Research ArticleHematologyOncology
Open Access |  10.1172/jci.insight.193779
10.1172/jci.insight.193779
Characterization of the clonal hierarchy and immunophenotype of PTPN11 mutations in acute myeloid leukemia
Sydney Fobare,1,2 Chia Sharpe,2 Kate Quinn,2 Kinsey Bryant,2 Linde A. Miles,3,4,5 Robert L. Bowman,6 Carolyn Cheney,2 Casie Furby,2 Marissa Long,2 Kaytlynn Fyock,2 Ben Wronowski,2 James R. Lerma,2 Krzysztof Mrózek,7 Deedra Nicolet,7,8 Thomas M. Sesterhenn,2 Megan E. Johnstone,2 Jianmin Pan,9,10,11 Shesh N. Rai,9,10,11 Chandrashekhar Pasare,4 Nives Zimmermann,12 Wen-Mei Yu,13 Cheng-Kui Qu,13 Andrew Carroll,14 Richard Stone,15 Eunice S. Wang,16 Jonathan Kolitz,17 Bayard Powell,18 John P. Perentesis,5 Ann-Kathrin Eisfeld,7,19 Erin Hertlein,2 and John C. Byrd2
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Fobare, S. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Sharpe, C. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Quinn, K. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Bryant, K. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Miles, L. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Bowman, R. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Cheney, C. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Furby, C. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Long, M. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Fyock, K. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Wronowski, B. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Lerma, J. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by
Mrózek, K.
in:
PubMed
|
Google Scholar
|

1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Nicolet, D. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Sesterhenn, T. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Johnstone, M. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Pan, J. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Rai, S. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Pasare, C. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Zimmermann, N. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Yu, W. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by
Qu, C.
in:
PubMed
|
Google Scholar
|

1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Carroll, A. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Stone, R. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Wang, E. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Kolitz, J. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Powell, B. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by
Perentesis, J.
in:
PubMed
|
Google Scholar
|

1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Eisfeld, A. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Hertlein, E. in: PubMed | Google Scholar
1Medical Scientist Training Program, The Ohio State University, Columbus, Ohio, USA.
2Department of Internal Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
3Division of Experimental Hematology and Cancer Biology, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
4Department of Pediatrics, Cincinnati Children’s Hospital Medical Center, Cincinnati, Ohio, USA.
5Cancer and Blood Diseases Institute, Cincinnati Children’s Hospital, Cincinnati, Ohio, USA.
6Department of Cancer Biology, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
7Clara D. Bloomfield Center for Leukemia Outcomes Research, and
8Alliance Statistics and Data Management Center, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
9Biostatistics and Informatics Shared Resource, University of Cincinnati Cancer Center, Cincinnati, Ohio, USA.
10Cancer Data Science Center, and
11Department of Biostatistics, Health Informatics and Data Sciences, University of Cincinnati College of Medicine, Cincinnati, Ohio, USA.
12Pathology and Laboratory Medicine, University of Cincinnati, Cincinnati, Ohio, USA.
13Aflac Cancer and Blood Disorders Center, Department of Pediatrics, Winship Cancer Institute, Emory University School of Medicine, Atlanta, Georgia, USA.
14Department of Genetics, University of Alabama at Birmingham, Birmingham, Alabama, USA.
15Dana-Farber/Partners CancerCare, Boston, Massachusetts, USA.
16Roswell Park Comprehensive Cancer Center, Buffalo, New York, USA.
17Monter Cancer Center, Zucker School of Medicine at Hofstra/Northwell, Lake Success, New York, USA.
18Wake Forest Baptist Comprehensive Cancer Center, Winston-Salem, North Carolina, USA.
19Division of Hematology, Department of Internal Medicine, The Ohio State University Comprehensive Cancer Center, Columbus, Ohio, USA.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Find articles by Byrd, J. in: PubMed | Google Scholar
Authorship note: SF and CS contributed equally to this work and have been designated as co–first authors. EH and JCB contributed equally to this work and have been designated as co–senior authors.
Published February 23, 2026 - More info
JCI Insight. 2026;11(4):e193779. https://doi.org/10.1172/jci.insight.193779.
© 2026 Fobare et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: March 27, 2025; Accepted: January 2, 2026
-
Abstract
Mutations in protein tyrosine phosphatase non-receptor type 11 (PTPN11) have been considered late acquired mutations in acute myeloid leukemia (AML) development. Using single-cell DNA sequencing, we found that PTPN11 mutations can occur as initiating events in some patients with AML when accompanied by strong oncogenic drivers, commonly NPM1 mutations. The resulting AML has a diverse set of variably differentiated myeloid cells with few myeloid cells that lack leukemic mutations. The role of Ptpn11 as a codriver was confirmed in a murine model that exhibits an AML phenotype with a comparable immune diversity that is serially engraftable and reconstituted from early precursor cells. Furthermore, lineage-negative bone marrow cells from these mice reconstitute the full diversity of mature myeloid cells, and these cells exhibit an altered cytokine response after physiologic stimulation. Our work highlights how PTPN11-mutated AML is derived from a multitude of codominant and late acquired aberrations that have a previously unrecognized differentiated myeloid clonal expansion potentially contributing to pathogenesis of the disease.
-
Introduction
Acute myeloid leukemia (AML) is the most common diagnosed acute leukemia in adults and for most patients it has extremely poor outcomes. In younger patients, leukemogenesis most commonly involves early acquisition of a distinct balanced translocation or mutations with strong oncogenic potential, followed by acquisition of additional late genetic mutations in signaling genes (e.g., KIT, FLT3, NRAS, KRAS, NF1, CBL, and PTPN11) (1). In contrast, in most older adults, AML is characterized by the early acquisition of a variety of clonal hematopoiesis (CH) mutations (e.g., DNMT3A, TET2, and ASXL1) that are not by themselves transforming but rather promote inflammation and confer a stem cell/progenitor growth advantage and increased risk of hematologic malignancy and multiple other complications (2). CH mutations are then followed by acquisition of additional signaling mutations or cytogenetic abnormalities that lead to the spectrum of neoplastic development: cytopenias without dysplasia, myelodysplastic syndrome, and/or ultimately AML (3). Characterization of the impact of gene mutations and cytogenetic abnormalities on clinical outcome in AML has been mostly derived from treatment trials with intensive chemotherapy, and as a result a distinct group of driver and late mutations are well characterized as predictors of poor outcome in the 2022 European LeukemiaNet (ELN) classification of AML (4).
The development of now approved targeted agents toward FMS-like tyrosine kinase 3 (FLT3) (midostaurin, gilteritinib, and quizartinib), IDH1 (ivosidenib, olutasidenib), IDH2 (enasidenib), and BCL2 (venetoclax) have transformed the treatment landscape in AML. Within these new targeted therapeutic approaches, many established mutations associated with poor prognosis, such as TP53 mutations, have continued to predict poor outcomes. However, additional mutations such as MAPK-pathway-activating mutations (e.g., NRAS, KRAS, CBL, NF1, and PTPN11) have emerged as specifically impacting the outcomes of patients receiving targeted therapies (5–8). Indeed, patients with PTPN11-mutated AML had the lowest response among patients carrying other genomic abnormalities who received IDH2 inhibitors (9) or venetoclax/azacitidine (10, 11). Studies to identify the pathogenic features of clonal evolution and compound genetic models of PTPN11-mutated AML are lacking.
The PTPN11 gene encodes Src homology region 2–containing (SH2-containing) protein tyrosine phosphatase 2 (SHP2). SHP2 contains 2 SH2 domains and a tyrosine phosphatase domain. SHP2 is capable of self-regulation, as the N-terminal SH2 domain can bind to the active phosphatase domain to prevent dephosphorylation of SHP2 targets (12). Pathogenic PTPN11 mutations tend to occur in regions involved in its self-regulation, and thus, these gain-of-function mutations result in activation of numerous signaling pathways (e.g., RAS/ERK1/2, FLT3, JAK/STAT, PI3K/AKT, and NF-κB), mediates immune evasion via PD-1, and induces the overexpression of antiapoptotic proteins such as MCL-1 (13–15). In humans, gain-of-function PTPN11 mutations can occur as germline mutations in Noonan syndrome and are associated with a higher frequency of juvenile myelomonocytic leukemia, suggesting these mutations can contribute to the initiation of myeloid malignancies (16, 17). To study the role of PTPN11 gain-of-function mutations, Xu et al. developed the Ptpn11E76K mouse that mimics the most common hotspot mutation in both Noonan syndrome and myeloid malignancies (18). They found that constitutive expression of Ptpn11E76K is embryonically lethal in mice, but conditional, heterozygous expression of the Ptpn11E76K mutation under the control of Mx-Cre promotes the development of multilineage leukemias (approximately equal frequency of myeloid, B cell, and T cell leukemias).
These findings have prompted several groups, including our own, to examine the prognostic impact of PTPN11 mutations in AML. These analyses suggest that PTPN11 mutations co-occur most commonly with either NPM1 mutations or with chromosome translocations or inversions and typically confer poor outcome (19–21). In the absence of an NPM1 mutation, PTPN11 mutations often co-occur with cytogenetic abnormalities associated with dysregulation of EVI1. Inconsistencies between studies suggest that the pathogenic contribution of PTPN11 mutations may depend on its context among other driver mutations (19). Notably, in some patients, PTPN11 mutations occur at a high variant allele frequency (VAF), suggesting they could be initiating or co-initiating events in leukemogenesis. However, due to limitations of previous sequencing technologies it has not been possible to determine when PTPN11 mutations arise during AML development. Understanding the clonal evolution of PTPN11 mutations is clinically relevant, as previous reports have suggested that in patients treated with intensive chemotherapy, dominant PTPN11 mutations are associated with better outcomes compared with subclonal mutations, although this is confounded by the fact that patients with subclonal mutations tend to have a higher comutational burden (20). However, as an early driver mutation PTPN11 has serious implications for predicting therapeutic response, and as an acquired subclonal mutation PTPN11 can contribute to the emergence of resistance to therapy, both resulting in poor outcome. Given the lack of knowledge related to clonal evolution of PTPN11 mutations and characterization of myeloid immunophenotype of this disease, we initiated studies in both primary human samples and developed what we believe is the first murine model of concurrent Npm1 and Ptpn11 mutations to address these questions. Herein, we demonstrate that PTPN11 can be an early acquired mutation, commonly co-occurring with a strong oncogene (NPM1 mutation) to promote AML development. The NPM1- and PTPN11-mutated subtype of AML has a diverse immune profile in patients that was reproduced in our Npm1cA/Ptpn11E76K animal model that exhibits capacity for serial engraftment. This model provides a system to understand the role of early PTPN11 mutations as they relate to clonal architecture and influence on AML immunophenotype.
-
Results
PTPN11-mutated AML demonstrates multiple-ancestor origins. Previous work from our group demonstrated that PTPN11 mutations occur at a wide range of VAFs (0.05 up to 0.54), suggesting that PTPN11 mutations may occur early in leukemogenesis (19). Previous studies that have evaluated PTPN11 mutations using single-cell sequencing techniques have mainly included low-frequency PTPN11 mutations (22–24). To further elucidate the genomic characteristics of dominant PTPN11 mutations, we assembled a cohort of patients enriched with those that had a VAF of greater than 0.3 and mutations that fell within the PTPN11 amplicons generated by the Tapestri single-cell DNA sequencing (scDNA-seq) Myeloid Clonal Evolution Published Panel (Mission Bio). All Alliance patient samples that we had access to and met these 2 requirements were included in the study. Four patients with confirmed PTPN11 mutations without a co-occurring NPM1 mutation by bulk sequencing and 9 with concurrent PTPN11 and NPM1 mutations were analyzed. The demographics and mutation profile identified by bulk sequencing (VAF > 0.05) of the included patients are summarized in Table 1. All 4 patients without NPM1 mutations had a strong driver translocation or inversion involving EVI1 together with monosomy 7 (cases AML-1 to AML-3) or t(6;11) (q27;q23) (case AML-5). However, chromosomal changes are not captured by this scDNA-seq panel, so in these samples a single PTPN11-mutated clone was identified (representative case AML-3; Figure 1A and Supplemental Figure 1; supplemental material available online with this article; https://doi.org/10.1172/jci.insight.193779DS1).
 Figure 1
Figure 1Clonal architecture of PTPN11-mutated AML patient samples. Clonographs demonstrating the frequency of co-occurring mutations determined using scDNA-seq in representative samples. (A) PTPN11 mutation without NPM1 mutation (AML-3; total n = 4). (B) Subclonal PTPN11 mutation (AML-16, total n = 4). (C) Inferred clonal relationship for B. (D) Early PTPN11 mutation (AML-9, total n = 5). (E) Inferred clonal relationship for D.
Among the 9 patients with both PTPN11 and NPM1 mutations, we identified that half the patients had subclonal PTPN11 mutations and the other half had PTPN11 mutations in the first leukemic clone identified. Two patient samples (AML-11 and AML-16) showed evidence of an NPM1-mutated clone that lacked a PTPN11 mutation (representative case AML-16; Figure 1B and Supplemental Figure 1), suggesting that the NPM1 mutation preceded the PTPN11 mutation. However, the double-mutant cells still generated the dominant clone (Figure 1C). Two patient samples (AML-13 and AML-15) had the acquisition of a TET2 or DNMT3A mutation followed by PTPN11 and then NPM1 mutations. The clonal architecture in these 4 samples aligns with the predominant understanding of PTPN11 mutations as a late acquired signaling mutation akin to FLT3 or RAS family mutations and is supported by previous reports of scDNA-seq performed on samples from patients with AML (22, 23). In the remaining 5 patients (AML 6–10), we observed PTPN11 mutations in the earliest detectable clone (representative case AML-9; Figure 1D and Supplemental Figure 1). In 4 of these 5 patients, no single-mutant clone was identified, which precluded the ability to confirm the initiating mutation in these samples. Nevertheless, these data support the early emergence of a PTPN11 mutation in the clonal evolution of these patients (Figure 1E). These data align with previous single-cell sequencing studies where signaling mutations can occur prior to NPM1 mutations (24). In 1 patient sample, we identified a PTPN11 single-mutant clone in addition to a PTPN11 and DNMT3A double-mutant clone and a PTPN11, DNMT3A, and NPM1 triple-mutant clone. This suggests that in some patients PTPN11 mutations may be an early event in AML leukemogenesis and can occur prior to the acquisition of other mutations typically described as driver mutations (AML-8; Supplemental Figure 1). Collectively, these data support the conclusion that while PTPN11 mutations are frequently a late acquired event, in contrast to other RAS family mutations, they can also serve as a driver or codriver mutations in AML when coupled with mutations or chromosomal abnormalities that are strongly associated with myeloid oncogenesis.
Most myeloid cells in PTPN11-mutated AML have leukemic mutations at diagnosis. Concurrent with the scDNA-seq results (Figure 1), we performed simultaneous single-cell immunophenotyping in primary AML samples using an antibody panel against 42 antigens (Mission Bio Tapestri). Distinct immunophenotypic clusters were identified using shared nearest neighbor clustering. We first generated a uniform manifold approximation and projection (UMAP) for data reduction based on the surface marker expression. As expected, the AML clones clustered separately from the wild-type (WT) cells within a patient sample (AML-6; Figure 2A). Heatmaps were also generated to visualize the coexpression of cell surface markers. Here we can identify clusters of lymphoid cells without leukemic mutations (yellow, teal, tan, and dark green clusters). Within the clusters associated with leukemic mutations, there are cells that express very few myeloid markers (light green cluster), immature myeloid markers such as CD117, CD34, and CD123 (navy cluster), and clusters that express mature myeloid markers such as CD14, CD11b, and HLA-DR (pink cluster) or CD11c and CD141 (red cluster, Figure 2B). However, we could not identify a cluster with mature myeloid markers composed of predominately WT cells, thus verifying that the mature myeloid cells in these patients are leukemia derived (Figure 2C). This trend was consistent across all 13 samples analyzed (Supplemental Figure 2). Therefore, our data suggest that AML patients with PTPN11 mutations have a diverse immunophenotype, with leukemic cells expressing a variety of mature myeloid markers and very few WT myeloid cells at diagnosis.
 Figure 2
Figure 2Mutated AML cells comprise most myeloid cells at diagnosis. Immunophenotypic clusters were identified using shared nearest neighbor clustering on a representative early PTPN11-mutated sample (AML-6). (A) UMAP overlaid with WT, NPM1287/PTPN11D61V, and NPM2288*fs/PTPN11D61V/DNMT3AR882H clone identity and nearest neighbor clustering. (B) Heatmap of select lineage defining markers. A total of 9 samples were analyzed. (C) UMAP overlaid with the identity of the clusters from the heatmap in B.
The diverse immunophenotype of PTPN11-mutated AML is recapitulated with flow cytometry. To further validate the diverse immunophenotype of the AML cells as suggested by the single-cell immunophenotype, we developed a spectral flow cytometry immunophenotyping antibody panel against 38 antigens (gating strategy in Supplemental Figure 3). AML leukemic stem cells are canonically defined as CD34+ cells (25), but interestingly, we did not identify a large portion of CD34+ cells within the PTPN11-mutated samples (Figure 3A). This is consistent with previous data showing that NPM1 mutations are associated with decreased CD34 expression (26). We did, however, note the presence of cells committed to a mature myeloid lineage. In our cohort, we found that about half the patients had an expansion of CD14+ monocytes above the mean; however, this was not statistically significant (Figure 3B). There was also a trend toward an increase in pre-conventional dendritic cells (pre-cDCs) in bone marrow of patients with PTPN11 mutations at diagnosis (FDR-adjusted P = 0.0781; Figure 3C). There were a couple of patients with increased plasmacytoid DCs (pDCs) in the peripheral blood but not sufficient to be characterized as pDC-AML (Figure 3D). There was also an expansion of immature DCs in both the peripheral blood and bone marrow compared with healthy controls (FDR-adjusted P = 0.0004 and FDR-adjusted P = 0.0681, respectively; Figure 3E). Overall, we found that the tumor-derived myeloid cells exhibit a range of early differentiated cells predominantly of monocytic and DC lineages. When these cells are stimulated with CpG overnight, they have a reduced capacity to produce IFNs compared with healthy controls, highlighting a functional difference from normal, nonmutant cells (Supplemental Figure 4). Therefore, the combination of scDNA-seq and flow cytometry data highlights the presence of mutated leukemia cells with a variable immune phenotype representing both immature and mature populations.
 Figure 3
Figure 3Immunophenotypic diversity of primary PTPN11- and NPM1-mutated AML peripheral blood and bone marrow samples at diagnosis. (A) Percentage of CD34+CD38– and CD34+CD38+ cells in healthy and leukemic peripheral blood (PB) and bone marrow (BM) with representative flow plots. (B) Percentage of CD14+ monocytes in healthy and leukemic PB and BM with representative flow plots. (C) Percentage of pre-conventional dendritic cells (pre-cDCs) in healthy and leukemic PB and BM with representative flow plots. (D) Percentage of plasmacytoid DCs (pDCs) in healthy and leukemic PB and BM with representative flow plots. (E) Percentage of CD1c+ DCs, CD141+ DCs, and immature DCs in healthy and leukemic PB and BM, respectively, with representative flow plots. Percentages are based on CD45+ cells. Data are presented as mean ± SD of 2 independent experiments. Statistical analysis by 1-way ANOVA with Benjamini-Hochberg FDR correction applied. *FDR-adjusted P ≤ 0.05, ***FDR-adjusted P ≤ 0.001.
Npm1cA/Ptpn11E76K mice consistently develop AML. Our AML patient data suggest that PTPN11 mutations can constitute a driving transformative event and can produce a diverse myeloid phenotype. Therefore, we aimed to develop a Ptpn11-mutated AML mouse model that would allow further investigation of the role of Ptpn11 mutations in leukemogenesis and pathogenesis. Analysis of co-occurring mutations with PTPN11 aberrations demonstrated NPM1 to be most co-associated in AML. We therefore combined Ptpn11E76K (18) and Npm1cA (27) mutant alleles, which are both driven by Mx-Cre (28) and are inducible with IFN or polyinosinic-polycytidylic acid [poly(I:C)]. Mice expressing both mutant alleles (Npm1cA/Ptpn11E76K) exhibited a fully penetrant lethal disease phenotype with a median overall survival of 13.4 weeks after induction, significantly faster than either Ptpn11E76K/Mx-Cre (Ptpn11E76K; 39.2 weeks) or Npm1cA/Mx-Cre (Npm1cA; 35 weeks) mice (Figure 4A). When comparing overall survival of Npm1cA/Ptpn11E76K mice based on sex, female mice had a significantly shorter survival compared with male mice (Supplemental Figure 5).
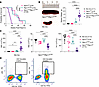 Figure 4
Figure 4Characterization of the Npm1cA/Ptpn11E76K mouse model. (A) Overall survival of Npm1cA, Ptpn11E76K, and Npm1cA/Ptpn11E76K mice. (B) Representative picture of a spleen collected from a Npm1cA/Ptpn11E76K mouse at survival endpoint compared with age-matched Mx-Cre mouse spleen. Summary of spleen weights (in grams) of Npm1cA/Ptpn11E76K mice at time of death compared with aged-matched controls. (C) White blood cell (WBC), red blood cell (RBC), and platelet (PLT) counts of Npm1cA/Ptpn11E76K mice at time of death compared with aged-matched controls. (D) Representative peripheral blood flow cytometry plot (gated on CD45+CD3–CD19–NK1.1– cells; percentage based on total CD45+ cells) of Mx-Cre and Npm1cA/Ptpn11E76K mice 4 weeks after poly(I:C) induction (total n of 11 Mx-Cre mice and 14 Ptpn11E76K/Npm1cA mice). Data are presented as mean ± SD. Statistical analysis by 1-way ANOVA with Benjamini-Hochberg FDR correction applied. *FDR-adjusted P ≤ 0.05, ***FDR-adjusted P ≤ 0.001, ****FDR-adjusted P ≤ 0.0001.
At time of sacrifice, Npm1cA/Ptpn11E76K mice had significant splenomegaly compared with age-matched Mx-Cre and single-gene control mice (Figure 4B). Npm1cA/Ptpn11E76K mice also exhibited significant leukocytosis, anemia, and thrombocytopenia compared with age-matched single-gene controls (Figure 4C). There was no difference in splenomegaly, leukocytosis, anemia, or thrombocytopenia between sexes at time of death (Supplemental Figure 5). As previously published, the Ptpn11E76K mouse can develop multilineage leukemias; therefore, we wanted to confirm that the addition of an Npm1cA mutation would result in a consistent myeloid phenotype. We performed flow cytometric analysis, showing that all Npm1cA/Ptpn11E76K mice have a clear expansion of CD11b+c-Kit+ cells in the blood as early as 4 weeks after induction, which progressively expanded throughout disease development (Figure 4D).
Broad immunophenotyping identifies diverse myeloid populations in the Npm1cA/Ptpn11E76K mice reflective of human AML. As we noted a diverse immunophenotype in primary human AML samples at diagnosis, we comprehensively characterized the Npm1cA/Ptpn11E76K murine disease immunophenotype to see how well this represented human disease. To this end, we developed a spectral flow cytometry immunophenotyping antibody panel against 35 antigens to characterize both the lymphoid and myeloid populations in the spleen (gating strategy in Supplemental Figure 6). Compared with age-matched single-gene controls, the Npm1cA/Ptpn11E76K mice exhibited an expansion of multiple distinct myeloid populations, including neutrophils (CD11b+Ly6G+) and monocytic cells (CD11b+Ly6C+) (Figure 5A). As PTPN11-mutated AML is associated with a monocytic phenotype (21), it was unsurprising to find that the Npm1cA/Ptpn11E76K mice have an increase in the percentage of monocytes compared with control mice. However, in addition to the expected expansion of the CX3CR1–c-Kit+ population of “immature” monocytic cells, we also observed a significant expansion of the CX3CR1+c-Kit– population of “mature” monocytes (Figure 5, B and C). This suggests that while Npm1 mutations have been associated with differentiation block, in the context of a co-occurring Ptpn11 mutation, there is an expansion of variably differentiated myeloid cells, including those immunophenotypically consistent with mature monocytes.
 Figure 5
Figure 5Immunophenotyping of Npm1cA/Ptpn11E76K splenocytes. Flow cytometric analysis of splenocytes from Mx-Cre (n = 5), Npm1cA (n = 6), Ptpn11E76K (n = 5), and Npm1cA/Ptpn11E76K (n = 10) mice. Percentages were calculated as a fraction of total CD45.2+ cells and all control mice were age matched. (A) Percentage of immune subsets in the spleen. (B) Representative flow plots CX3CR1 and c-Kit expression on myeloid cells in a representative Mx-Cre and Npm1cA/Ptpn11E76K mouse. (C) Percentage of immature myeloid cells and monocytes. (D) Representative flow plots of dendritic cell (DC) gating from a Npm1cA/Ptpn11E76K mouse. (E) Percentages of conventional DC 1 (cDC1), cDC2, and immature DC populations. (F) Percentages of immature and mature plasmacytoid DC (pDC) populations. Data are presented as mean ± SD from 3 independent experiments. Statistical analysis by 1-way ANOVA with Benjamini-Hochberg FDR correction applied. *FDR-adjusted P ≤ 0.05, **FDR-adjusted P ≤ 0.01, ***FDR-adjusted P ≤ 0.001, ****FDR-adjusted P ≤ 0.0001. NK cells, natural killer cells.
In addition to the expansion of monocytes, we also saw an expansion of variably differentiated DCs in the murine splenocytes comparable to the primary AML samples (Figure 3). Specifically, the Npm1cA/Ptpn11E76K mice have an expansion of cDC1s and cDC2s in the spleen. There was also an expansion of immature DCs (CD11c+MHClo/–) compared with Mx-Cre and Npm1cA control mice but not to the Ptpn11E76K mice, suggesting a relationship to the Ptpn11 mutation (Figure 5, D and E). There was also an increased percentage of immature (CD11c+B220+) and mature pDCs (CD11c+B220+Siglec H+CD317+) in the Npm1cA/Ptpn11E76K mice compared with controls (Figure 5F). There was no difference in the immunophenotype of the splenocytes from the male and female Npm1cA/Ptpn11E76K mice, suggesting that the difference in survival is not related to an expansion of a specific myeloid cell (data not shown). Therefore, our Npm1cA/Ptpn11E76K mouse can recapitulate the monocytic and cDC expansion of human PTPN11-mutated AML and also exhibits an expansion of pDCs.
Npm1cA/Ptpn11E76K splenocytes can recapitulate a leukemic phenotype in primary and secondary recipients. Thus far, we have demonstrated that co-induction of Npm1cA and Ptpn11E76K in mice is comparable to our human data and that these 2 mutations are sufficient to develop a 100% penetrant, aggressive AML. However, a hallmark of a true AML model requires that it can be adoptively transferred with preservation of its immunophenotypic features. We therefore wanted to determine whether AML cells from the Npm1cA/Ptpn11E76K mice formed a lethal leukemia in a primary recipient. We found that these splenocytes, when injected via the tail vein of NCG recipients, were able to recapitulate a leukemic phenotype with an average overall survival of 8.2 weeks after engraftment (individual donors [n = 8] ranged from 7.5 to 17 weeks, Figure 6A). While all individual AML donors formed a lethal leukemia in some recipients, 2 of the 8 donors tested (S0283 and S0500) had slightly more inconsistent engraftment, resulting in failed engraftment in 2 of 5 and 1 of 4 recipients, respectively. We then took passaged splenocytes from 3 donors (S0200, S0437, and S0500) and engrafted them into additional recipients (n = 5 recipients per donor) and found they were capable of engrafting into a secondary recipient with an average overall survival of 8.5 weeks after engraftment (individual donors [n = 3] ranged from 7.6 to 9.7 weeks, Figure 6B). The consistency increased with the secondary engraftment, with all 5 recipients from each donor resulting in a lethal leukemia. To confirm that the leukemic cells developing in the recipients phenotypically matched the original donor cells, we performed the immunophenotyping panel on splenocytes from mice with overt leukemia, as described above. We found the overall diversity of cell subsets shows a similar profile to that of the donor in both the primary and secondary engraftment. The variability of differentiated monocytes and DCs was maintained at similar proportions in both the primary and secondary recipients (Figure 6C), suggesting stability of the diverse immunophenotype of Npm1cA/Ptpn11E76K leukemia cells.
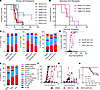 Figure 6
Figure 6Engraftment of Npm1cA/Ptpn11E76K splenocytes into immunodeficient mice. (A) Overall survival in primary NCG engraftment in 3 independent experiments. (B) Overall survival in secondary NCG engraftment. (C) Summary of spleen immune subsets as a proportion of total CD45.2+ cells in primary AML donors compared to primary and secondary NCG recipients meeting the survival endpoint from 2 independent experiments. (D) Frequency of donor-derived CD45.2+ cells over time in the peripheral blood from NCG mice engrafted with LSK cells. (E) Immune cell subsets in spleen from 1 donor (S0200) and recipients of either bulk splenocytes (n = 4 recipients) or LSK cells (n = 3 recipients) at endpoint. FACS-isolated CD45+, CD11c+, or CD11c–CD11b+ cells were sorted from 3 donors. (F) Frequency of donor-derived CD45.2+ cells over time in the peripheral blood. (G) White blood cell (WBC) and (H) overall survival of NCG mice engrafted with CD45+ (n = 12), CD11c+ (n = 11), or CD11c–CD11b+ (n = 8) cells. Data are presented as the mean. pDCs, plasmacytoid dendritic cells; cDC2s, conventional dendritic cells 2; cDC1s, conventional dendritic cells 1; DCs, dendritic cells; NK cells, natural killer cells.
Npm1cA/Ptpn11E76K LSKs can recapitulate the immune subset diversity in primary recipients. With Mx-Cre–driven expression, all hematopoietic lineage cells in the Npm1cA/Ptpn11E76K mice express the driver mutations. However, not all leukemia populations are expected to have the same capacity to engraft into primary recipients. Given the relative stability of the immune diversity of the Npm1cA/Ptpn11E76K leukemic cells, we wanted to determine whether lineage–Sca-1+cKit+ (LSK) cells, which previous studies in mice have identified as containing leukemia-initiating cells (29), were capable of generating diversity of immune subsets identified in the transgenic mice. Therefore, LSKs were sorted from the spleens of individual donors (n = 3) and injected via the tail vein into NOD-Prkdcem26Cd52Il2rgem26Cd22/NjuCrl (NCG) recipients (n = 3; sorting strategy in Supplemental Figure 7). There was considerable variability in the engraftment of the LSK cells into NCG recipients, with donors producing a lethal leukemia in 1 of 3, 2 of 3, and 3 of 3 recipients (Figure 6D). We again performed immune profiling of the splenocytes from recipient mice that developed a lethal leukemia, and the immune subset diversity in LSK-engrafted mice was comparable to bulk-engrafted recipients (Figure 6E). This suggests that the diverse mature myeloid populations in Npm1cA/Ptpn11E76K AML are derived from a stem or progenitor population within the LSK cells.
CD11c+ Npm1cA/Ptpn11E76K splenocytes form a lethal leukemia in primary recipients. We observed that the rate of engraftment failure in recipients of sorted LSK cells was higher than when bulk splenocytes were engrafted. We therefore hypothesized that a population expressing a lineage marker may also contribute to engraftment. To test this, we sorted 3 populations: (a) total CD45+, (b) CD11c+ cells (DC lineage) and (c) CD11c–CD11b+ (monocyte/macrophage lineage) from Npm1cA/Ptpn11E76K splenocytes (n = 3; sorting strategy in Supplemental Figure 8). The frequency of donor-derived (CD45.2+) engraftment in NCG recipients shows that both the CD45+ and CD11c+ populations engrafted at an equivalent rate, while the CD11c–CD11b+ population exhibited highly sporadic and short-lived engraftment (Figure 6F). Both the CD45+ and CD11c+ populations, but not CD11c–CD11b+ cells, exhibited increased white blood cell (WBC) counts (Figure 6G), leading to the formation of a lethal leukemia (Figure 6H). Importantly, the survival in mice engrafted with the CD11c+ population was similar to NCG mice engrafted with total bulk spleen cells (as in Figure 6A).
FLT3 ligand differentiates Npm1cA/Ptpn11E76K bone marrow cells into monocytes and DCs that have an altered cytokine response to CpG. Given the expansion of a DC population in these mice, we wanted to determine whether the leukemic cells isolated from Npm1cA/Ptpn11E76K mice can respond to antigen by stimulating Npm1cA/Ptpn11E76K bone marrow cells with CpG. We first differentiated the cells for 9 days in vitro with FLT3 ligand to ensure that there was a sufficient population of DCs, particularly in the Mx-Cre mice where these cells are more limited. The Mx-Cre control mice developed mainly mature pDCs and cDC1s, with a smaller proportion of cDC2s, immature pDCs, immature DCs, and very few monocytes. The Npm1cA bone marrow cells developed into mainly mature pDCs, while the Ptpn11E76K bone marrow cells differentiated into mainly cDC1s. Finally, the Npm1cA/Ptpn11E76K bone marrow cells differentiated into monocytes, followed by immature pDCs and immature DCs (Figure 7A). After overnight stimulation with CpG, the Npm1cA/Ptpn11E76K cells produced an equal amount of IFN-α compared to Mx-Cre controls but less IFN-γ and TNF-α (Figure 7, B–D). Compared with Mx-Cre, Npm1cA, and Ptpn11E76K controls, the Npm1cA/Ptpn11E76K cells produced more immunosuppressive IL-10 after CpG stimulation (Figure 7E). These results suggest that the bone marrow cells from Npm1cA/Ptpn11E76K mice can be differentiated into monocytes and immature DCs with a potentially immunosuppressive phenotype.
 Figure 7
Figure 7Murine Npm1cA/Ptpn11E76K bone marrow DC differentiation and in vitro CpG stimulation. (A) Immune subsets of bone marrow cells from Mx-Cre (n = 3), Npm1cA (n = 3), Ptpn11E76K (n = 3), and Npm1cA/Ptpn11E76K (n = 3) mice after 9 days of differentiation with FLT3 ligand in vitro. (B) IFN-α (pg/mL), (C) IFN-γ (pg/mL), (D) TNF-α (pg/mL), and (E) IL-10 (pg/mL) in the supernatant of DC-differentiated Mx-Cre (n = 3), Npm1cA (n = 3), Ptpn11E76K (n = 3), and Npm1cA/Ptpn11E76K (n = 3) bone marrow cells after overnight stimulation with 0.1 μg/μL of CpG. Data are presented as mean ± SD. Statistical analysis by 1-way ANOVA with Benjamini-Hochberg FDR correction applied. *FDR-adjusted P ≤ 0.05, **FDR-adjusted P ≤ 0.01, ***FDR-adjusted P ≤ 0.001, ****FDR-adjusted P ≤ 0.0001.
-
Discussion
Using scDNA-seq of AML patient samples, we present data that demonstrate for the first time to our knowledge that PTPN11 can act as an early driver mutation in AML when accompanied by a strong oncogene such as NPM1 mutation or rearrangement of EVI1 or KMT2A. The co-occurring single-cell surface immunophenotype showed that most myeloid cells, independent of immature or mature myeloid marker expression, had leukemic mutations. These data highlight that there are few WT myeloid cells irrespective of differentiation state at diagnosis in PTPN11-mutated AML. We further characterized the immunophenotype of leukemic myeloid cells with spectral flow cytometry and identified variably differentiated monocytes and DCs. We then developed a transgenic mouse model concurrently expressing the Ptpn11E76K and Npm1cA alleles to perform validation studies and further characterize the role of a strong driver gene (mutant NPM1) together with a PTPN11 mutation in driving propagation of disease. Mice harboring both Ptpn11E76K and Npm1cA mutations rapidly developed a fully penetrant lethal myeloid neoplasm that recapitulates many of the features of PTPN11-mutated AML in humans. Primary transgenic Npm1cA/Ptpn11E76K mice present with an expansion of variably differentiated myeloid cells, including monocytes and DCs. This diverse immunophenotype was stable upon engraftment, and all immune populations could be derived from the LSK cells. When differentiated in culture with FLT3 ligand, the leukemic cells from the Npm1cA/Ptpn11E76K mice could be pushed toward a DC lineage that exhibits an altered cytokine response to CpG oligodeoxynucleotide stimulation, with increased IL-10 release suggestive of an immunosuppressive phenotype.
Prior work from our group showing a high VAF of PTPN11 mutations in some patients with AML suggested that PTPN11 mutations may be an early co-event in leukemogenesis. The findings of our scDNA-seq analysis corroborate this, confirming that PTPN11 can be both a late acquired subclonal mutation or an early acquired driving mutation together with another strong oncogene (EVI1, KMT2A, or NPM1 mutation). However, a few limitations of the scDNA-seq technology utilized hindered our ability to fully understand the clonal evolution in these patients. First, we are unable to integrate information about chromosomal rearrangements with genomic mutations at a single-cell level. This was particularly pertinent in the PTPN11-mutated patients without a concurrent NPM1 mutation, who frequently carried chromosome rearrangements but had few co-occurring mutations based on the bulk sequencing analysis. In these patients, we could not infer the order of acquisition of the PTPN11 mutation relative to the chromosome rearrangement. Secondly, due to the limited coverage of the scDNA-seq panel, multiple high-VAF mutations identified in the bulk sequencing were not captured in the single-cell data. Therefore, we can only infer the order of acquisition for the captured mutations. Conversely, the scDNA-seq reliably captured some low-frequency mutations not identified in the bulk sequencing, including NRAS, KRAS, and WT1 mutations. Lastly, while single-cell sequencing allows us to look at an individual cell, there may be small clones missed due to cellular capture and allelic dropout limitations of the MissionBio Tapestri platform. We manually analyzed each sample to mitigate these limitations. Despite these limitations, our data support the notion that a PTPN11 mutation can occur early in AML development.
The Npm1cA/Ptpn11E76K mouse model described herein consistently develops an AML phenotype, as evidenced by accumulation of myeloid cells in the peripheral blood and progressive anemia, thrombocytopenia, and leukocytosis at time of death. The Npm1cA/Ptpn11E76K model has a median survival of approximately 4 months from induction, which is shorter than the Npm1cA/Dnmt3AR878H model (30), longer than the Npm1cA/Flt3ITD model, and comparable to the Npm1cA/NrasG12D model (31). In addition to similar disease kinetics, the Npm1cA/Ptpn11E76K and Npm1cA/NrasG12D models also share a propensity for an AML with maturation immunophenotype. One limitation of the Npm1cA/Ptpn11E76K mouse model is that it is only able to inform about the biology of early PTPN11 mutations and not subclonal/late PTPN11 mutations. Comprehensive immune profiling of splenocytes showed considerable diversity in the immunophenotype of Npm1cA/Ptpn11E76K mice. PTPN11 mutations have been previously associated with a monocytic phenotype (21), and we indeed identified a significant expansion of a population of mature and immature monocytic cells. However, we also identified expansion of cells with a mature immunophenotype, including an expansion of cDC2s and mature pDCs as well as partially differentiated DCs and pDCs. Furthermore, we found that when engrafting bulk splenocytes, mature myeloid populations were consistently present in both immunodeficient primary and secondary recipients and importantly could be derived from the LSK stem/progenitor population. Interestingly, engraftment of LSK cells was more inconsistent than unselected splenocytes from the same donors, suggesting contribution of the more mature myeloid cell populations in efficient pathogenesis of Ptpn11E76K/Npm1cA AML. Finally, we found that isolated CD11c+ cells were capable of engraftment and the establishment of a lethal leukemia with equivalent efficiency to a total CD45+ sorted population. This supports the hypothesis that in this mouse model, leukemia-initiating cells are not exclusively found within the LSK population. While the canonical immunophenotypic definition of hematopoietic stem cells excludes CD11c-expressing cells, it has been suggested that stem cells may express CD11c (32). Our immunophenotyping data also show that multiple AML populations express CD11c, including those that express the additional markers that define DCs but also a population of cells lacking MHCII that we identified as immature DCs. Further work is needed to delineate the specific CD11c+ population responsible for engraftment and how the mature immune populations that comprise the AML contribute to the engraftment of LSK cells.
When comparing the populations of myeloid cells across patient samples and the murine model, we were able to identify similar populations of mature and immature monocytes, cDCs, and immature DCs that expressed CD11c but not terminally differentiated DC markers. While monocytes were not significantly upregulated in our patient samples, the scDNA-seq data indicate that these are leukemic monocytes. One major difference is that we were not able to identify a population of neutrophils in the patient samples, which is likely related to a difference in cell processing as the human blood and bone marrow samples were mononuclear cells isolated by density gradient centrifugation, whereas the mouse cells were not. Finally, we looked for the presence of pDCs in the human samples, which were present in a couple of samples but were not above the 2%-of-cells threshold to be defined as pDC AML (33). The pDC expansion in the murine model is likely related to a combination of Mx-Cre driving the expression of mutated Ptpn11 and Npm1 in all hematopoietic stem cells and to the fact that primary AML samples tend to have mutations in more genes than PTPN11 and NPM1. These additional mutations could interfere with an expansion of pDCs.
While CD11c has been recognized as being aberrantly expressed on AML cells for many years (34, 35), DC AML is a very rare entity (36). However, it has been noted that patients with internal tandem duplication of FLT3 (FLT3-ITD) have elevated numbers of cDCs and pDCs. Interestingly, Rickmann et al. noted that the intensity of expression of HLA-DR was decreased in FLT3-ITD–positive samples compared with FLT3-ITD–negative samples (37). This same group published a follow-up paper demonstrating a higher frequency of precursor DCs, defined as lineage–HLA-DR+CD11c+CD123+, and a decreased frequency of terminally differentiated DCs, defined as CD1c+, CD141+, or CD303+ DCs (38). These results are similar to ours, as we detected an expansion of immature DCs that are CD11c+HLA-DR+ but lack CD1c and CD141 expression. However, none of the patients in our cohort had an alteration in FLT3. To our knowledge, we are the first to describe an expansion of variably matured DCs in PTPN11-mutated AML and can confirm using scDNA-seq that these cells have leukemic mutations. While CpG stimulation in vitro resulted in altered IFN production in both murine and primary PTPN11- and NPM1-mutated AML, more work is needed to understand the full functionality of these cells and how they are contributing to leukemogenesis and treatment resistance.
In conclusion, we have demonstrated with both human and murine genetic studies that mutated PTPN11 can have a divergent clonal hierarchy and can be a co-initiating factor with other strong oncogenes. Our murine model of Npm1- and Ptpn11-mutated AML can serve to study the role of DC expansion in both disease progression and immunosuppression associated with this disease. Previous work from Loberg et al. (30) has shown how order of acquisition of DNMT3A and NPM1 mutations can impact disease trajectory. Our work provides the foundation for future studies regarding how mutated NPM1 and SHP2 proteins interact to drive the disease.
-
Methods
Sex as a biological variable. Our study examined male and female primary samples and animals, and sex-dimorphic effects are reported.
Patient samples. scDNA-seq was performed on samples from 13 newly diagnosed patients from the Alliance for Clinical Trials in Oncology with de novo AML and PTPN11 mutations. Five samples were peripheral blood and 8 were bone marrow samples. All patients had a PTPN11 mutation as a dominant mutation (VAF > 0.3), with 9 patients having a co-occurring mutation in the NPM1 gene. Flow cytometry for immunophenotyping was performed on 15 primary AML patient samples, of which 5 had PTPN11-only mutations (5 peripheral blood and 2 bone marrow samples) and 10 had PTPN11 and NPM1 mutations (10 peripheral blood and 6 bone marrow samples). Five healthy donor peripheral blood and 3 bone marrow samples were used as controls for the flow cytometry.
Sequencing reagents. The TotalSeq-D Heme Oncology Panel of oligo-conjugated reagents was purchased from BioLegend (clone information can be found in Supplemental Methods). Primers for the Myeloid Clonal Evolution Panel were purchased from MissionBio (22).
scDNA and protein library preparation and sequencing. Cryopreserved samples were thawed and counted using the Cellaca PLX Image Cytometry System (Nexcelom). A Dead Cell Removal Kit (Miltenyi Biotec) was then used to collect live cells. Cells were recounted and resuspended in Cell Staining Buffer (MissionBio). The remaining staining, cell encapsulation, barcoding, target PCR amplification, PCR cleanup, and quantification steps were performed according to MissionBio’s Tapestri Single-Cell DNA + Protein Sequencing Protocol. Libraries were quantified using a TapeStation (Agilent) prior to pooling. Four pooled samples were run on one S1 lane on an Illumina NovaSeq at the Cincinnati Children’s Hospital Medical Center DNA Sequencing and Genotyping Core.
Data processing. FASTQ files were analyzed through the Tapestri Pipeline, and the subsequent output was analyzed in R as previously described (22). Briefly, reads were aligned to the hg19 genome build (39, 40) and to cell barcodes. Genotypes were called using the GATKv3.7 (41–43). Any intronic changes were removed from the analysis and only nonsynonymous variants were included. The aligned reads of all called variants were visually inspected in the Integrative Genomics Viewer (https://www.broadinstitute.org/scientific-community/software/integrative-genomics-viewer). All variants with VAFs of less than 0.01 and genotyped in less than 50% of cells were excluded from the analysis. Clones were manually examined to exclude those that have high allele dropout rates, low genotype quality, low read depth, exhibit loss of heterozygosity, or improper amplification of mutant alleles. Further filtering and mutational order were determined by current biological knowledge of clonal evolution. For each variant, the number of cells in each genotype was counted and VAF was calculated as VAF = 2n(Het) + 2n(Homo)/2[2n(Het) + 2n(Homo) + 2n(WT)], where n = number of cells identified with relevant mutation. Cells with missing genotyping calls were excluded. A comparison of the bulk and single-cell sequencing VAF is presented in Supplemental Table 1. For the protein analysis, normalization was performed using DSB normalization (44). Dimension reduction, differential expression markers, and data visualization was performed using Seurat in R, as previously described (22, 45). To generate unsupervised heatmaps of protein expression in R, the ComplexHeatmap package from Bioconductor was used (46, 47).
Primary AML sample flow cytometry. For the primary AML immunophenotyping flow cytometry, cryopreserved peripheral blood and bone marrow cells were thawed and 1 × 106 cells were obtained for staining. Cells were first stained with a Live/Dead Fixable Blue Dead Cell Stain Kit (Thermo Fisher Scientific) for 30 minutes. Cells were then washed and blocked with BD Pharmingen Human BD Fc Block (BD Biosciences) and True-Stain Monocyte Blocker (BioLegend) for 5 minutes. Antibody master mix (Supplemental Methods) with Brilliant Stain Buffer Plus (BD Biosciences) in FACS buffer (PBS, 2% FBS, and 0.1% sodium azide) was then added and cells were incubated for 30 minutes. Cells were then washed 3 times and flow cytometry data were immediately acquired on a Cytek Aurora (Cytek Biosciences) with 100,000 events collected per sample.
Experimental animals. Ptpn11E76K and Npm1cA mice were gifts from Cheng-Kui Qu (18) at Emory University (Atlanta, Georgia, USA) and George Vassiliou (27) from Cambridge Stem Cell Institute (Cambridge, United Kingdom), respectively. B6.Cg-Tg(Mx1-cre)1Cgn/J mice (referred to as Mx-Cre mice; strain 003556) (28) were purchased from The Jackson Laboratory. NOD-Prkdcem26Cd52Il2rgem26Cd22/NjuCrl (referred to as NCG mice; strain 572) were purchased from Charles River Laboratories. To induce expression of the transgenes, mice were injected 5 times (once every other day) with 20 μg/g of poly(I:C) (Sigma Aldrich) via intraperitoneal injection at 6–10 weeks old. Mice were bled biweekly to monitor disease development via complete blood counts (CBCs) obtained on the Element HT5 (Heska) and signs of early removal criteria (ERC). ERC was defined as 20% weight loss, partial or full hind limb paralysis, dehydration, anorexia, anemia, hunched posture, inactivity, lethargy, difficulty breathing, or rough hair coat.
Murine cell isolation. When a mouse met ERC, spleen, femurs, and tibias were collected in PBS and immediately processed. Spleens were dissociated in a gentleMACS Dissociator (Miltenyi Biotec) and bone marrow cells were isolated via centrifugation. Both splenocytes and bone marrow cells underwent red blood cell (RBC) lysis with 1× RBC Lysis buffer (Thermo Fisher Scientific). Samples were then sterilely cryopreserved.
Murine peripheral blood and splenocyte flow cytometry. For the peripheral blood flow cytometry, 20 μL of blood was blocked with TrueStain FcX (anti–mouse CD16/32) (BioLegend) and True-Stain Monocyte Blocker (BioLegend) for 5 minutes prior to staining with the 20-antibody cocktail (Supplemental Methods) in Brilliant Stain Buffer Plus (BD Biosciences) and FACS buffer. After 30 minutes of incubation, samples were lysed twice with 1× RBC Lysis Buffer. Fifty thousand events per sample were immediately collected on the Cytek Aurora.
For the splenocyte immunophenotyping flow cytometry, cryopreserved splenocytes were thawed and 1 × 106 were obtained for staining. Cells were first stained with the Live/Dead Fixable Blue Dead Cell Stain Kit for 30 minutes. Cells were then washed and blocked with TrueStain FcX (anti-mouse CD16/32) and True-Stain Monocyte Blocker for 5 minutes. Antibody master mix for the splenocyte panel (Supplemental Methods) with Brilliant Stain Buffer Plus in FACS buffer was then added and cells were incubated for 30 minutes. Cells were then washed 3 times prior to paraformaldehyde fixation for 20 minutes. Cells were then washed and stored at 4°C overnight. Flow cytometry data were acquired the next day on the Cytek Aurora, with 100,000 events per sample being collected.
In vivo bulk splenocyte, LSK, and lineage+ engraftment. One million bulk splenocytes from Ptpn11E76K/Npm1cA mouse spleens were injected via the tail vein into NCG mice. Splenocytes from these primary recipients were collected as described above and 1 × 106 cells were subsequently transplanted into a secondary NCG recipient. For the LSK and lineage+ engraftment, splenocytes were sorted on the BigFoot Spectral Cell Sorter (Thermo Fisher Scientific), and LSKs or lineage+ populations (total CD45+, CD11c+ [CD45+CD3–CD19–Nk1.1–Ly6G–CD11c+] or CD11c–CD11b+ [CD45+CD3–CD19–Nk1.1–Ly6G–CD11c–CD11b+]) were collected for engraftment (Supplemental Methods and Supplemental Figures 7 and 8). Five thousand to 30,000 LSKs were engrafted per NCG mouse via tail vein injection. For each lineage+ donor, between 1 and 6 NCG recipients were engrafted per population with 3 × 105 sorted cells. Disease development was monitored through weekly bleeds via CBCs and flow cytometry for CD45.2 (clone 104; BD Biosciences, 561875) and CD45.1 (clone A20; BD Biosciences, 553775). At ERC, splenocytes and bone marrow cells were cryopreserved as described above.
In vitro murine bone marrow DC differentiation. Cryopreserved bone marrow samples from age-matched Mx-Cre, Npm1cA, Ptpn11E76K, and Npm1cA/Ptpn11E76k mice were thawed and cultured for 9 days in RPMI 1640 supplemented with 10% fetal calf serum (Gibco), 1 mM sodium pyruvate (Gibco), 1× MEM non-essential amino acids (Gibco), 10 mM HEPES (Gibco), 1× GlutaMAX (Gibco), 55 μM 2-mercaptoethanol (Gibco), 1× penicillin-streptomycin (Gibco), and 100 ng/mL FLT3 ligand (Peprotech). Cells were then collected either for flow cytometry or CpG stimulation. Cells collected for flow cytometry were stained with the Live/Dead Fixable Blue Dead Cell Stain Kit for 30 minutes. Cells were then washed and blocked with TrueStain FcX (anti–mouse CD16/32) and True-Stain Monocyte Blocker for 5 minutes. Antibody master mix (Supplemental Methods) with Brilliant Stain Buffer Plus in FACS buffer was then added and cells were incubated for 30 minutes. Cells were washed 3 times with FACS buffer and flow cytometry data were acquired immediately on the Cytek Aurora.
In vitro murine bone marrow CpG stimulation. After 9 days of growth in vitro as described above, bone marrow cells from age-matched Mx-Cre, Npm1cA, Ptpn11E76K, and Npm1cA/Ptpn11E76k mice were resuspended at 1 × 106 cells/mL and cultured overnight with 0.1 μg/μL of CpG (ODN 1585, Invivogen). Supernatant was collected and stored at –80°C. Supernatant was then thawed and cytokines were analyzed using the Mouse Anti-Virus Response LEGENDplex Panel (BioLegend). Samples were run on the Cytek Auroa, and 4,000 events were collected per sample.
Statistics. Data are presented as mean ± SD, unless otherwise specified. One-way ANOVA for log(%) was used to compare the frequency of various cell populations from the broad immunophenotyping panels for both primary AML samples and murine splenocytes. One-way ANOVA for log(weight) or log(count) using contrast for each pair of groups was used to determine differences in spleen weight, WBC, RBC, and platelet (PLT) counts at ERC. One-way ANOVA for log(concentration) using contrast for each pair of groups was used for cytokine production analysis. The Shapiro-Wilk method was used for normality. For multiple-comparisons, the Benjamini-Hochberg method was used to adjust P values. To analyze differences in overall survival time, we fit a Cox’s proportional hazards model for group and used contrast to compare each pair of groups. A P value of less than 0.05 was used to determine significance.
Study approval. All patients provided written informed consent prior to participation in the following protocols for the collection of pretreatment AML peripheral blood and bone marrow samples: CALGB 8461 (cytogenetic studies only, no tissue obtained), CALGB 9665 (leukemia tissue bank, blood and bone marrow obtained for this), and CALGB 20202 (patient consent to perform molecular studies). All protocols were approved by the Institutional Review Board at each center. For the healthy donor bone marrow samples, all donors provided written informed consent prior to participation, and protocols were approved by the IRB at the University of Cincinnati (UC). Healthy donor peripheral blood samples were purchased from the Hoxworth Blood Center (UC). All animal experiments were conducted after approval by UC and The Ohio State University (OSU) Institutional Animal Care and Use Committees (IACUC). Both male and female mice were used. No work was done related to this paper at the University of Pittsburgh (JCB moved here after acceptance of this paper).
Data availability. All supporting data including means and SD reported in the text are available in the Supporting Data Values file which can be found in the Supplemental Material. Raw sequencing data are available in NCBI’s SRA (SUB15641400). Individuals desiring data related to this work can correspond with one of the corresponding authors.
-
Author contributions
SF and CS have equally contributed to the design and execution of the presented data and preparation and revision of the manuscript and therefore were designated as co-first authors. SF, CS, EH, and JCB conceived and designed the study. SF, CS, KB, CC, CF, ML, KF, BW, JRL, and TMS performed experiments. SF, CS, KQ, LAM, and RLB analyzed the single cell sequencing data. SF and CS analyzed all non-sequencing data. CP provided experimental methods for dendritic cell differentiation. NZ helped with immunophenotype definitions. KM, DN, and AKE provided patient samples and corresponding demographics, cytogenetics, and outcomes. JP and SNR performed the statistics. WMY and CKQ provided the Ptpn11E76K mouse. SF, CS, EH, and JCB wrote the original draft of manuscript. MEJ and JPP contributed to experimental design. AC, RS, ESW, JK, and BP were involved in the Alliance studies that collected the primary samples used in this manuscript. All authors reviewed and edited the manuscript. JCB and EH obtained funding for this study. EH and JCB supervised this study.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- NIH grants F30CA265281 (to SF), R35CA197734 and UG1 CA233338 (to JCB).
- Department of Defense grants HT9425-24-1-0821 (to EH).
- Pelotonia graduate fellowship (to SF).
- National Cancer Institute grant R00CA252005 (to LAM).
- American Society of Hematology Junior Faculty Scholar award (to LAM).
- NIH grant P30CA016058 (to The Ohio State University Comprehensive Cancer Center Leukemia Tissue Bank Shared Resource).
-
Acknowledgments
We would like to acknowledge Christopher Manring, Jean Truxall, and Shelley Orwick at The Ohio State Comprehensive Cancer Center. All flow cytometry–assisted sorting was performed with the help of and using equipment maintained by the Research Flow Cytometry Facility in the Division of Rheumatology and the sequencing was performed by the Genomic Sequencing Facility both at Cincinnati Children’s Hospital Medical Center.
Address correspondence to: John C. Byrd, UPMC Hillman Cancer Center, Associate Vice Chancellor for Cancer Affairs, University of Pittsburgh, UPMC Cancer Pavilion, Suite 500, 5150 Centre Avenue, Pittsburgh, Pennsylvania, 15232, USA. Phone: 412.623.3205; Email: byrdjc@upmc.edu. Or to: Erin Hertlein, Division of Hematology-Oncology, University of Cincinnati College of Medicine, 231 Albert Sabin Way, ML 0551, Room 7009, Cincinnati, Ohio, 45267, USA. Phone: 513.558.0858; Email: hertleek@ucmail.uc.edu.
-
Footnotes
JCB’s present address is: UPMC Hillman Cancer Center and Division of Malignant Hematology and Medical Oncology, Department of Medicine, University of Pittsburgh, Pittsburgh, Pennsylvania, USA.
Conflict of interest: LAM and RLB had previously received honoraria for speaking arrangements and had previously served on a Speakers Bureau for Mission Bio, Inc.
Copyright: © 2026, Fobare et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(4):e193779.https://doi.org/10.1172/jci.insight.193779.
-
References
- Kishtagari A, Levine RL. The role of somatic mutations in acute myeloid leukemia pathogenesis. Cold Spring Harb Perspect Med. 2021;11(4):a034975.
- Weeks LD, Ebert BL. Causes and consequences of clonal hematopoiesis. Blood. 2023;142(26):2235–2246.
- Welch JS, et al. The origin and evolution of mutations in acute myeloid leukemia. Cell. 2012;150(2):264–278.
- Döhner H, et al. Diagnosis and management of AML in adults: 2022 recommendations from an international expert panel on behalf of the ELN. Blood. 2022;140(12):1345–1377.
- Smith CC, et al. Molecular profile of FLT3-mutated relapsed/refractory patients with AML in the phase 3 ADMIRAL study of gilteritinib. Blood Adv. 2022;6(7):2144–2155.
- Wang F, et al. Leukemia stemness and co-occurring mutations drive resistance to IDH inhibitors in acute myeloid leukemia. Nat Commun. 2021;12(1):2607.
- Cai SF, et al. A study to assess the efficacy of enasidenib and risk-adapted addition of azacitidine in newly diagnosed IDH2-mutant AML. Blood Adv. 2024;8(2):429–440.
- McMahon CM, et al. Clonal selection with RAS pathway activation mediates secondary clinical resistance to selective FLT3 inhibition in acute myeloid leukemia. Cancer Discov. 2019;9(8):1050–1063.
- Choe S, et al. Molecular mechanisms mediating relapse following ivosidenib monotherapy in IDH1-mutant relapsed or refractory AML. Blood Adv. 2020;4(9):1894–1905.
- Lee D, et al. PTPN11 mutations confer adverse outcomes and therapy resistance in older patients with acute myeloid leukemia (AML). Blood. 2022;140(suppl 1):3208–3209.View this article via: CrossRef Google Scholar
- Chyla B, et al. Genetic biomarkers of sensitivity and resistance to venetoclax monotherapy in patients with relapsed acute myeloid leukemia. Am J Hematol. 2018;93(8):E202–E205.
- Barford D, Neel BG. Revealing mechanisms for SH2 domain mediated regulation of the protein tyrosine phosphatase SHP-2. Structure. 1998;6(3):249–254.
- Chen L, et al. Mutated Ptpn11 alters leukemic stem cell frequency and reduces the sensitivity of acute myeloid leukemia cells to Mcl1 inhibition. Leukemia. 2015;29(6):1290–1300.
- Nabinger SC, et al. The protein tyrosine phosphatase, Shp2, positively contributes to FLT3-ITD-induced hematopoietic progenitor hyperproliferation and malignant disease in vivo. Leukemia. 2013;27(2):398–408.
- Patsoukis N, et al. Interaction of SHP-2 SH2 domains with PD-1 ITSM induces PD-1 dimerization and SHP-2 activation. Commun Biol. 2020;3(1):128.
- Strullu M, et al. Juvenile myelomonocytic leukaemia and Noonan syndrome. J Med Genet. 2014;51(10):689–697.
- Kratz CP, et al. The mutational spectrum of PTPN11 in juvenile myelomonocytic leukemia and Noonan syndrome/myeloproliferative disease. Blood. 2005;106(6):2183–2185.
- Xu D, et al. Non-lineage/stage-restricted effects of a gain-of-function mutation in tyrosine phosphatase Ptpn11 (Shp2) on malignant transformation of hematopoietic cells. J Exp Med. 2011;208(10):1977–1988.
- Fobare S, et al. Molecular, clinical, and prognostic implications of PTPN11 mutations in acute myeloid leukemia. Blood Adv. 2022;6(5):1371–1380.
- Stasik S, et al. Impact of PTPN11 mutations on clinical outcome analyzed in 1529 patients with acute myeloid leukemia. Blood Adv. 2021;5(17):3279–3289.
- Alfayez M, et al. The clinical impact of PTPN11 mutations in adults with acute myeloid leukemia. Leukemia. 2021;35(3):691–700.
- Miles LA, et al. Single-cell mutation analysis of clonal evolution in myeloid malignancies. Nature. 2020;587(7834):477–482.
- Morita K, et al. Clonal evolution of acute myeloid leukemia revealed by high-throughput single-cell genomics. Nat Commun. 2020;11(1):5327.
- Schwede M, et al. Mutation order in acute myeloid leukemia identifies uncommon patterns of evolution and illuminates phenotypic heterogeneity. Leukemia. 2024;38(7):1501–1510.
- Lapidot T, et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature. 1994;367(6464):645–648.
- Alcalay M, et al. Acute myeloid leukemia bearing cytoplasmic nucleophosmin (NPMc+ AML) shows a distinct gene expression profile characterized by up-regulation of genes involved in stem-cell maintenance. Blood. 2005;106(3):899–902.
- Vassiliou GS, et al. Mutant nucleophosmin and cooperating pathways drive leukemia initiation and progression in mice. Nat Genet. 2011;43(5):470–475.
- Kühn R, et al. Inducible gene targeting in mice. Science. 1995;269(5229):1427–1429.
- Krivtsov AV, et al. Cell of origin determines clinically relevant subtypes of MLL-rearranged AML. Leukemia. 2013;27(4):852–860.
- Loberg MA, et al. Sequentially inducible mouse models reveal that Npm1 mutation causes malignant transformation of Dnmt3a-mutant clonal hematopoiesis. Leukemia. 2019;33(7):1635–1649.
- Dovey OM, et al. Molecular synergy underlies the co-occurrence patterns and phenotype of NPM1-mutant acute myeloid leukemia. Blood. 2017;130(17):1911–1922.
- Hou L, et al. CD11c regulates hematopoietic stem and progenitor cells under stress. Blood Adv. 2020;4(24):6086–6097.
- Xiao W, et al. Plasmacytoid dendritic cell expansion defines a distinct subset of RUNX1-mutated acute myeloid leukemia. Blood. 2021;137(10):1377–1391.
- Scott CS, et al. Flow cytometric analysis of membrane CD11b, CD11c and CD14 expression in acute myeloid leukaemia: relationships with monocytic subtypes and the concept of relative antigen expression. Eur J Haematol. 1990;44(1):24–29.
- Bahia DM, et al. Aberrant phenotypes in acute myeloid leukemia: a high frequency and its clinical significance. Haematologica. 2001;86(8):801–806.View this article via: PubMed Google Scholar
- Ferran M, et al. Acute myeloid dendritic cell leukaemia with specific cutaneous involvement: a diagnostic challenge. Br J Dermatol. 2008;158(5):1129–1133.
- Rickmann M, et al. Elevated frequencies of leukemic myeloid and plasmacytoid dendritic cells in acute myeloid leukemia with the FLT3 internal tandem duplication. Ann Hematol. 2011;90(9):1047–1058.
- Rickmann M, et al. Monitoring dendritic cell and cytokine biomarkers during remission prior to relapse in patients with FLT3-ITD acute myeloid leukemia. Ann Hematol. 2013;92(8):1079–1090.
- Langmead B, et al. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10(3):R25.
- Kim D, et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14(4):R36.
- McKenna A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20(9):1297–1303.
- Depristo MA, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43(5):491–498.
- Van der Auwera GA, et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics. 2013;43(1110):11.10.1–11.10.33.View this article via: PubMed Google Scholar
- Mulè MP, et al. Normalizing and denoising protein expression data from droplet-based single cell profiling. Nat Commun. 2022;13(1):2099.
- Hao Y, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184(13):3573–3587.
- Gu Z. Complex heatmap visualization. Imeta. 2022;1(3):e43.
- Gu Z, et al. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32(18):2847–2849.
-
Version history
- Version 1 (February 23, 2026): Electronic publication












