King Lab for useful discussions. We thank Marcus A. Florez, Effa Longe, Daniel Morales-Mantilla, Hannah Yan, Daniel E. Kennedy II, Arushana Maknojia, Mara Willis, and Spencer Redding for technical assistance with the work. We thank Joel Sederstrom and the Flow Cytometry and Cell Sorting Core at Baylor College of Medicine for their assistance. We would also like to thank Ajibike Lapite for help with editing. Finally, we thank Chad Shaw for discussion and conceptualization of this project. Figures were generated in BioRender.
Address correspondence to: Katherine Y. King, Department of Pediatrics, Division of Infectious Diseases, 1102 Bates Street Suite 1150, Houston, Texas, 77030 USA. Phone: 832.824.4330; Email: kyk@bcm.edu.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2026, Tran et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(2):e193686.https://doi.org/10.1172/jci.insight.193686.
-
References
- Netea MG, et al. Trained immunity: a program of innate immune memory in health and disease. Science. 2016;352(6284):aaf1098.
- Ochando J, et al. Trained immunity - basic concepts and contributions to immunopathology. Nat Rev Nephrol. 2023;19(1):23–37.
- Saeed S, et al. Epigenetic programming of monocyte-to-macrophage differentiation and trained innate immunity. Science. 2014;345(6204):1251086.
- Cheng SC, et al. MTOR- and HIF-1α-mediated aerobic glycolysis as metabolic basis for trained immunity. Science. 2014;345(6204):1250684.
- Arts RJW, et al. Immunometabolic pathways in BCG-induced trained immunity. Cell Rep. 2016;17(10):2562–2571.
- Quintin J, et al. Candida albicans infection affords protection against reinfection via functional reprogramming of monocytes. Cell Host Microbe. 2012;12(2):223–232.
- Kleinnijenhuis J, et al. Bacille Calmette-Guérin induces NOD2-dependent nonspecific protection from reinfection via epigenetic reprogramming of monocytes. Proc Natl Acad Sci U S A. 2012;109(43):17537–17542.
- Arts RJW, et al. BCG vaccination protects against experimental viral infection in humans through the induction of cytokines associated with trained immunity. Cell Host Microbe. 2018;23(1):89–100.
- Moorlag SJCFM, et al. β-glucan induces protective trained immunity against mycobacterium tuberculosis infection: a key role for IL-1. Cell Rep. 2020;31(7):107634.
- Benn CS, et al. A small jab - a big effect: nonspecific immunomodulation by vaccines. Trends Immunol. 2013;34(9):431–439.
- Kleinnijenhuis J, et al. Long-lasting effects of BCG vaccination on both heterologous Th1/Th17 responses and innate trained immunity. J Innate Immun. 2014;6(2):152–158.
- King KY, Goodell MA. Inflammatory modulation of HSCs: viewing the HSC as a foundation for the immune response. Nat Rev Immunol. 2011;11(10):685–692.
- Takizawa H, et al. Demand-adapted regulation of early hematopoiesis in infection and inflammation. Blood. 2012;119(13):2991–3002.
- Ding Y, et al. Transcriptional and epigenetic control of hematopoietic stem cell fate decisions in vertebrates. Dev Biol. 2021;475:156–164.
- Pietras EM. Blood spotlight inflammation: a key regulator of hematopoietic stem cell fate in health and disease. Blood. 2017;130(15):1693–1698.
- Cao R, et al. Forged in the fire: Lasting impacts of inflammation on hematopoietic progenitors. Exp Hematol. 2024;134:104215.
- Hormaechea-Agulla D, et al. Chronic infection drives Dnmt3a-loss-of-function clonal hematopoiesis via IFNγ signaling. Cell Stem Cell. 2021;28(8):1428–1442.
- de Laval B, et al. C/EBPβ-dependent epigenetic memory induces trained immunity in hematopoietic stem cells. Cell Stem Cell. 2020;26(5):657–674.
- Sun SJ, et al. BCG vaccination alters the epigenetic landscape of progenitor cells in human bone marrow to influence innate immune responses. Immunity. 2024;57(9):2095–2107.
- Cirovic B, et al. BCG vaccination in humans elicits trained immunity via the hematopoietic progenitor compartment. Cell Host Microbe. 2020;28(2):322–334.
- Kaufmann E, et al. BCG educates hematopoietic stem cells to generate protective innate immunity against tuberculosis. Cell. 2018;172(1-2):176–190.
- Kain BN, et al. Hematopoietic stem and progenitor cells confer cross-protective trained immunity in mouse models. iScience. 2023;26(9):107596.
- Mitroulis I, et al. Modulation of myelopoiesis progenitors is an integral component of trained immunity. Cell. 2018;172(1-2):147–161.
- Khan N, et al. M. tuberculosis reprograms hematopoietic stem cells to limit myelopoiesis and impair trained immunity. Cell. 2020;183(3):752–770.
- Le DT, et al. BATF2 promotes HSC myeloid differentiation by amplifying IFN response mediators during chronic infection. iScience. 2023;26(2):106059.
- Cheong JG, et al. Epigenetic memory of coronavirus infection in innate immune cells and their progenitors. Cell. 2023;186(18):3882–3902.
- Tian C, et al. Impaired histone inheritance promotes tumor progression. Nat Commun. 2023;14(1):3429.
- Flury V, Groth A. Safeguarding the epigenome through the cell cycle: a multitasking game. Curr Opin Genet Dev. 2024;85:102161.
- Reverón-Gómez N, et al. Accurate recycling of parental histones reproduces the histone modification landscape during DNA replication. Mol Cell. 2018;72(2):239–249.
- Escobar TM, et al. Active and repressed chromatin domains exhibit distinct nucleosome segregation during DNA replication. Cell. 2019;179(4):953–963.
- Ludin A, et al. Reactive oxygen species regulate hematopoietic stem cell self-renewal, migration and development, as well as their bone marrow microenvironment. Antioxid Redox Signal. 2014;21(11):1605–1619.
- Brookes PS. Mitochondrial H(+) leak and ROS generation: an odd couple. Free Radic Biol Med. 2005;38(1):12–23.
- Gogoleva VS, et al. Current perspectives on the role of TNF in hematopoiesis using mice with humanization of TNF/LT system. Front Immunol. 2021;12:661900.
- Tie R, et al. Interleukin-6 signaling regulates hematopoietic stem cell emergence. Exp Mol Med. 2019;51(10):1–12.
- Netea MG, et al. The role of trained immunity in COVID-19: Lessons for the next pandemic. Cell Host Microbe. 2023;31(6):890–901.
- Hernández-Malmierca P, et al. Antigen presentation safeguards the integrity of the hematopoietic stem cell pool. Cell Stem Cell. 2022;29(5):760–775.
- Song R, et al. IRF1 governs the differential interferon-stimulated gene responses in human monocytes and macrophages by regulating chromatin accessibility. Cell Rep. 2021;34(12):108891.
- Singh RK, et al. Immunoregulatory cytokines in bone marrow and peripheral blood stem cell products. Bone Marrow Transplant. 1999;23(1):53–62.
- Yan H, et al. The bacterial microbiota regulates normal hematopoiesis via metabolite-induced type 1 interferon signaling. Blood Adv. 2022;6(6):1754–1765.
- Baldridge MT, et al. Quiescent haematopoietic stem cells are activated by IFN-gamma in response to chronic infection. Nature. 2010;465(7299):793–797.
- Tretina K, et al. Interferon-induced guanylate-binding proteins: guardians of host defense in health and disease. J Exp Med. 2019;216(3):482–500.
- Kim B-H, et al. A family of IFN-γ-inducible 65-kD GTPases protects against bacterial infection. Science. 2011;332(6030):717–721.
- Nakamura MC. CIITA: a master regulator of adaptive immunity shows its innate side in the bone. J Bone Miner Res. 2014;29(2):287–289.
- Aparicio-Siegmund S, Garbers C. The biology of interleukin-27 reveals unique pro- and anti-inflammatory functions in immunity. Cytokine Growth Factor Rev. 2015;26(5):579–586.
- Silva MJA, et al. The association between CCL5/RANTES SNPs and susceptibility to HIV-1 infection: a meta-analysis. Viruses. 2023;15(9):1958.
- Wang Z, et al. Interplay between cofactors and transcription factors in hematopoiesis and hematological malignancies. Signal Transduct Target Ther. 2021;6(1):24.
- Yu X, Buck MJ. Pioneer factors and their in vitro identification methods. Mol Genet Genomics. 2020;295(4):825–835.
- Huang Q, et al. Mechanistic insights into the interaction between transcription factors and epigenetic modifications and the contribution to the development of obesity. Front Endocrinol (Lausanne). 2018;9:370.
- Rice G, et al. Sox9 marks limbal stem cells and is required for asymmetric cell fate switch in the corneal epithelium [preprint]. https://doi.org/10.1101/2024.04.08.588195 Posted on bioRxiv May 2, 2024.
- Gonzales KAU, et al. Stem cells expand potency and alter tissue fitness by accumulating diverse epigenetic memories. Science. 2021;374(6571):eabh2444.
- Matatall KA, et al. Type II interferon promotes differentiation of myeloid-biased hematopoietic stem cells. Stem Cells. 2014;32(11):3023–3030.
- Matatall KA, et al. Chronic infection depletes hematopoietic stem cells through stress-induced terminal differentiation. Cell Rep. 2016;17(10):2584–2595.
- Kalvakolanu DV. Alternate interferon signaling pathways. Pharmacol Ther. 2003;100(1):1–29.
- Li H, et al. The Interferon signaling network and transcription factor C/EBP-β. Cell Mol Immunol. 2007;4(6):407–418.View this article via: PubMed Google Scholar
- Méndez-Samperio P, et al. Mycobacterium bovis Bacillus Calmette-Guérin (BCG)-induced activation of PI3K/Akt and NF-kB signaling pathways regulates expression of CXCL10 in epithelial cells. Cell Immunol. 2009;256(1-2):12–18.
- Chavez C, et al. IRF1 cooperates with ISGF3 or GAF to form innate immune de novo enhancers in macrophages. Sci Signal. 2025;18(868):eado8860.
- Starr T, et al. The phorbol 12-myristate-13-acetate differentiation protocol is critical to the interaction of THP-1 macrophages with Salmonella Typhimurium. PLoS One. 2018;13(3):e0193601.
- Chen S, et al. Mitochondrial control of innate immune responses. Front Immunol. 2023;14:1166214.
- Kaufmann E, et al. BCG vaccination provides protection against IAV but not SARS-CoV-2. Cell Rep. 2022;38(10):110502.
- Li Y, et al. Roles of trained immunity in the pathogenesis of periodontitis. J Periodontal Res. 2023;58(5):864–873.
- Christ A, et al. Western diet triggers NLRP3-dependent innate immune reprogramming. Cell. 2018;172(1-2):162–175.
- Ochando J, et al. Trained immunity in organ transplantation. Am J Transplant. 2020;20(1):10–18.
- Mills TS, et al. A distinct metabolic and epigenetic state drives trained immunity in HSC-derived macrophages from autoimmune mice. Cell Stem Cell. 2024;31(11):1630–1649.
- Bottero V, et al. NF-kappaB and the regulation of hematopoiesis. Cell Death Differ. 2006;13(5):785–797.
- Smith JNP, et al. Hematopoietic stem cell regulation by Type I and II interferons in the pathogenesis of acquired aplastic anemia. Front Immunol. 2016;7:330.
- Lama C, et al. Type 1 interferon perturbates clonal competition by reshaping human blood development [preprint]. https://doi.org/10.1101/2022.09.28.509751 Posted on bioRxiv August 27, 2024.
- Ziogas A, et al. Long-term histone lactylation connects metabolic and epigenetic rewiring in innate immune memory. Cell. 2025;188(11):2992–3012.
- Barth TK, Imhof A. Fast signals and slow marks: the dynamics of histone modifications. Trends Biochem Sci. 2010;35(11):618–626.
- Yang Y, et al. The pioneer factor SOX9 competes for epigenetic factors to switch stem cell fates. Nat Cell Biol. 2023;25(8):1185–1195.
- Kamada R, et al. Interferon stimulation creates chromatin marks and establishes transcriptional memory. Proc Natl Acad Sci U S A. 2018;115(39):E9162–E9171.
- Kalafati L, et al. Innate immune training of granulopoiesis promotes anti-tumor activity. Cell. 2020;183(3):771–785.
- Nilsson AJSR, et al. IRF1 regulates self-renewal and stress responsiveness to support hematopoietic stem cell maintenance. 2023;9(43):eadg5391.
- Nathan CF, et al. Identification of interferon-gamma as the lymphokine that activates human macrophage oxidative metabolism and antimicrobial activity. J Exp Med. 1983;158(3):670–689.
- Rosain J, et al. Human IRF1 governs macrophagic IFN-γ immunity to mycobacteria. Cell. 2023;186(3):621–645.
- Chu Y biao, et al. Irf1- and Egr1-activated transcription plays a key role in macrophage polarization: A multiomics sequencing study with partial validation. Int Immunopharmacol. 2021;99:108072.
- Langlais D, et al. The macrophage IRF8/IRF1 regulome is required for protection against infections and is associated with chronic inflammation. J Exp Med. 2016;213(4):585–603.
- Wang L, et al. The multiple roles of interferon regulatory factor family in health and disease. Signal Transduct Target Ther. 2024;9(1):282.
- Moorlag SJCFM, et al. Multi-omics analysis of innate and adaptive responses to BCG vaccination reveals epigenetic cell states that predict trained immunity. Immunity. 2024;57(1):171–187.
- Shi T, et al. Role of GBP1 in innate immunity and potential as a tuberculosis biomarker. Sci Rep. 2022;12(1):11097.
- Praefcke GJK. Regulation of innate immune functions by guanylate-binding proteins. Int J Med Microbiol. 2018;308(1):237–245.
- Schelle L, et al. Evolutionary and functional characterization of lagomorph guanylate-binding proteins: a story of gain and loss and shedding light on expression, localization and innate immunity-related functions. Front Immunol. 2024;15:1303089.
- Siwek W, et al. Activation of clustered IFNγ target genes drives cohesin-controlled transcriptional memory. Mol Cell. 2020;80(3):396–409.
- Mikulski P, et al. Heritable maintenance of chromatin modifications confers transcriptional memory of interferon-γ signaling. Nat Struct Mol Bio. 2025;32(7):1255–1267.
- Riksen NP, Netea MG. Immunometabolic control of trained immunity. Mol Aspects Med. 2021;77:100897.
- Benaoudia S, et al. A genome-wide screen identifies IRF2 as a key regulator of caspase-4 in human cells. EMBO Rep. 2019;20(9):e48235.
- Skene PJ, Henikoff S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. Elife. 2017;6:e21856.
- Zhang Y, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9(9):R137.
- Lerdrup M, et al. An interactive environment for agile analysis and visualization of ChIP-Seq data. Nat Struct Mol Biol. 2016;23(4):349–357.
- Ross-Innes CS, et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature. 2012;481(7381):389–393.
- McLean CY, et al. GREAT improves functional interpretation of cis-regulatory regions. Nat Biotechnol. 2010;28(5):495–501.
- Gu Z, Hübschmann D. rGREAT: an R/bioconductor package for functional enrichment on genomic regions. Bioinformatics. 2023;39(1):btac745.
- Love MI, et al. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 2014;15(12):550.
- Choi J, et al. Haemopedia RNA-Seq: a database of gene expression during haematopoiesis in mice and humans. Nucleic Acids Res. 2019;47(d1):D780–D785.
- Yoshida H, et al. The cis-regulatory atlas of the mouse immune system. Cell. 2019;176(4):897–912.
- Hao Y, et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat Biotechnol. 2024;42(2):293–304.
- Stuart T, et al. Single-cell chromatin state analysis with Signac. Nat Methods. 2021;18(11):1333–1341.
-
Version history
- Version 1 (December 9, 2025): In-Press Preview
- Version 2 (January 23, 2026): Electronic publication












