Research ArticleGastroenterology
Open Access |  10.1172/jci.insight.192953
10.1172/jci.insight.192953
Characteristics of anti–integrin αvβ6 autoantibodies in patients with ulcerative colitis
Ikuhisa Takimoto,1 Masahiro Shiokawa,1 Yoshihiro Nishikawa,1 Takeshi Kuwada,1 Sakiko Ota,1 Darryl Joy C. Juntila,1 Takafumi Yanaidani,1 Kenji Sawada,1 Ayako Hirata,1 Muneji Yasuda,1 Koki Chikugo,1 Risa Nakanishi,1 Masataka Yokode,1 Yuya Muramoto,1 Shimpei Matsumoto,1 Tomoaki Matsumori,1 Tsutomu Chiba,1,2 and Hiroshi Seno1
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Takimoto, I. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Shiokawa, M. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by
Nishikawa, Y.
in:
PubMed
|
Google Scholar
|

1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by
Kuwada, T.
in:
PubMed
|
Google Scholar
|

1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Ota, S. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Juntila, D. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Yanaidani, T. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Sawada, K. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Hirata, A. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Yasuda, M. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Chikugo, K. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Nakanishi, R. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Yokode, M. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by
Muramoto, Y.
in:
PubMed
|
Google Scholar
|

1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Matsumoto, S. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by
Matsumori, T.
in:
PubMed
|
Google Scholar
|

1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Chiba, T. in: PubMed | Google Scholar
1Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, Sakyo-ku, Kyoto, Japan.
2Kansai Electric Power Hospital, Fukushima-ku, Osaka, Japan.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
Authorship note: MS and YN contributed equally to this work.
Find articles by Seno, H. in: PubMed | Google Scholar
Authorship note: MS and YN contributed equally to this work.
Published January 8, 2026 - More info
JCI Insight. 2026;11(4):e192953. https://doi.org/10.1172/jci.insight.192953.
© 2026 Takimoto et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: March 3, 2025; Accepted: January 2, 2026
-
Abstract
Ulcerative colitis (UC) is a chronic inflammatory condition of the colon that primarily affects the mucosal layer. Previously, we identified autoantibodies against integrin αvβ6 in patients with UC. In this study, we established monoclonal antibodies (mAbs) from patients with UC to reveal the features and functions of these anti–integrin αvβ6 autoantibodies. We identified two shared heavy chain complementarity-determining region 3 (CDR3) amino acid sequences among different patients with UC. Notably, several mAbs contained the RGD sequence in their heavy chain CDR3 that mimicked the key recognition sequence of integrin αvβ6 ligands such as fibronectin. Almost all mAbs selectively reacted with integrin αvβ6 in the presence of divalent cations (Ca2+ and Mg2+) and blocked fibronectin–integrin αvβ6 binding. MAbs that shared the same heavy chain CDR3 amino acid sequence showed differences in reactivity to integrin αvβ6, indicating that the reactivity of these mAbs is also affected by the light chain. Some of the mAbs showed varying degrees of cross-reactivity with integrin αvβ3. The identification of shared CDR3 amino acid sequences in anti–integrin αvβ6 antibodies from several patients with UC suggests a common mechanism underlying their production, which may help elucidate the pathogenesis of UC.
-
Introduction
Ulcerative colitis (UC) is a chronic inflammatory bowel disease marked by continuous mucosal inflammation that begins in the rectum and spreads toward the proximal colon (1–3). The etiology of UC is not yet clearly understood; however, it is hypothesized that genetic predisposition and environmental factors drive immune dysregulation, leading to various grades of epithelial damage along the intestinal tract (1–3).
We previously reported that most patients with UC have autoantibodies targeting integrin αvβ6, and the antibody titer correlates with disease severity; furthermore, IgG from patients with UC inhibits binding between integrin αvβ6 and fibronectin (4). Subsequently, a nationwide multicenter study in Japan confirmed the diagnostic utility of anti–integrin αvβ6 antibodies for UC (5), and several groups have confirmed the presence of anti–integrin αvβ6 antibodies in patients with UC from various countries (6–8). Importantly, a study revealed that these antibodies can be detected up to a decade before a clinical diagnosis of UC, and that elevated antibody titers are linked to poor UC-related prognoses (6). However, the pathophysiological significance of anti–integrin αvβ6 antibodies in UC has not yet been fully elucidated.
Integrins are transmembrane receptors that facilitate interactions between cells and the extracellular matrix (9, 10). Integrin heterodimers are composed of α and β subunits, which are noncovalently linked (9). In mammals, the integrin family comprises 18 α subunits and 8 β subunits, forming a total of 24 distinct heterodimeric combinations (9). Among them, integrin αvβ6 is expressed exclusively on epithelial cells, where it functions as a receptor for extracellular matrix proteins, including fibronectin (10). Additionally, integrin αvβ6 binds to latency-associated protein (LAP) that is in complex with TGF-β, thereby facilitating the activation of TGF-β (10, 11). Thus, integrin αvβ6 plays a pivotal role in maintaining epithelial barrier integrity, protecting against pathogenic infections, and modulating inflammation through TGF-β signaling activation (10, 12–14).
Taking into consideration the correlation between anti–integrin αvβ6 antibody titers and both the severity and outcomes of UC, as well as the critical role of integrin αvβ6 in maintaining colon epithelial integrity, it is likely that this antibody has a substantial impact on UC pathogenesis. Recent studies have also highlighted the importance of B cells in the pathogenesis of UC (15–17). Therefore, we aimed to evaluate the pathophysiological function of anti–integrin αvβ6 antibodies in UC. In this study, we established anti–integrin αvβ6 monoclonal antibodies (mAbs) from the patients with UC and evaluated their characteristics.
-
Results
Anti–integrin αvβ6 antibodies from patients with UC share common CDR3 amino acid sequences. To investigate the properties of anti–integrin αvβ6 autoantibodies from patients with UC, we initially established 15 anti–integrin αvβ6 mAbs using peripheral blood mononuclear cells (PBMCs) or lymph node samples of 7 patients with UC with anti–integrin αvβ6 antibodies. The clinical characteristics of the patients are summarized in Supplemental Table 1 (supplemental material available online with this article; https://doi.org/10.1172/jci.insight.192953DS1). As shown in Figure 1, the 7 patients were given unique identification numbers, P1 to P7, and the obtained mAbs were given unique identification numbers (UC1-1 to UC7-1) along with serial numbers (nos. 1–15). To characterize each mAb, we analyzed complementarity-determining region 3 (CDR3) sequences, which are crucial for antigen specificity, along with variable region gene usage patterns (Figure 1) and the frequency of somatic hypermutations (SHMs) in the V gene region (Supplemental Table 2). SHM frequency was calculated by counting of the number of mutated sequences against the annotated germline sequences of V region using IgBLAST. Using enzyme-linked immunosorbent assay (ELISA), we confirmed that these mAbs reacted with integrin αvβ6 (Figure 2A).
 Figure 1
Figure 1Gene usage and CDR3 amino acid sequence of each mAb. The variable regions of the heavy and light chains of the antibodies were analyzed using IgBLAST. The V, D, and J gene usage, along with the amino acid sequences of the complementarity-determining region 3 (CDR3) for each mAb, is presented.
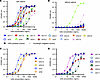 Figure 2
Figure 2Evaluation of reactivity of each mAb with integrin αvβ6. (A) The reactivity of each mAb to integrin αvβ6 was evaluated using ELISA in the presence of Mg2+ and Ca2+. All mAbs reacted with integrin αvβ6, albeit with different reactivity. (B) In ELISA in the absence of Mg2+ and Ca2+, all mAbs except UC4 lost reactivity to integrin αvβ6. (C and D) ELISA to estimate the reactivity against integrin αvβ6 of antibodies with a common heavy chain CDR3 amino acid sequence — UC1-1, UC2-3, UC2-4, UC3, and UC6 (carrying the CDR-H1 sequence) (C) and UC1-4, UC2-1, UC2-2, UC5-1, UC5-2, and UC7-1 (carrying the CDR-H2 sequence) (D).
Notably, we identified 2 distinct CDR3 amino acid sequences of the heavy chain that were shared among some of the mAbs obtained from the 7 patients with UC: one was AKVIPRIRGSGKAGIKDYYYGMDV (CDR-H1), encoded by IGHV3-30*18/IGHD3-10*01/IGHJ6*02 (shown in red in Figure 1); and the other was ARDRGFRGDTAMIKGGMDV (CDR-H2), encoded by IGHV1-18*01/IGHD5-18*01/IGHJ6*02 (shown in blue in Figure 1). MAbs no. 1 (UC1-1), no. 7 (UC2-3), no. 8 (UC2-4), no. 9 (UC3), and no. 13 (UC6) shared the first sequence, whereas mAbs no. 4 (UC1-4), no. 5 (UC2-1), no. 6 (UC2-2), no. 11 (UC5-1), no. 12 (UC5-2), and no. 14 (UC7-1) shared the second sequence. Notably, among the mAbs that shared the second sequence in the heavy chain CDR3, UC5-1 had a one-residue variation.
Interestingly, some mAbs had RGD sequences (shown in yellow in Figure 1) in the CDR3 region of the heavy chain. Integrin αvβ6 recognizes RGD sequences of its physiological ligands, including fibronectin and LAP (9). The presence of RGD sequences in the CDR3 regions of these mAbs suggests that they compete with the ligands (e.g., fibronectin and LAP) for binding to integrin αvβ6.
Analysis of CDR3 amino acid lengths and SHM frequencies of the V region gene (Supplemental Table 2) showed that these mAbs had longer CDR3 regions and lower SHM rates in the V region gene than do those reported in healthy individuals (7.47%) (18). Overall SHM frequencies were low; however, mutations were enriched in the CDRs, with higher rates than those in the framework regions (FR) (Supplemental Figure 1A). Amino acid mutations followed a similar distribution (Supplemental Figure 1B).
Most antibodies with CDR-H1 or CDR-H2 carried only 1–3 replacement mutations in the heavy chain CDRs and lacked silent mutations (Supplemental Figure 1C). In the FR regions, antibodies with CDR-H1 tended to show lower replacement/silent ratios, while those with CDR-H2 often showed few replacement mutations and no silent ones (Supplemental Figure 1C). These findings suggest that selective introduction of minimal but functional mutations may be sufficient for antigen binding, and that FR mutations may also contribute to binding in antibodies containing CDR-H2.
SHM levels in light chains varied widely. Some antibodies had heavily mutated CDRs, whereas others retained entirely germline light chains. Notably, several antibodies showed integrin αvβ6 reactivity without any light chain mutations, suggesting that such mutations are not always required for binding.
Taken together, these findings suggest that B cells using germline VH genes with intrinsic autoreactive potential acquire or enhance integrin αvβ6 binding through limited, functionally targeted replacement mutations.
All mAbs except UC4 react against integrin αvβ6 in a cation-dependent manner. Reactivity and affinity of each mAb to integrin αvβ6 were assessed using ELISA and biolayer interferometry (BLI), respectively. To validate results obtained in ELISA, we used the mouse anti–human integrin αvβ6 antibody 10D5, which is known to inhibit both fibronectin and LAP binding (19), as a positive control, and IgG purified from healthy individuals as negative controls. The reactivity of all the mAbs against integrin αvβ6 was confirmed using ELISA (Figure 2A), and the half-maximal effective concentration (EC50) values of the mAbs were calculated (6.30–1,537 ng/mL) (Table 1).
In our previous study, we showed that the serum titer of anti–integrin αvβ6 antibodies in patients with UC increased significantly with the addition of Ca2+ and Mg2+ (4). Therefore, we compared the binding of mAbs to integrin αvβ6 in the presence and absence of Ca2+ and Mg2+. The mAbs except UC4 showed marked loss of reactivity in ELISA in the absence of Ca2+ and Mg2+ (Figure 2B). Ca2+ and Mg2+ are important for integrin heterodimer formation, activation of integrins, and binding to its ligands (20–22). This cation-dependent binding suggests that these antibodies recognize integrin αvβ6 in its active conformation maintained by divalent cations. In other words, these mAbs were directed toward active integrin αvβ6. On the other hand, only UC4 reacted with integrin αvβ6 in a cation-independent manner, indicating that its binding site is different from that of other mAbs.
To investigate differences in reactivity with integrin αvβ6 caused by differences in light chains, we chose mAbs with a common CDR3 amino acid sequence in the heavy chain (Figure 2C: CDR-H1; and Figure 2D: CDR-H2) but with different light chains and analyzed their reactivities. Notably, UC2-3 and UC6 had identical CDR3 amino acid sequences in the light chain, and the light chain CDR3 of UC5-1 and UC7-1 had highly similar amino acid sequences. The other antibodies with common heavy chain CDR3 amino acid sequences had different light chain CDR3 amino acid sequences (UC1-1, UC2-4, and UC3; UC1-4, UC2-1, UC2-2, and UC5-2). UC2-3 and UC6 as well as UC5-1 and UC7-1 showed similar reactivities, consistent with sequence similarity; however, the mAbs with common heavy chain CDR3 amino acid sequences but different light chain amino acid sequences showed different reactivities in ELISA (Figure 2, C and D), suggesting that the light chain amino acid sequence is also important for determining binding specificity to integrin αvβ6.
Next, the affinity of each mAb to integrin αvβ6 was measured using BLI (Table 2 and Supplemental Figure 2A). First, we conducted BLI in the presence of cations in the buffer. Most mAbs had sufficient affinity for integrin αvβ6, similar to the results seen in ELISA; however, some of the mAbs that had shown high EC50 values (UC5-1, UC7-1, and UC7-2) in ELISA showed low affinity in BLI. In particular, UC4 did not bind sufficiently to integrin αvβ6 in BLI. To further characterize the binding properties of UC4, which showed unique cation-independent reactivity in ELISA, we performed BLI measurements using both UC4 and UC1-1 in the absence of cations (Supplemental Figure 2, B and C). We used UC1-1 as a representative cation-dependent antibody. In BLI measurements without cations, neither antibody bound to integrin αvβ6. Notably, UC4 showed no binding in BLI even in the presence of cations, despite showing reactivity in ELISA. The reason for this discrepancy remains to be determined.
 Table 2
Table 2Measurement of the dissociation constant, association rate constant, and dissociation rate constant for the association of each mAb with integrin αvβ6 using biolayer interferometry
Effect of mAbs on integrin αvβ6–fibronectin and integrin αvβ6–LAP binding. To investigate the function of each mAb, we examined blocking activities of the mAbs on integrin αvβ6–fibronectin and integrin αvβ6–LAP binding. Both fibronectin and LAP bind to the RGD binding site of integrin αvβ6 via their RGD motifs (9). In a solid-phase binding assay, 8 of the 15 mAbs firmly blocked integrin αvβ6–fibronectin binding in a dose-dependent manner (Figure 3A). The calculated half-maximal inhibitory concentration (IC50) values are shown in Table 3. However, some mAbs with high EC50 values (UC4, UC5-1, UC7-1, and UC7-2) showed very limited blocking activity. UC2-2, UC5-2, and UC6 did not have sufficient blocking activity to reach a plateau within the measured concentration range, and the IC50 could not be calculated. Notably, mAbs with common heavy chain CDR3 amino acid sequences showed varying degrees of blocking activity (Figure 3, B and C). Only marginal inhibition of integrin αvβ6–LAP binding was observed for the patient-derived mAbs (Figure 3D), in contrast to the clear blocking of integrin αvβ6–fibronectin binding. To better understand this differential blocking effect, we compared the affinities of fibronectin and LAP to integrin αvβ6 using BLI (Supplemental Figure 3, A and B, and Supplemental Table 3). Fibronectin showed measurable binding kinetics (dissociation constant [KD] = 47.18 nM), whereas LAP demonstrated extremely stable binding with negligible dissociation (dissociation rate constant [Koff] < 1.0 × 10–7 1/s); this indicated a substantially higher affinity to integrin αvβ6. The control antibody 10D5 demonstrated much higher affinity for integrin αvβ6 (Supplemental Figure 3C) and was able to effectively block LAP binding. These results suggest that the affinity of patient-derived mAbs for integrin αvβ6 is not sufficient to overcome the strong binding of LAP, thereby limiting their ability to inhibit integrin αvβ6–LAP interaction.
 Figure 3
Figure 3Blocking of integrin αvβ6–fibronectin binding or integrin αvβ6–LAP binding by each mAb in ELISA. (A) Blocking activity of each mAb on integrin αvβ6–fibronectin binding was evaluated using a solid-phase integrin αvβ6 binding assay. MAbs with low EC50 blocked integrin αvβ6–fibronectin binding in a concentration-dependent manner. MAbs with high EC50 concentrations (UC4, UC5-1, UC7-1, and UC7-2) showed little blocking activity. UC2-2, UC5-2, and UC6 did not have sufficient blocking activity to reach a plateau within the measured concentration range. (B and C) Blocking activity on integrin αvβ6–fibronectin binding was compared between antibodies with a common amino acid sequence in the heavy chain CDR3 — UC1-1, UC2-3, UC2-4, UC3, and UC6 (carrying the CDR-H1 sequence) (B) and UC1-4, UC2-1, UC2-2, UC5-1, UC5-2, and UC7-1 (carrying the CDR-H2 sequence) (C). (D) Blocking activity of each mAb against integrin αvβ6–LAP binding was evaluated using the same method. (E and F) Effect of RGDS peptides (E) and control RGES peptides (F) at various concentrations on the binding of mAbs to integrin αvβ6. LAP, latency-associated protein.
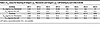 Table 3
Table 3IC50 values for blocking of integrin αvβ6–fibronectin and integrin αvβ6–LAP binding by each mAb in ELISA
Integrin αvβ6 binds to the RGD sequence in its ligands (9). Based on our results and those of a previous study (4), we hypothesized that the mAbs interact with the RGD binding site in integrin αvβ6. Therefore, we evaluated whether RGDS peptides could inhibit the binding of each of the studied mAbs with integrin αvβ6. In line with a previous report (4), RGDS peptides inhibited the binding of all mAbs, except that of UC4, to integrin αvβ6 in a dose-dependent manner (Figure 3E), whereas control RGES peptides did not show such effects (Figure 3F). The binding of UC4 with integrin αvβ6, which showed reactivity with integrin αvβ6 in the absence of cations in ELISA, was inhibited by neither RGDS nor RGES, indicating that this antibody reacts with integrin αvβ6 in an RGD-independent manner. The other mAbs were suggested to be critically dependent on the RGD binding site for binding to integrin αvβ6.
Cross-reactivity of anti–integrin αvβ6 antibodies with integrin αvβ3. Previously, we reported that sera of certain patients with UC contained antibodies against both integrin αvβ6 and αvβ3, with anti-αvβ3 antibody titers being consistently lower than those of anti-αvβ6 antibodies (4). However, it remained unclear whether anti–integrin αvβ3 antibodies existed independently or whether anti–integrin αvβ6 antibodies cross-reacted with integrin αvβ3. Therefore, using ELISA, we tested whether each of the mAbs used in this study reacts with integrin αvβ1, αvβ3, αvβ5, and αvβ8, all of which have αv chains and recognize RGD peptides in their physiological ligands (9) (Figure 4). UC4 showed almost identical reactivity to all integrins used for testing. Therefore, this antibody might react with either the αv chain or a common region of the β chain, such as the β tail domain. UC5-1 and UC5-2 showed reactivity not only to integrin αvβ6 but also to integrin αvβ3. Other mAbs reacted slightly to integrin αvβ3 at high concentrations and had no notable reaction to other integrins. These results suggested that anti–integrin αvβ6 antibodies potentially cross-react with integrin αvβ3.
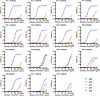 Figure 4
Figure 4Cross-reaction of mAbs to other integrins. Using ELISA, each mAb was tested to evaluate whether it reacted with integrin αvβ1, αvβ3, αvβ5, and αvβ8.
IgG and mAbs derived from patients with UC exhibit similar blocking activity on integrin αvβ6–fibronectin/LAP binding. We previously reported that IgG derived from patients with UC blocked integrin αvβ6–fibronectin binding (4); however, we had not tested whether it blocks integrin αvβ6–LAP binding. Using the IgG derived from 42 patients with UC who were part of the training group of our previous study and IgG from 8 control patients (4), we evaluated anti–integrin αvβ6 IgG titer of patients with UC, and then evaluated blocking activity of these IgG samples on integrin αvβ6–fibronectin and –LAP binding (Figure 5, A–C, and Supplemental Table 4).
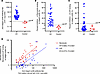 Figure 5
Figure 5Evaluation of anti–integrin αvβ6 antibodies using IgG from patients with UC. (A) Presence of anti–integrin αvβ6 antibodies in IgG derived from patients with UC. (B and C) Inhibitory effect of patient-derived IgG on integrin αvβ6–fibronectin binding (B) and integrin αvβ6–LAP binding (C). (D) Correlation of percentage inhibition of integrin αvβ6–fibronectin binding (B) and integrin αvβ6–LAP binding with antibody titers (fibronectin: R2 = 0.6463, P < 0.0001; LAP: R2 = 0.5775, P < 0.0001).
Consistent with our previous results (4), anti–integrin αvβ6 IgG titers were positive in 100% of cases (42 of 42) and showed a substantial inhibitory effect on integrin αvβ6–fibronectin binding in 76.2% of cases (32 of 42). Further, 23.8% cases (10 of 42) showed a substantial inhibitory effect on integrin αvβ6–LAP binding. Both inhibitory effects were correlated with antibody titers (Figure 5D). The patient-derived IgG samples showed blocking activity similar to that of the previously evaluated mAbs, based on their ability to inhibit integrin αvβ6–fibronectin and integrin αvβ6–LAP binding. In both cases, inhibition of integrin αvβ6–fibronectin binding was stronger than that of integrin αvβ6–LAP binding. This similarity suggests that these mAbs capture key features of anti–integrin αvβ6 antibodies in patients with UC.
In contrast to the mAbs, a small proportion of IgG samples from patients exhibited a strong inhibitory effect on integrin αvβ6–LAP binding. It is possible that these individuals possess high-affinity anti–integrin αvβ antibodies, comparable to the 10D5 antibody. However, given that the majority of IgG samples showed limited or no inhibition of integrin αvβ6–LAP binding, such high-affinity antibodies appear to be relatively uncommon and are unlikely to constitute the predominant anti–integrin αvβ6 antibody population in patients with UC.
-
Discussion
In this study, we established 15 anti–integrin αvβ6 mAbs from PBMCs or lymph node samples of patients with UC. We identified 2 predominant CDR3 amino acid sequences in the heavy chain that were shared among different patients. One sequence was encoded by IGHV3-30*18/IGHD3-10*01/IGHJ6*02 and the other by IGHV1-18*01/IGHD5-18*01/IGHJ6*02. Almost all mAbs reacted with integrin αvβ6 in the presence, but not in the absence, of cations. These mAbs inhibited integrin αvβ6–fibronectin binding, but showed substantially lower inhibitory effect against integrin αvβ6–LAP binding. The binding of mAbs with integrin αvβ6 was inhibited by RGDS peptide. Notably, several mAbs contained the RGD sequence in their heavy chain CDR3. Taken together, these data suggest that these mAbs bind to integrin αvβ6 in an RGD site–dependent manner. Evaluation via ELISA revealed that some of the anti–integrin αvβ6 mAbs cross-reacted against integrin αvβ3. The functional properties of these mAbs are similar to those observed in IgG samples, suggesting that these mAbs capture key features of anti–integrin αvβ6 antibodies in UC.
Our analysis revealed that 2 distinct CDR3 amino acid sequences were particularly prevalent among different patients with UC. The first sequence, encoded by IGHV3-30*18/IGHD3-10*01/IGHJ6*02, was found in 5 mAbs from 4 patients, whereas the second sequence, encoded by IGHV1-18*01/IGHD5-18*01/IGHJ6*02, was shared among 6 mAbs from 4 patients. The presence of such shared heavy chain CDR3 amino acid sequences has been observed in other autoimmune conditions, including anti-desmoglein antibodies in patients with pemphigus (23, 24) and anti-ADAMTS13 antibodies in those with thrombotic thrombocytopenic purpura (25). Indeed, it is increasingly recognized that patients with diverse genetic backgrounds and immunological histories produce stereotyped B cell receptors (BCRs) in response to a specific antigen (26). The identification of these stereotyped anti–integrin αvβ6 antibodies in patients with UC suggests common humoral immune mechanisms underlying the production of these antibodies.
Our antibodies exhibited low SHM levels, often with a few replacement mutations in the CDRs, yet retained antigen reactivity. This is in line with recent findings suggesting that antibody affinity can be optimized not through extensive mutation, but via selective, minimal functional changes to germline VH genes (27). According to a previous report, in plasma cells in the intestinal tract of patients with UC, the IGHJ6 usage is increased, CDR3 length is longer, and the SHM frequency is lower than in healthy controls (17). In this study, most mAbs were found to have a long CDR3; in particular, mAbs encoded by IGHV3-30*18/IGHD3-10*01/IGHJ6*02 had many features, such as increased IGHJ6 usage and low SHM rate, that match earlier reports regarding characteristics of antibodies in patients with UC. The agreement between the current results and those of previous studies (17) may reflect the widespread sharing of these mAbs among patients with UC. The mechanisms of production of these mAbs and the roles of such mAbs in the pathology of UC are yet unknown and require further research.
The mAbs established in this study shared many characteristics with antibodies present in the sera or with IgG from patients with UC who were positive for anti–integrin αvβ6 antibodies (4). Furthermore, some new insights were also gained. Notably, the mAbs in this study comprised both cation-dependent and cation-independent antibodies. The cation-dependent antibodies were expected to bind via the RGD binding site in integrin αvβ6. In contrast, UC4 was the unique antibody that reacted with integrin αvβ6 in a cation-independent manner, with evidence suggesting that it may be reacting with integrin monomers independent of the RGD binding site.
We observed in this study that some anti–integrin αvβ6 mAbs cross-reacted with integrin αvβ3. We have reported earlier that some patients with UC have antibodies that react with integrin αvβ3 (4). Although we cannot ignore the possibility that anti–integrin αvβ3–specific autoantibodies coexist independent of anti–integrin αvβ6 autoantibodies in the sera of patients with UC, our current findings suggest that the previously observed reactivity to integrin αvβ3 of the sera of patients with UC may be explained by cross-reactivity of the anti–integrin αvβ6 antibodies to integrin αvβ3.
Using both mAbs and patient-derived IgG samples, we found that inhibition of integrin αvβ6–LAP binding was consistently weaker than that of fibronectin binding. This likely reflects the stronger affinity of LAP for integrin αvβ6, as shown by our BLI analysis. Accordingly, most mAbs exhibited only limited ability to interfere with LAP binding. Interestingly, a subset of patient IgG samples demonstrated appreciable inhibition of LAP binding; this suggested that high-affinity anti-αvβ6 antibodies are present in a fraction of patients, although they do not appear to represent the predominant anti–integrin αvβ6 antibody population in UC. While the inhibition of fibronectin binding was common, LAP inhibition was generally modest. The in vivo relevance of either effect remains unclear and warrants further investigation.
The major limitation of this study is that both the number of mAbs and the number of patients from whom those mAbs were obtained were rather small. Moreover, patient-derived anti–integrin αvβ6 antibodies were randomly collected; therefore, the results may not reveal the complete picture of anti–integrin αvβ6 antibodies in patients with UC. Given this limited dataset, the broad representativeness of the identified shared CDR3 sequences across patients with UC still remains to be further examined. Experiments using samples from a larger number of patients with UC will be necessary to prove universality of the characteristics of anti–integrin αvβ6 antibodies. In addition, we have not been able to examine whether these anti–integrin αvβ6 antibodies are directly involved in the pathogenesis of UC. In particular, the presence of an RGD motif or the ability to recognize the RGD binding site may enhance the functional impact of certain anti-αvβ6 antibodies by interfering with ligand binding or downstream signaling. Further studies using in vivo models are needed to clarify whether such antibodies contribute distinctively to UC pathogenesis. Classification based on these features may offer new insights into disease mechanisms. Furthermore, we obtained mAbs only at one point for each patient; therefore, sequential samples from the same patient were lacking. Longitudinal studies with serial sampling from a larger number of patients are warranted to explore potential changes in the qualitative and functional characteristics of anti–integrin αvβ6 antibodies during the disease course in individual patients, and to provide important insights into their roles in disease onset and exacerbation.
In conclusion, the mAbs established from patients with UC mainly recognized integrin αvβ6 in its active conformation. This study identified 2 predominant CDR3 amino acid sequences in the heavy chain of mAbs. Most mAbs demonstrated cation-dependent binding to integrin αvβ6 and selective inhibition of integrin αvβ6–fibronectin binding. The functional properties of these mAbs are similar to those observed in IgG samples from 42 patients with UC; this suggests that these mAbs capture key features of anti–integrin αvβ6 antibodies in UC. Taken together, these data indicate that anti–integrin αvβ6 antibodies are deeply involved in the pathophysiology of UC, and these results may provide potential leads for the development of new therapeutic strategies. As this study characterized a limited number of mAbs, further investigation with larger antibody panels will be needed to fully understand the diversity and clinical significance of these autoantibodies.
-
Methods
Sex as a biological variable. This study included samples from 4 male and 3 female patients with UC. Sex was not considered a biological variable in the analyses because the study focused on the molecular and functional characterization of anti–integrin αvβ6 antibodies, which do not show sex-related differences. Therefore, the findings are expected to be relevant to both sexes.
Study design. The purpose of this study was to generate and characterize patient-derived anti–integrin αvβ6 antibodies. We used PBMC or mesenteric lymph node samples from patients with UC to identify B cells expressing anti–integrin αvβ6 antibody by B cell immortalization or single-cell sorting using flow cytometry. The gene of the variable region of the antibody was sequenced to produce an anti–integrin αvβ6 mAb. These sequences were annotated using IgBLAST (28) to identify gene usage and amino acid sequences. Based on our previous reports (4), we performed experiments such as ELISA and solid-phase integrin αvβ6 binding assay to characterize the antibodies.
Patients. A total of 7 patients with UC undergoing treatment at Kyoto University Hospital participated in this study. The clinical characteristics of the patients are summarized in Supplemental Table 1. Diagnosis of patients was based on a combination of their symptoms, endoscopic findings, histological features, and the lack of alternative diagnoses (29, 30). Patients were considered eligible if they were positive for anti–integrin αvβ6 antibodies, regardless of clinical severity or treatment. The study was conducted in accordance with the Declaration of Helsinki and was authorized by the Ethics Committee of the Graduate School of Medicine, Kyoto University. Written informed consent was obtained from all patients after an explanation of the nature and the possible outcomes of the study. Sample sources (lymph nodes or PBMCs) were selected solely on the basis of clinical availability and procedural feasibility. Lymph node samples were obtained from patients undergoing surgical procedures, which facilitated the collection of a sufficient number of B cells. PBMC samples were obtained from patients who did not undergo surgery. Each patient contributed only one type of sample. No notable differences were observed in the characteristics of antibodies derived from PBMCs and those derived from lymph nodes.
Sample preparation. To generate mAbs, blood or lymph node samples were collected from the patients. PBMCs were isolated from whole blood by gradient centrifugation using BD Vacutainer CPT (Becton, Dickinson and Co.). The lymph node samples were obtained from the mesenteric lymph nodes attached to the surgical specimen during total colectomy. Lymph node samples were chopped with a clean scalpel, mashed on a 70 μm mesh using a plunger, and washed with PBS. After centrifugation of the cell suspension at 300g for 5 minutes, the supernatant was removed, and 1× RBC Lysis Buffer (pluriSelect) was added. After 3 minutes, the reaction was stopped by addition of PBS with 0.1% BSA. The resulting cell suspension was subjected to centrifugation at 300g for 5 minutes to obtain lymph node–derived cells, which were preserved at –80°C for further use.
Preparation of Epstein-Barr virus reagent. The Epstein-Barr virus (EBV) reagent was prepared and stocked as a culture supernatant harvested from the B95-8 marmoset cell line obtained from the Japanese Collection of Research Bioresources Cell Bank (cell ID: JCRB9123). B95-8 cells were cultured in RPMI 1640 medium (Fujifilm Wako Pure Chemical) containing 10% FBS at 37°C in 5% CO2 until reaching confluence. The culture medium was centrifuged at 400g for 10 minutes to remove all residue, and the supernatant was divided into aliquots and stored at –80°C until used for transduction.
Establishment of lymphoblastoid cell lines by EBV transduction. Lymphoblastoid cell lines (LCLs) were established as previously reported (31, 32). Briefly, IgM+ B cells were removed from PBMCs using magnetic cell sorting with anti-human IgM MicroBeads following the manufacturer’s instructions (Miltenyi Biotec). PBMCs obtained after removal of IgM+ B cells were suspended in EBV stock (1 × 107 cells/mL) containing 2.5 μg/mL ODN2006 (Alpha Diagnostic International) and kept at 37°C under 5% CO2 for 1 hour. The cells were suspended in RPMI 1640 (2 × 105 cells/mL) containing 20% FBS, streptomycin/penicillin, 2.5 μg/mL ODN2006 CpG, 50 IU recombinant human IL-2 (R&D Systems), and 500 ng/mL cyclosporin A (Tokyo Chemical Industry Co) and seeded at 200 μL/well in round-bottomed 96-well plates. After culture for 2 weeks, ELISA was performed using culture supernatants to identify LCLs producing anti–integrin αvβ6 antibodies, and RNA isolation was performed from these LCLs. Total RNA was extracted from LCLs using an RNeasy Mini Kit (catalog 74106, QIAGEN) following the manufacturer’s protocol.
An alternative method was to scale up the LCLs producing the antibody of interest and then use a cell array method to identify single cells producing anti–integrin αvβ6 antibodies. This method was supported by EVEC Inc. Briefly, antigen-immobilized microbeads were seeded onto microarray chips together with LCLs and allowed to react for several hours. Array chips were washed and stained with anti-human IgG-R-phycoerythrin, and wells with positive beads were identified using fluorescence microscopy. RNA isolation was performed from cells in positive wells.
Single-cell sorting. MAbs were generated from antigen-specific B cells using a single-cell sorting protocol based on a previously reported method (33), with minor modifications as detailed below. PBMCs or lymph node cells were resuspended at 1 × 106 cells per 100 μL in FACS buffer, which comprised 49 mL of Dulbecco’s PBS and 1 mL of FBS. Cells were stained on ice for 20 minutes with the following antibodies: Alexa Fluor 700–mouse anti–human CD20 (1:80; clone L27; catalog 560631, Becton, Dickinson and Co.), APC–mouse anti-human IgG (1:20; clone G18-145; catalog 550931, Becton, Dickinson and Co.), and DAPI (1:100; Thermo Fisher Scientific).
To detect integrin αvβ6–specific B cells, a bait complex was prepared by preincubation of biotinylated integrin αvβ6 (IT6-H82E4, ACROBiosystems) with phycoerythrin-conjugated (PE-conjugated) NeutrAvidin (Thermo Fisher Scientific) in FACS buffer at a fixed volumetric ratio (31:1:18 μL, respectively). This bait complex was applied at an approximate cell-to-bait ratio of 20:1. It enables the detection of integrin αvβ6–specific B cells through fluorescent signal amplification. Appropriate negative controls were included to assess background staining. The gating strategy for detection of integrin αvβ6–specific B cells using PBMCs or lymph node cells by flow cytometry is shown in Supplemental Figure 4.
Before sorting, each well of a 96-well PCR plate was preloaded with 4 μL of sorting buffer, comprising 3.1 μL of nuclease-free H2O, 0.2 μL of RNasin (40 U/μL; Promega), 0.1 μL of RNaseOUT (40 U/μL; Thermo Fisher Scientific), 0.2 μL of 10× PBS (resulting in 0.5× PBS), and 0.4 μL of 100 mM DTT (final concentration 10 mM; Thermo Fisher Scientific). Single viable B cells (CD20+, IgG+, DAPI–, and PE+ for αvβ6 binding) were individually sorted into the wells using a BD FACSAria II cell sorter (Becton, Dickinson and Co.). After sorting, the plates were immediately frozen and stored at –80°C until RNA extraction.
Single-cell cDNA synthesis and PCR for amplifying variable regions of BCRs. To characterize the antibody repertoire at the single-cell level, reverse transcription (RT) and semi-nested PCR were performed to amplify the variable regions of Ig heavy and light chain genes from individual B cells.
Sorted single B cells were thawed on ice and first incubated at 65°C for 2.5 minutes with a random-hexamer primer mix comprising 5.6 μL of nuclease-free H2O, 0.75 μL of random-hexamer primers (200 ng/μL; Thermo Fisher Scientific), 0.5 μL of NP-40 (10%; Thermo Fisher Scientific), and 0.15 μL of RNaseOUT (40 U/μL; Thermo Fisher Scientific). After incubation, the samples were placed on ice for at least 2 minutes.
Subsequently, an RT mix containing 2.05 μL of nuclease-free H2O, 3 μL of 5× RT buffer (Thermo Fisher Scientific), 0.5 μL of dNTP mix (25 mM; Thermo Fisher Scientific), 1 μL of DTT (100 mM), 0.1 μL of RNasin (40 U/μL; Promega), 0.1 μL of RNaseOUT, and 0.25 μL of SuperScript IV reverse transcriptase (200 U/μL; Thermo Fisher Scientific) was added to each well (final volume of RT mix: 7 μL). The RT reaction was performed using the following thermal protocol: 42°C for 10 minutes, 25°C for 10 minutes, 50°C for 10 minutes, and 94°C for 5 minutes, followed by a hold at 4°C. Following RT, the resulting cDNA was diluted with 16 μL of nuclease-free water before PCR amplification.
The variable regions of Ig heavy and light chains were amplified using a semi-nested PCR strategy with Platinum Taq DNA Polymerase or Platinum Taq Green Hot Start DNA Polymerase (Thermo Fisher Scientific), using exactly the same optimized primer set described in the previous study (33), which was specifically designed for efficient amplification of Ig variable regions.
For both the first- and second-round PCRs, reactions were conducted in a 25 μL total volume using Platinum Taq DNA Polymerase (Thermo Fisher Scientific). The master mix comprised the following components per reaction: 14.68 μL of nuclease-free H2O, 2.05 μL of 10× Platinum Taq PCR buffer, 1.23 μL of KB Extender (6%), 0.61 μL of MgCl2 (50 mM), 0.16 μL of dNTP mix (25 mM), and 0.09 μL each of forward and reverse primers (50 μM), using the same primer sets previously optimized and described in detail (33). For the first-round PCR, 6 μL of diluted cDNA was used as a template; subsequently, 1 μL of the first-round PCR product was used as the template in the second-round PCR. For second-round PCR, Platinum Taq Green Hot Start DNA Polymerase (Thermo Fisher Scientific) was used to enable direct visualization of PCR products on agarose gels via the included tracking dye.
The first-round PCR was performed under the following conditions: 94°C for 2 minutes, followed by 50 cycles at 94°C for 30 seconds, 57°C for 30 seconds, and 72°C for 55 seconds. The second-round PCR used similar conditions, with an extended step involving 72°C for 45 seconds. Second-round PCR products were analyzed by agarose gel electrophoresis, and samples of the correct size (approximately 500 bp for heavy chains and 450 bp for light chains) were subjected to Sanger sequencing. The same protocol was applied to RNA derived from LCLs.
Analysis of the V region gene sequence of BCRs. Aliquots of the second-round PCR products were analyzed by 2% agarose gel electrophoresis to confirm the presence of amplicons of the expected size — approximately 500 bp for heavy chain and 450 bp for light chain variable regions. PCR products of the correct size were subjected to Sanger sequencing, and the resulting sequences were annotated using IgBLAST (28) with the IMGT reference database to determine V(D)J gene usage, CDR3 sequences, and repertoire characteristics and to infer clonal relationships. Sequences containing stop codons or out-of-frame rearrangements (i.e., nonproductive sequences) were excluded from further analysis.
Cloning of the V region gene sequence of BCRs. For cloning of BCR variable regions, the first-round PCR product was reamplified using KOD -Plus- Neo polymerase (TOYOBO) and the same optimized primer set with vector-compatible overhangs as described in a previous report (33). PCR comprised the following thermal cycle: 98°C for 30 seconds; 35 cycles of 98°C for 10 seconds, 65°C for 30 seconds, and 72°C for 30 seconds; and 72°C for 2 minutes. Before cloning, amplified PCR products were purified using a PCR purification kit (QIAquick PCR Purification Kit, QIAGEN).
AbVec2.0-IGHG1 (IgG1) (Addgene plasmid 80795), AbVec1.1-IGKC (Igk) (Addgene plasmid 80796), and AbVec1.1-IGLC2-XhoI (Igl) (Addgene plasmid 99575) were used as human antibody expression vectors as previously reported (33). EcoRI and SalI were used for IgG1, EcoRI and BsiWI for IgK, and EcoRI and XhoI for IgL to linearize each vector. PCR products and expression vectors were cloned using In-Fusion Snap Assembly Master Mix (Takara Bio Inc.).
Expression plasmids were obtained by transforming of Stellar Competent Cells (Takara Bio) and purified using the QIAprep Spin Miniprep Kit (QIAGEN). To screen for reactivity to integrin αvβ6, antibodies were produced in HEK293 cells (originally obtained from ATCC, CRL-1573) by transfection with Lipofectamine 3000 (Thermo Fisher Scientific). Transfected cells were maintained in DMEM containing 2% FBS for 5 days, and then the supernatant was used for ELISA. Plasmid DNA from reactive clones was transformed into Stellar Competent Cells and purified using the QIAGEN Plasmid Maxi Kit.
Antibody production. Recombinant antibodies were transiently expressed using the ExpiCHO Expression System (Thermo Fisher Scientific). ExpiCHO cells were cotransfected with a mixture of expression vectors for the heavy and light chains of the antibodies following the manufacturer’s protocol.
After 10 days of culturing of the transfected cells, the clarified culture supernatant was loaded into Ab-Capcher (ProteNova), and mAbs were purified in accordance with the manufacturer’s instructions. Disposable plastic columns (Thermo Fisher Scientific) were used according to the manufacturer’s recommended protocol to obtain solubilized recombinant mAbs.
Preparation of human IgG. Ab-Rapid SPiN (P-013, ProteNova) was used to purify IgG from the sera of patients with UC and controls. The purified IgG was then dialyzed against PBS (pH 7.2), and concentrated by ultrafiltration with an Amicon Ultrafilter (UFC805024, Millipore) followed by storage at –20°C. Purified IgG concentrations were measured using a NanoDrop 2000c spectrophotometer (Thermo Fisher Scientific).
ELISA. ELISA starter accessory kits (E101, Bethyl Laboratories) were used in accordance with the instructions of the manufacturer. Microtiter plates were coated using carbonate-bicarbonate buffer (coating buffer) with 100 μL of 2 μg/mL recombinant human integrin αvβ6 heterodimer proteins (IT6-H52E1, ACROBiosystems), incubated overnight at 4°C, washed 3 times with TBS containing 0.05% Tween 20 (wash solution), and blocked with TBS containing 1% BSA for 30 minutes at approximately 25°C. After 3 washes with wash solution, 10-fold serial dilutions of mAbs starting at 10 μg/mL and 3 μg/mL (diluted with TBS with 0.05% Tween 20 and 1% BSA) were added, and plates were incubated for 1 hour at approximately 25°C. Plates were washed 5 times and incubated with 100 μL goat anti-human IgG antibody conjugated with horseradish peroxidase (1:50,000; ab6759, Abcam) at 25°C for 60 minutes. After washing, the bound antibodies were detected by incubation with 3,30,5,50-tetramethylbenzidine for 10 minutes. Absorbance was noted at 450 nm. EC50 values were calculated by nonlinear regression analysis on the binding curves using GraphPad Prism version 10 (GraphPad Software). ELISA was carried out in the presence or absence of MgCl2 and CaCl2 (1 mM each). MgCl2 and CaCl2 were added to buffer for washing, blocking, and dilution of antibodies. The same method was used to assess reactivity with other integrins. The mAb 10D5 (ab77906, Abcam) was used as the positive control and IgG derived from healthy individuals (143-09501, Fujifilm Wako Pure Chemical) as the negative control.
To examine whether the RGD (Arg-Gly-Asp) peptide blocked the binding of each mAb to integrin αvβ6, the RGDS (Arg-Gly-Asp-Ser) peptide (A9041, Sigma-Aldrich) or the control RGES (Arg-Gly-Glu-Ser) peptide (A5686, Sigma-Aldrich) was added to each mAb before incubation. In experiments with peptides, the final concentration of mAb was 3 μg/mL, and each peptide was adjusted to 5 concentrations: 0 μg/mL, 12.5 μg/mL, 25 μg/mL, 50 μg/mL, and 100 μg/mL.
We used an anti–integrin αvβ6 ELISA kit (catalog 5288, Medical & Biological Laboratories) for detecting anti–integrin αvβ6 IgG antibody titers from patients with UC according to the manufacturer’s instructions.
Biolayer interferometry. The affinity between each mAb and integrin αvβ6 was measured using biolayer interferometry (BLI) with an Octet RED96 (ForteBio). Biotinylated integrin αvβ6 (IT6-H82E4, ACROBiosystems) was loaded at 25 nM in kinetics buffer (0.1% BSA, 0.6 M sucrose, 0.02% Tween 20, 1 mM MgCl2, and CaCl2 in TBS) for 300 seconds onto a SAX2 biosensor (ForteBio). The association of integrin αvβ6 and mAbs at 200, 50, 12.5, 3.13, and 0.78 nM was measured in kinetics buffer for 300 seconds. The measurement range was adjusted according to the dissociation constant (KD) value of each mAb. Dissociation in kinetics buffer was measured for 300 seconds. The on-rate constant (Kon), off-rate constant (Koff), and KD values were calculated using a global fit to a 1:1 binding model.
Solid-phase integrin αvβ6 binding assay. For this assay, 96-well microtiter plates were previously coated using coating buffer with either 5 μg/mL of fibronectin (F0985, Sigma-Aldrich) or 0.5 μg/mL of LAP (LAP-H5213, ACROBiosystems) (100 μL/well, 4°C, overnight). After removal of the coating solution, the plates were blocked by TBS containing 1% BSA. In another 96-well plate, 60 μL/well of a 2× stock (4 μg/mL of integrin αvβ6 with His-tag for fibronectin or 0.4 μg/mL integrin αvβ6 with His-tag for LAP) was combined with 60 μL/well of a 2× stock of each mAb diluted in the same way as for ELISA and incubated for 1 hour. After the ligand-coated plates were washed, 100 μL of the mAb–integrin αvβ6 mixture was transferred to the ligand-coated plate and incubated for 1 hour. After washing of the plate with wash solution, an Anti-His-tag mAb-HRP-DirecT (1:5,000; D291-7, Medical & Biological Laboratories) was added followed by incubation for 60 minutes. After the wash, bound antibodies were incubated with 3,30,5,50-tetramethylbenzidine for 10 minutes for detection. The absorbance was measured at 450 nm.
Statistics. Statistical analysis was conducted using GraphPad Prism (version 10). The correlation between anti–integrin αvβ6 IgG titers and blocking activity of integrin αvβ6–fibronectin or integrin αvβ6–LAP binding was evaluated using the Pearson product-moment correlation. A P value less than 0.05 was considered to indicate statistical significance. For experiments using patient-derived IgG, the cutoff OD values for the antibody titer and the inhibitory effect were defined as the mean value of control IgG plus 3 SD.
Study approval. The study was conducted in accordance with the Declaration of Helsinki and was approved by the Ethics Committee of the Graduate School of Medicine, Kyoto University, Kyoto, Japan (protocol R1004). Written informed consent was obtained from all patients after they were provided with a full explanation of the nature and possible outcomes of the study. Recombinant DNA experiments were performed following approval by the Kyoto University Recombinant DNA Experiment Safety Committee (approval 230103).
Data availability. Values for all data points shown in graphs and values behind any reported means are provided in the Supporting Data Values XLS file accompanying this article. No next-generation sequencing data were generated in this study. All BCR variable region sequences were obtained by Sanger sequencing, which does not fall under the MINSEQE guidelines. All data supporting the findings of this study are included in the article or its supplemental materials. Additional information is available upon reasonable request.
-
Author contributions
IT, YN, MS, and TK conceptualized the study. IT and YN developed methodology. IT and YN performed formal analysis. IT, YN, and DJCJ performed investigation. MS acquired funding. IT and YN provided resources. TC and HS supervised the study. IT performed visualization. IT and YN prepared the original draft of the manuscript. MS, TK, SO, TY, KS, AH, M Yasuda, KC, RN, M Yokode, YM, SM, TM, TC, and HS reviewed and edited the manuscript.
-
Funding support
- Japan Society for the Promotion of Science KAKENHI grant 24K02437 (to MS).
- Japan Agency for Medical Research and Development (23ek0109597 to MS).
-
Acknowledgments
We express our gratitude to the patients who provided blood or lymph node samples for this study. We also thank Shino Yamaguchi and Taichi Ito for their valuable technical support and Reiko Shinkura for writing a review. We are grateful to Medical & Biological Laboratories Co. Ltd. for providing the anti–integrin αvβ6 ELISA kit used in this study. Finally, we thank Editage for English language editing.
Address correspondence to: Masahiro Shiokawa or Yoshihiro Nishikawa, Department of Gastroenterology and Hepatology, Kyoto University Graduate School of Medicine, 54 Shogoin-Kawahara-cho, Sakyo-ku, Kyoto 606-8507, Japan. Phone: 81.75.751.4319; Email: machan@kuhp.kyoto-u.ac.jp (MS); nishi328@kuhp.kyoto-u.ac.jp (YN).
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2026, Takimoto et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(4):e192953.https://doi.org/10.1172/jci.insight.192953.
-
References
- Kobayashi T, et al. Ulcerative colitis. Nat Rev Dis Primers. 2020;6(1):74.
- Ordas I, et al. Ulcerative colitis. Lancet. 2012;380(9853):1606–1619.
- Ramos GP, Papadakis KA. Mechanisms of disease: inflammatory bowel diseases. Mayo Clin Proc. 2019;94(1):155–165.
- Kuwada T, et al. Identification of an anti-integrin αvβ6 autoantibody in patients with ulcerative colitis. Gastroenterology. 2021;160(7):2383–2394.
- Okabe M, et al. Anti-integrin αvβ6 antibody as a biomarker for diagnosing ulcerative colitis: a nationwide multicenter validation study. J Gastroenterol. 2025;60(1):86–95.
- Livanos AE, et al. Anti-integrin αvβ6 autoantibodies are a novel biomarker that antedate ulcerative colitis. Gastroenterology. 2023;164(4):619–629.
- Marafini I, et al. Diagnostic value of anti-integrin αvβ6 antibodies in ulcerative colitis. Dig Liver Dis. 2024;56(1):55–60.
- Rydell N, et al. Measurement of serum IgG anti-integrin αvβ6 autoantibodies is a promising tool in the diagnosis of ulcerative colitis. J Clin Med. 2022;11(7):1881.
- Barczyk M, et al. Integrins. Cell Tissue Res. 2010;339(1):269–280.
- Koivisto L, et al. Integrin αvβ6: structure, function and role in health and disease. Int J Biochem Cell Biol. 2018;99:186–196.
- Annes JP, et al. Integrin alphaVbeta6-mediated activation of latent TGF-beta requires the latent TGF-beta binding protein-1. J Cell Biol. 2004;165(5):723–734.
- Chen X, et al. Intestinal epithelial cell-derived integrin αβ6 plays an important role in the induction of regulatory T cells and inhibits an antigen-specific Th2 response. J Leukoc Biol. 2011;90(4):751–759.
- Knight PA, et al. Enteric expression of the integrin alpha(v)beta(6) is essential for nematode-induced mucosal mast cell hyperplasia and expression of the granule chymase, mouse mast cell protease-1. Am J Pathol. 2002;161(3):771–779.
- White JB, et al. ImmunoPET imaging of αvβ6 expression using an engineered anti-αvβ6 Cys-diabody site-specifically radiolabeled with Cu-64: considerations for optimal imaging with antibody fragments. Mol Imaging Biol. 2018;20(1):103–113.
- Castro-Dopico T, et al. Targeting B cells for inflammatory bowel disease treatment: back to the future. Curr Opin Pharmacol. 2020;55:90–98.
- Frede A, et al. B cell expansion hinders the stroma-epithelium regenerative cross talk during mucosal healing. Immunity. 2022;55(12):2336–2351.
- Uzzan M, et al. Ulcerative colitis is characterized by a plasmablast-skewed humoral response associated with disease activity. Nat Med. 2022;28(4):766–779.
- Kitaura K, et al. Different somatic hypermutation levels among antibody subclasses disclosed by a new next-generation sequencing-based antibody repertoire analysis. Front Immunol. 2017;8:389.
- Weinreb PH, et al. Function-blocking integrin alphavbeta6 monoclonal antibodies: distinct ligand-mimetic and nonligand-mimetic classes. J Biol Chem. 2004;279(17):17875–17887.
- Hall ER, Slack RJ. The effect of divalent metal cations on the αv integrin binding site is ligand and integrin specific. Biomed Pharmacother. 2019;110:362–370.
- Luo BH, et al. Structural basis of integrin regulation and signaling. Annu Rev Immunol. 2007;25:619–647.
- Tiwari S, et al. Divalent cations regulate the folding and activation status of integrins during their intracellular trafficking. J Cell Sci. 2011;124(pt 10):1672–1680.
- Yamagami J, et al. Homologous regions of autoantibody heavy chain complementarity-determining region 3 (H-CDR3) in patients with pemphigus cause pathogenicity. J Clin Invest. 2010;120(11):4111–4117.
- Cho MJ, et al. Shared VH1-46 gene usage by pemphigus vulgaris autoantibodies indicates common humoral immune responses among patients. Nat Commun. 2014;5:4167.
- Schaller M, et al. The splenic autoimmune response to ADAMTS13 in thrombotic thrombocytopenic purpura contains recurrent antigen-binding CDR3 motifs. Blood. 2014;124(23):3469–3479.
- Henry Dunand CJ, Wilson PC. Restricted, canonical, stereotyped and convergent immunoglobulin responses. Philos Trans R Soc Lond B Biol Sci. 2015;370(1676):20140238.
- Merkenschlager J, et al. Regulated somatic hypermutation enhances antibody affinity maturation. Nature. 2025;641(8062):495–502.
- Ye J, et al. IgBLAST: an immunoglobulin variable domain sequence analysis tool. Nucleic Acids Res. 2013;41(web server issue):W34–W40.
- Kornbluth A, et al. Ulcerative colitis practice guidelines in adults: American College Of Gastroenterology, Practice Parameters Committee. Am J Gastroenterol. 2010;105(3):501–523.
- Maaser C, et al. ECCO-ESGAR guideline for diagnostic assessment in IBD Part 1: initial diagnosis, monitoring of known IBD, detection of complications. J Crohns Colitis. 2019;13(2):144–164.
- Minamitani T, et al. Novel neutralizing human monoclonal antibodies against tetanus neurotoxin. Sci Rep. 2021;11(1):12134.
- Valgardsdottir R, et al. Identification of human SARS-CoV-2 monoclonal antibodies from convalescent patients using EBV immortalization. Antibodies (Basel). 2021;10(3):26.
- Gieselmann L, et al. Effective high-throughput isolation of fully human antibodies targeting infectious pathogens. Nat Protoc. 2021;16(7):3639–3671.
-
Version history
- Version 1 (January 8, 2026): In-Press Preview
- Version 2 (February 23, 2026): Electronic publication