Research ArticleCell biologyDermatology Free access | 10.1172/jci.insight.193017
USP16 drives psoriasis progression by deubiquitinating and stabilizing NLRP3 in keratinocytes
Nan Wang,1 Fangqian Guan,1 Yifan Lin,1 Bohao Sun,2 Jindan Dai,1 Xiejun Xu,1,3 Weibo Tang,1,4 Yanhua Ren,5 Xuliang Huang,6 Wenjie Gao,1 Xixi Chen,1,7 Litai Jin,1,8 Weitao Cong,1 and Zhongxin Zhu1
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Wang, N. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Guan, F. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Lin, Y. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Sun, B. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Dai, J. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Xu, X. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Tang, W. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Ren, Y. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Huang, X. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Gao, W. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Chen, X. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Jin, L. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Cong, W. in: PubMed | Google Scholar
1School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China.
2Department of Pathology, the Second Affiliated Hospital, School of Medicine, Zhejiang University, Hangzhou, China.
3Shusheng Key Laboratory of Medical Chemical Pharmaceutical Research, Ningbo Second Hormone Factory, Ningbo, China.
4Laboratory of Tumor Targeted Therapy and Translational Medicine, Jilin Medical University, Jilin, China.
5School of Basic Medical Sciences, Wenzhou Medical University, Wenzhou, China.
6Department of Anaesthesia, the First Affiliated Hospital of Wenzhou Medical University, Wenzhou, China.
7Department of Pharmacy, Taizhou Central Hospital, Taizhou, Zhejiang, China.
8Ningbo Key Laboratory of Skin Science, Ningbo College of Health Sciences, Ningbo, China.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
Authorship note: NW and FG contributed equally to this work.
Find articles by Zhu, Z. in: PubMed | Google Scholar
Authorship note: NW and FG contributed equally to this work.
Published January 27, 2026 - More info
JCI Insight. 2026;11(5):e193017. https://doi.org/10.1172/jci.insight.193017.
© 2026 The American Society for Clinical Investigation
Received: March 3, 2025; Accepted: January 14, 2026
-
Abstract
Psoriasis is a chronic inflammatory dermatosis characterized by pathological keratinocyte hyperproliferation and dysregulated immune activation. While ubiquitin-specific peptidase 16 (USP16) has been implicated in modulating multiple cellular signaling pathways, its functional role in psoriatic pathogenesis remains poorly understood. Our investigation revealed pronounced upregulation of USP16 expression in psoriatic epidermis compared with normal controls. Keratinocyte-specific USP16 knockdown demonstrated remarkable therapeutic efficacy, significantly ameliorating characteristic psoriatic phenotypes including epidermal hyperplasia and inflammatory infiltration. RNA-seq analysis showed that USP16 has substantial effects on cell cycle transition and keratinocytes proliferation. Through KEGG analysis, it was found that USP16 primarily regulates the NLRP3 signaling pathway, leading to enhanced cell proliferation and inflammation. Mechanically, USP16 directly binds to the NLRP3 protein to eliminate K48 ubiquitination modification, enhancing the stability of the NLRP3 protein, activating inflammasome activity. Further studies showed that the therapeutic effects of reducing USP16 on psoriasis progression were counteracted by an NLRP3 activator and keratinocyte-specific NLRP3 overexpression adenovirus. Collectively, these results shed light on how USP16 promotes NLRP3 signaling in keratinocytes, exacerbating psoriasis development. This positive regulation highlights the potential of USP16 as a therapeutic target for psoriasis.
-
Introduction
Psoriasis is a prevalent chronic inflammatory skin condition distinguished by the aberrant proliferation of keratinocytes (KCs) and the infiltration of immune cells (1, 2). The immune system dysfunction, a primary driver of psoriasis, is chiefly initiated by various immune factors and sustained by T cells (3). Nevertheless, the significant contribution of KCs to the pathogenesis of psoriasis is increasingly acknowledged (4). The emergence of targeted drugs aimed at KCs holds promise as a novel treatment approach for psoriasis. Nonetheless, the precise mechanism driving the abnormal behavior of KCs in psoriasis remains incompletely elucidated.
The ubiquitin proteasome system serves as the primary mechanism for protein degradation within eukaryotic cells, facilitating the breakdown of aged or damaged intracellular proteins (5, 6). This process involves the coordinated action of ubiquitin activating enzyme (E1), ubiquitin binding enzyme (E2), and ubiquitin ligase (E3) (7). Ubiquitination is a dynamic and reversible process, wherein deubiquitination can eliminate the ubiquitin chain from a substrate protein, thereby erasing the ubiquitination marker. USP16, a member of the ubiquitin-specific protease family, features a Zn-F domain and 2 UBP domains (8). USP16 catalyzes the removal of ubiquitin from the ubiquitin-ubiquitin junction site on a substrate protein through a cysteine-mediated mechanism at position C205 (9). Numerous studies have indicated that USP16 plays a crucial role in regulating diverse aspects of signaling pathways, cell cycle progression, cell proliferation, and inflammation in a wide array of diseases, such as cancer, infectious diseases, and immune disorders (10–12). However, the function of USP16 in psoriatic KCs has not been clearly defined.
Inflammasomes are key regulators of inflammation, with the NLRP3 inflammasome being extensively studied, especially in the central nervous system (13, 14). This complex comprises the pattern recognition receptor NLRP3, the adaptor protein ASC, and an inactive form of the catalytic enzyme Caspase-1 (15). Upon activation by various signals like oxidative stress, pathogens, and tissue damage products, the NLRP3 inflammasome processes pro–IL-1β into its active form through Caspase-1 (16). NLRP3 serves as a sensor for different danger signals, influencing immune responses and being implicated in diverse inflammatory and autoimmune skin disorders such as dermatophytosis, acne, urticaria, and bullous pemphigoid (17–19). Recent studies also have shown that the abnormal activation of the NLRP3 inflammasome linked to enhanced cellular proliferation and inflammation in psoriasis (20, 21).
This research demonstrates elevated levels of USP16 in psoriatic lesions, imiquimod-induced (IMQ-induced) and IL-23–induced psoriasis mouse models. Furthermore, the deficiency or knockdown of USP16 decreases the aberrant proliferation and inflammation of KCs, leading to the alleviation of psoriasis phenotype. Our mechanistic investigation reveals that USP16 interacts with NLRP3, eliminating the K48 ubiquitination of NLRP3. This action stabilizes NLRP3, ultimately activating the NLRP3 signaling pathway. In essence, our study uncovers a mechanism through which USP16 contributes to psoriasis progression by modulating the NLRP3 signaling pathway.
-
Results
USP16 is upregulated in both psoriatic skin and in IMQ-induced psoriasis mouse models. To explore the function of USP16 in psoriasis, we initially examined publicly accessible transcriptomic data and noted a significant elevation in USP16 gene expression in psoriasis models (GSE14905), compared with the respective controls (Figure 1A). Subsequently, our investigations in psoriasis patients revealed that the expression level of USP16 was notably elevated compared with that in normal skin tissues, as indicated by IHC staining (Figure 1B). Following this, we induced psoriasis in a mouse model using IMQ and observed a significant increase in USP16 expression in the psoriatic epidermis compared with the mice coated with Vaseline determined by IHC staining (Figure 1C). In line with these findings, Western blot analysis of skin tissue revealed a notable upregulation of USP16 expression in mice with IMQ-induced psoriasis compared with mice treated with Vaseline (Figure 1D). To better characterize the cell types that primarily express USP16, we analyzed the recently published single-cell RNA-seq dataset from normal human skin tissue. The results show that USP16 was mainly expressed in KCs (Supplemental Figure 1A; supplemental material available online with this article; https://doi.org/10.1172/jci.insight.193017DS1). To replicate the characteristics of psoriatic KCs in an in vitro setting, we utilized a cytokine cocktail called pso-mix (M5), comprising IL-17A, IL-22, OSM, IL-1α, and TNF-α. This combination created an inflammatory environment that closely resembles the conditions observed in psoriasis (22, 23). In addition, upon treating KCs with M5 at various time points, we observed that the expression of USP16 peaked at 24 hours (Figure 1E and Supplemental Figure 1B). Moreover, after treating KCs with varying concentrations of M5 for 24 hours to induce psoriasis model in vitro, we discovered that the mRNA level of USP16 was most pronounced at 10 ng/mL for 24 hours (Figure 1F). Collectively, these findings indicate that the upregulation of USP16 in lesions is linked to the advancement of psoriasis.
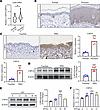 Figure 1
Figure 1USP16 is upregulated in both psoriatic skin and in IMQ-induced psoriasis mouse models. (A) The mRNA level of USP16 in the skin from normal and lesional skin of psoriasis patients was analyzed using the Gene Expression Omnibus (GEO) database. (B) Representative IHC staining for USP16 expression in the skin from healthy Normal and Psoriatic patients (n = 5). Scale bar: 100 μm. (C) IHC staining and quantitative analysis of USP16 in the skin from Vaseline and IMQ-mice (n = 5). Scale bar: 100 μm. (D) Immunoblotting analysis and quantitative analysis of USP16 expression in the skin from Vaseline and IMQ-mice. β-Actin was used as a loading control (n = 6). (E) Immunoblotting analysis and quantitative analysis of USP16 protein level in HaCaT cells stimulates with M5 treatment for 0, 12, 24, and 48 hours at 10 ng/mL dose. β-Actin was used as a loading control (n = 3). (F) qPCR analysis for USP16 mRNA level in HaCaT cells stimulates with M5 for 0, 5, 10, 15 ng/mL with 24 hours treatment. β-Actin was used as a loading control (n = 3). Data are shown as the mean ± SEM. *P < 0.05; **P < 0.01; ***P < 0.001. The P value was determined using unpaired, 2-tailed Student’s t test (A–D) and 1-way ANOVA (E and F). All numbers (n) are biologically independent experiments.
USP16 ablation in KCs ameliorates the psoriasiform phenotype in IMQ-induced psoriasis mouse models. To explore the function of USP16 in psoriasis, we engineered an adeno-associated virus carrying a USP16 plasmid tagged with GFP. This construct was designed to knockdown USP16 specifically in mouse skin KCs by utilizing a KC-specific Krt14 promoter (Supplemental Figure 2, A and B). Through this approach, we aimed to investigate the effect of USP16 in KCs on the development and progression of psoriasis. H&E staining revealed that, under normal conditions, the targeted knockdown of USP16 in KCs did not show a significant effect on the thickness of the skin’s epidermal layer (Supplemental Figure 2C). This indicates that, in the absence of external stimuli or pathological conditions, the depletion of USP16 does not have a significant effect on the normal structural integrity of the skin. However, following the induction of psoriasis using IMQ in mouse models (Supplemental Figure 2D), we observed that the psoriasis area and severity index (PASI) were notably higher in AAV-GFP mice compared with AAV-Cre mice (Figure 2, A and B). Subsequently, we conducted immunoblotting analysis on the back skin tissue of the mice to assess the levels of Cyclin A1, Cyclin E1, and Bcl-2, key regulators of the cell cycle known to have a crucial role in controlling cell proliferation. Our findings reveal a downregulation of these proteins in AAV-Cre mice in comparison with AAV-GFP mice (Figure 2C). H&E images were used to measure epidermal thickness, revealing that the skin thickness of mice with USP16 KC-specific knockdown was significantly reduced compared with the control mice (Figure 2D). Similarly, the expression of proliferation marker Ki-67+ KCs in IMQ-treated AAV-Cre mice was significantly decreased compared with IMQ-treated AAV-GFP mice (Figure 2E). Finally, we examined the expression levels of skin inflammation and hyperplasia markers, such as K6, through immunofluorescence analysis to study the effect of USP16 on IMQ-induced skin inflammation. As anticipated, the expression of K6 was higher in IMQ-treated AAV-GFP mice compared with IMQ-treated AAV-Cre mice (Figure 2F). These discoveries offer valuable insights into the potential involvement of USP16 in regulating KC proliferation and inflammation following IMQ treatment.
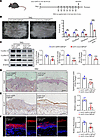 Figure 2
Figure 2USP16 ablation in keratinocytes ameliorates the psoriasiform phenotype. (A) Schematic diagram demonstrating the animal experiment design. (B) Representative images of the dorsal back from mice (left) and mice PASI scores were depicted (right) (n = 4). Image acquired with digital camera. (C) Immunoblotting and quantitative analysis of Cyclin A1, Cyclin E1, and Bcl-2 protein levels in AAV-GFP and AAV-Cre mice were treated with IMQ for 7 days. β-Actin was used as a loading control (n = 5). (D) Representative histological sections of the dorsal back from AAV-GFP and AAV-Cre mice treated by IMQ for 7 days stained with H&E (left), and quantification of the epidermal thickness stained with H&E (n = 5). Scale bars: 200 μm. (E) Representative IHC staining for Ki-67 expression in the skin from AAV-GFP and AAV-Cre mice were treated with IMQ for 7 days (n = 5). Scale bar: 100 μm. (F) Immunofluorescent labeling and quantitative analysis of K6+ in the skin from AAV-GFP and AAV-Cre mice were treated with IMQ for 7 days. Nuclei were stained with DAPI (blue) (n = 5). Scale bar: 50 μm. Data are shown as the mean ± SEM. **P < 0.01; ***P < 0.001. The P value was determined using unpaired, 2-tailed Student’s t test (B–F). All numbers (n) are biologically independent experiments.
USP16 is required for KC proliferation. Given that psoriasis is marked by the abnormal proliferation of KCs, we delved deeper into exploring the potential role of USP16 in modulating the proliferation of KCs. Firstly, we employed siRNA to silence endogenous USP16 in HaCaT cells (Supplemental Figure 3A) and subjected these cells to M5 stimulation and performed RNA-seq analysis. Gene Ontology (GO) enrichment analysis demonstrated that the silencing of USP16 resulted in significant alterations in cell cycle transformation and cell proliferation of KCs. Subsequently, gene set enrichment analysis (GESA) of the data further reveals a strong association between the loss of USP16 and cell cycle transformation. These findings provide evidence for the critical role of USP16 in regulating these processes in KCs (Figure 3, A and B). Based on our findings, we then investigated the effect of USP16 on cell cycle transformation and cell proliferation in KCs. Specifically, we detected the expression levels of key cell cycle–related proteins, such as Cyclin A1, Cyclin E1, and the cell survival–promoting protein Bcl-2, and the results show that USP16 knockdown treatment significantly reduced the expression levels of these proteins (Figure 3C). These results were confirmed in normal human epidermal keratinocytes (Supplemental Figure 4A). Moreover, to evaluate the effect of USP16 knockdown on cell proliferation, we conducted EdU and Ki-67 staining to assess the presence of actively proliferating cells. Subsequent to M5 treatment, we observed a significant decrease in the number of EdU+ and Ki-67+ cells in the si-USP16 treatment group compared with the si-Scr treatment group (Figure 3, D and E). In order to bolster our understanding of the role of USP16 in facilitating cell cycle progression, we conducted flow cytometry analysis to examine cell cycle distribution. Our results reveal that USP16 knockdown resulted in cell cycle arrest at the G0/G1 phase, whereas a greater percentage of cells in the si-Scr treatment group were found in the G2/M phase (Figure 3F). These outcomes collectively indicate that the interference of USP16 not only influences cell proliferation but also disrupts the G1/S cell cycle transition in KCs.
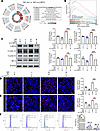 Figure 3
Figure 3USP16 is required for keratinocyte proliferation. (A) RNA-seq analysis of the differentially regulated genes between si-Scr group and si-USP16 group treated with M5 for 24 hours. Differentially regulated genes were enriched in GO biological process terms. (B) GSEA results for cell cycle transition. (C) Immunoblotting and quantitative analysis of USP16, Cyclin A1, Cyclin E1, and Bcl-2 protein levels in HaCaT cells that were treated with si-Scr or si-USP16 and stimulated with or without M5 for 24 hours. β-Actin was used as a loading control (n = 3). (D) Immunofluorescence and quantitative analysis of Ki-67+ in HaCaT cells that were treated with si-Scr or si-USP16 and stimulated with or without M5 for 24 hours. Nuclei were stained with DAPI (blue) (n = 3). Scale bar: 50 μm. (E) Immunofluorescent and quantitative analysis of EdU+ in HaCaT cells that were treated with si-Scr or si-USP16 and stimulated with or without M5 for 24 hours. Nuclei were stained with Hoechst (blue) (n = 3). Scale bar: 50 μm. (F) Flow cytometric plots of cell-cycle analysis performed with PI staining on HaCaT cells treated with si-Scr or si-USP16 and stimulated with or without M5 for 24 hours (left). Quantification the percentage of cells that fall into the sub G0/G1, S, or G2/M gates (right). Data are shown as the mean ± SEM. *P < 0.05; **P < 0.01; ***P < 0.001. The P value was determined using 1-way ANOVA (C–F). All numbers (n) are biologically independent experiments.
USP16 targets and regulates NLRP3 signaling in KCs. To explore the molecular mechanism behind USP16’s effect on KC proliferation and inflammation, we conducted RNA-seq analysis to identify genes involved in the altered signaling pathways (Figure 4A). Among these DEGs, the heatmap and KEGG analysis show that the NOD-like signaling pathway was downregulated upon USP16 silencing, suggesting that USP16 promotes KC proliferation and inflammation by activating this pathway (Figure 4, B and C). GO enrichment analysis indicated that the downregulated DEGs were mainly enriched in biological processes associated with signal transduction (Figure 4D). Nod-like receptor pyrin domain-containing protein 3 (NLRP3), a key member of the NOD-like receptor family, is typically triggered in response to infection or inflammation, leading to the formation of the NLRP3 inflammasome. Western blotting analysis confirmed that, despite the stimulation of KCs by M5, the activity of the NLRP3 signaling pathway was decreased upon silencing of USP16. This was evident from the reduced protein expression levels of NLRP3 and the secretion of cytokines, including cleaved-caspase-1, IL-18, and IL-1β (Figure 4E). Meanwhile, these results were confirmed in NHEKs (Supplemental Figure 5A). In addition, immunofluorescence analysis revealed that the downregulation of USP16 in KCs led to decreased expression levels of NLRP3 and C-Caspase-1 in the cytoplasm (Supplemental Figure 6, A and B).
 Figure 4
Figure 4USP16 targets and regulates NLRP3 signaling in keratinocytes. (A and B) Volcano map and heatmap analysis for the significantly differentially regulated gene by the interference of si-Scr or si-USP16 in HaCaT cells treated by M5. (C) The Top 15 significant KEGG signaling pathways are listed. (D) The downregulated DEGs were classified based on their gene ontology (GO) terms for biological process (BP), cellular component (CC), and molecular function (MF). (E) Immunoblotting and quantitative analysis of NLRP3, C-Caspase-1, IL-18, and IL-1β protein levels in HaCaT cells that were treated with si-Scr or si-USP16 and stimulated with or without M5 for 24 hours. β-Actin was used as a loading control (n = 3). Data are shown as the mean ± SEM. *P < 0.05; ***P < 0.001. The P value was determined using 1-way ANOVA (E). All numbers (n) are biologically independent experiments.
Furthermore, our in vivo experiments provided evidence that the activity of the NLRP3 signaling pathway was significantly diminished in USP16 KC-specific knockdown mice compared with the control group by Western blot assay (Supplemental Figure 7A). These findings were further supported by the immunofluorescence assay, which demonstrated a substantial decrease in NLRP3 expression within USP16-deficient KCs (Supplemental Figure 7B). In conclusion, our results reveal a close correlation between USP16 and the activity of the NLRP3 signaling pathway in the context of psoriasis.
USP16 interacted and stabilized NLRP3 by inhibiting its K48 ubiquitination. In the next step, we investigated the effect of USP16 on NLRP3 expression. The overexpression of USP16 notably elevated the levels of NLRP3 protein level in HEK293T cells (Figure 5A). Interestingly, silencing USP16 did not result in any changes in NLRP3 mRNA levels (Figure 5B). These findings indicate that USP16 modulates NLRP3 protein expression at the posttranslational level. Subsequently, we investigated whether USP16 influences the stability of NLRP3. A cycloheximide (CHX) chase assay revealed that the upregulation of USP16 delayed the degradation of NLRP3 expression in HEK293T cells (Figure 5C). Meanwhile, MG132 (proteasome inhibitor) had the ability to reverse the downregulation of NLRP3 caused by USP16 knockdown, while CQ (autophagic lysosome inhibitor) did not show any effect on this process (Figure 5D). Following that, we conducted immunofluorescence experiments to investigate the potential binding between USP16 and NLRP3. The results indicate a binding interaction between these 2 proteins (Figure 5E). Comparable results were also noted through immunoprecipitation experiments conducted on KCs under physiological conditions (Figure 5F).
 Figure 5
Figure 5USP16 interacted and stabilized NLRP3 by inhibiting its K48 ubiquitination. (A) Immunoblotting and quantitative analysis of NLRP3 protein level that Flag-USP16 or Vector were transfected in HaCaT cells. β-Actin was used as a loading control (n = 4). (B) qPCR analysis for NLRP3 mRNA level that si-Scr or si-USP16 were transfected in HaCaT cells treated with M5 for 24 hours (n = 3). (C) Immunoblotting and quantitative analysis of NLRP3 protein levels that Flag-USP16 or Vector were transfected in HaCaT cells treated with cyclohexamide (CHX) for 0 and 4 hours. β-Actin was used as a loading control (n = 3). (D) Immunoblotting analysis of NLRP3 protein level that si-Scr or si-USP16 were transfected in HaCaT cells treated with MG132 or CQ for 6 hours and stimulated M5 for 24 hours. β-Actin was used as a loading control (n = 3). (E) Immunofluorescent staining of USP16 and NLRP3 in HaCaT cells. Nuclei were stained with DAPI (blue). Scale bar: 10 μm. (F) The HaCaT cell lysates were immunoprecipitated with IgG and anti-USP16 antibody and the expression of USP16 and NLRP3 were detected by Western blot. (G) Molecular docking analysis of USP16 binding to NLRP3 (confidence score: 0.8593). (H) Coimmunoprecipitation experiment was carried out to determine the specific binding domain between Flag-USP16 and GST-NLRP3. (I) USP16 amino acid fragment diagram. (J) HEK293T cells were transfected with HA-NLRP3 and several Flag-USP16 mutants. The whole-cell lysates were immunoprecipitated with anti-HA beads and immunoblotted with anti-Flag and anti-HA antibodies. (K) The HEK293T cells were cotransfected with Flag-USP16 and HA-Ub plasmids. The cell lysates were immunoprecipitated by anti-NLRP3 antibody, and then Western blot assay with anti-HA, anti-NLRP3, and anti-Flag antibody. (L) The HEK293T cells were cotransfected with Flag-USP16 and HA-Ub-K48 plasmids. The cell lysates were immunoprecipitated by anti-NLRP3 antibody, and then Western blot assay with anti-HA, anti-NLRP3, and anti-Flag antibody. Data are shown as the mean ± SEM. *P < 0.05; ***P < 0.001. The P value was determined using unpaired, 2-tailed Student’s t test (A–C). All numbers (n) are biologically independent experiments.
We attempted to explore the molecular mechanism of the USP16-NLRP3 interaction using docking software. The results indicate that USP16 may bind to SER-855 of NLRP3 by hydrogen bonding (Figure 5G). The GST-pulldown assay yielded the same result (Figure 5H). USP16 comprises a peptidase domain and a zinc finger ubiquitin binding domain (Znf-Ubp). In order to pinpoint the critical domains responsible for the interactions between NLRP3 and USP16, we also generated various truncated forms of USP16 that comprised a peptidase domain (amino acids 1–142) and a zinc finger ubiquitin binding domain (ZnF-Ubp, amino acids 184–C-terminus). (11). The results of coimmunoprecipitation assays in HEK293T cells demonstrate that the interaction between NLRP3 and USP16 was specifically mediated by the peptidase domain of USP16 (Figure 5, I and J). Given that USP16 functions as a deubiquitinating enzyme, we proceeded to investigate its effect on the ubiquitination of NLRP3. Our findings reveal that the overexpression of USP16 led to a decrease in NLRP3 polyubiquitination in HEK293T cells (Figure 5K). Furthermore, our study indicates that the overexpression of USP16 has the capacity to eliminate the K48 ubiquitin chain modification on NLRP3 (Figure 5L). Although prior studies indicate that USP16 deubiquitinates IKKβ to activate NLRP3 (8, 24), our experiments show that NLRP3 expression remained unchanged following IKKβ knockdown (Supplemental Figure 8, A and B). This finding demonstrates that USP16 regulated NLRP3 independently of IKKβ. In summary, our findings demonstrate that USP16 interacts with NLRP3 and eliminates the K48 ubiquitin chain, thereby stabilizing its protein expression.
The psoriasis alleviation driven by USP16 knockdown depends on NLRP3. To explore the effect of NLRP3 on the alleviation of psoriatic KCs through USP16 knockdown, we employed an NLRP3 activator, BMS-986299, to induce the activation of the NLRP3 signaling pathway (25). In our in vitro experiments, Western blot analysis revealed that the NLRP3 activator notably counteracted the inhibitory effect of USP16 knockdown on cell proliferation markers, including Cyclin A1, Cyclin E1, and Bcl-2 (Supplemental Figure 9A). Additionally, immunofluorescence experiments confirmed that the NLRP3 activator significantly increased the percentage of Ki-67+ and EdU+ cells when USP16 expression was inhibited (Supplemental Figure 9, B and C). Moreover, our cell cycle analysis conducted through flow cytometry demonstrated that the NLRP3 activator effectively reversed the cell cycle inhibitory effects caused by interfering with USP16 (Supplemental Figure 9D). We also performed Western blot analysis in NHEKs to validate and confirm the previously observed results (Supplemental Figure 10A).
Following this, we conducted the corresponding experimental validation in mice (Figure 6A). As shown in the Figure 6B, the PASI demonstrated that the diminishing effect resulting from USP16 knockdown is reversed by the administration of the NLRP3 activator. This indicates that the NLRP3 activator counteracts the psoriasis alleviation brought about by USP16 knockdown. Furthermore, Western blot analysis indicated a reduction in the expression levels of Cyclin A1, Cyclin E1, and Bcl-2 following USP16 deletion, with partial restoration observed upon activation of the NLRP3 signaling pathway (Figure 6C). H&E staining demonstrated that the targeted knockdown of USP16 in KCs led to a decrease in epidermal thickness. Nonetheless, these effects were reversed upon the administration of BMS-986299 (Figure 6D). Similar findings were observed through IHC staining analysis of Ki-67 (Figure 6E). Immunofluorescence staining of K6 revealed that mice with USP16 deficiency exhibited diminished inflammation and decreased proliferation of KCs, with these outcomes predominantly reversed by the NLRP3 activator (Figure 6F). Moreover, we observed that the inflammatory cytokines IL-1β and IL-6 were restored in response to the NLRP3 activator, despite the knockdown of USP16 in KCs (Supplemental Figure 9E). To eliminate confounding effects from other cell types, we verified our findings using a KC-specific NLRP3 overexpression adenovirus. Subsequent analysis revealed that NLRP3 overexpression abrogated the therapeutic effect of USP16 knockdown on the psoriatic phenotype. This was jointly demonstrated by a return to a hyperproliferative state, as indicated by elevated levels of Cyclin A1, Cyclin E1, and Bcl-2, along with histological evidence from H&E staining, Ki-67 histochemistry, and K6 immunofluorescence (Supplemental Figure 11, A–F). These data establish that the alleviation of the psoriasis phenotype by USP16 knockdown is functionally dependent on the downregulation of NLRP3. These findings suggest that the NLRP3 signaling pathway plays a crucial role in mitigating the psoriasis phenotype following the deletion of USP16.
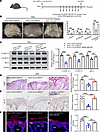 Figure 6
Figure 6The psoriasis alleviation driven by USP16 knockdown depends on NLRP3. (A) Schematic diagram demonstrating the animal experiment design. (B) Representative images of the dorsal back from mice (left) and mice PASI scores were depicted (right) (n = 4). (C) Immunoblotting and quantitative analysis of Cyclin A1, Cyclin E1, and Bcl-2 protein levels in the skin from AAV-GFP, AAV-Cre, and AAV-Cre-BMS-986299 mice treated by IMQ for 7 days. β-Actin was used as a loading control (n = 5). (D) Representative histological sections of the dorsal back from AAV-GFP, AAV-Cre and AAV-Cre-BMS-986299 mice treated by IMQ for 7 days stained with H&E, and quantification of the epidermal thickness and the infiltrating cells stained with H&E (n = 5). Scale bars: 200 μm. (E) Representative IHC staining for Ki-67 expression in the skin from AAV-GFP, AAV-Cre, and AAV-Cre-BMS-986299 mice stimulated by IMQ for 7 days (n = 5). Scale bars: 100 μm. (F) Immunofluorescence and quantitative analysis of K6+ in the skin from AAV-GFP, AAV-Cre, and AAV-Cre-BMS-986299 mice induced by IMQ for 7 days. Nuclei were stained with DAPI (blue) (n = 5). Scale bars: 50 μm. Data are shown as the mean ± SEM. *P < 0.05; **P < 0.01; ***P < 0.001. The P value was determined using 1-way ANOVA (B–F). All numbers (n) are biologically independent experiments.
KC specific knockdown of USP16 alleviates IL-23–induced psoriasis phenotype. In addition, the use of intradermal administration of IL-23 in mice offers another valuable approach to studying psoriasis (26, 27). This model specifically targets the clinically relevant IL-23/IL-17A signaling axis and effectively mimics various disease features observed in human psoriasis. Subsequently, we aim to investigate the potential role of USP16 in a mouse model of IL-23–induced psoriasis (Figure 7A). The increased expression of USP16 in the epidermal layer of the skin in IL-23–induced psoriasis models (Figure 7, B and C), as well as the beneficial effects observed upon KC-specific knockdown of USP16, highlight its importance in the pathogenesis of psoriasis. The downregulation of key cell proliferating proteins and the reduction in epidermal thickness, coupled with the decreased expression of Ki-67, strongly support the notion that USP16 is involved in the dysregulated proliferation and pathology of psoriatic KCs (Figure 7, D–F). The same regulatory effects of USP16 on NLRP3 signaling pathway was also verified in the IL-23–induced psoriasis mouse model (Supplemental Figure 12, A and B).
 Figure 7
Figure 7Keratinocyte-specific knockdown of USP16 alleviates IL-23–induced psoriasis phenotype. (A) Schematic diagram demonstrating the animal experiment design. (B) Immunoblotting and quantitative analysis of USP16 protein levels in PBS and IL-23–treated mice. β-Actin was used as a loading control (n = 5). (C) Representative IHC staining for USP16 expression in PBS and IL-23–induced psoriasis lesions (n = 5). Scale bars: 100 μm. (D) Immunoblotting and quantitative analysis of Cyclin A1, Cyclin E1, and Bcl-2 protein levels in the skin from AAV-GFP and AAV-Cre mice were treated with IL-23. β-Actin was used as a loading control (n = 5). (E) Representative histological sections of the dorsal back from AAV-GFP and AAV-Cre mice were treated with IL-23 and stained with H&E, and quantification of the epidermal thickness was analyzed (n = 5). Scale bars: 200 μm. (F) Representative IHC staining for Ki-67 expression in the skin from AAV-GFP and AAV-Cre mice treated with IL-23 (n = 5). Scale bar: 100 μm. Data are shown as the mean ± SEM. **P < 0.01, ***P < 0.001. The P value was determined using unpaired, 2-tailed Student’s t test (B–F). All numbers (n) are biologically independent experiments.
To further investigate the in vivo relationship between USP16 and NLRP3, we conducted additional validation using an IL-23–induced psoriasis mouse model. Specifically, the downregulation of proliferative protein expression, the reduction in epidermal thickness, and the decrease in the positive rate of Ki-67, which were observed upon USP16 knockdown, were all partially reversed by the administration of an NLRP3 activator (Supplemental Figure 13, A–C). Indeed, these observations provide compelling evidence for the significant role of USP16 in psoriatic KCs.
The expression of USP16 in psoriasis has been found to not only correlate with NLRP3 levels but also with the progression of the disease. To explore the potential clinical correlation between USP16 and NLRP3, we further assessed the expression of both proteins in 60 samples obtained from diverse psoriasis patients. IHC analysis revealed a positive correlation between the levels of USP16 and NLRP3 expression (Figure 8A). Immunostaining quantification and statistical analyses revealed the correlations to be significant (Figure 8B; r = 0.82, P < 0.001).
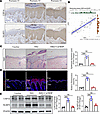 Figure 8
Figure 8The expression of USP16 in psoriasis has been found to not only correlate with NLRP3 levels but also with the progression of the disease. (A) IHC staining detected the expression of USP16 and NLRP3 in 60 psoriasis specimens. Images shown are 3 representative specimens. The inserts are high magnification images (n = 60). Scale bars: 100 μm. (B) Correlation of USP16 and NLRP3 expressions was significant (Pearson correlation test; r = 0.82, P < 0.001). (C) Representative histological sections of the dorsal skin from Vaseline, IMQ, and IMQ+Cal/BDP-treated mice stained with H&E, and quantification of the epidermal thickness (n = 5). Scale bars: 200 μm. (D) Immunofluorescent and quantitative analysis of USP16 in the dorsal skin from Vaseline, IMQ, IMQ+Cal/BDP-treated mice. Nuclei were stained with DAPI (blue) (n = 5). Scale bars: 100 μm. (E) Immunoblotting and quantitative analysis of USP16 in the dorsal skin from Vaseline, IMQ, IMQ+Cal/BDP-treated mice (n = 5). Data are shown as the mean ± SEM. ***P < 0.001. The P value was determined using 1-way ANOVA (C–E). All numbers (n) are biologically independent experiments.
Currently, several clinical drugs are available for the management of psoriasis, such as Calcipotriol and Betamethasone dipropionate gel (Cal/BDP), which has been widely used in clinical practice (28, 29). In our study, we administered Cal/BDP to IMQ-induced psoriasis mice for 6 consecutive days. As a result, we observed a remarkable alleviation of psoriasis phenotype, with the mice skin returning to a normal state (Supplemental Figure 14, A and B). Subsequently, we investigated the expression of USP16 and observed that its levels correlated with the progression of psoriasis phenotype (Figure 8, C–E). Successful treatment of psoriasis corresponded with a reduction in USP16 expression, indicating its involvement in the pathogenesis of psoriasis. These findings shed light on the dynamic changes in USP16 expression during the course of psoriasis and emphasize its potential as a target for therapeutic interventions.
-
Discussion
Psoriasis is a chronic skin condition marked by imbalanced innate and adaptive immune reactions, engaging various cell types including KCs, dendritic cells, and T cells (30). Epidermal hyperplasia, a hallmark of psoriasis, arises from the dysregulated proliferation and inflammation of KCs, the predominant cell type in the epidermis, playing a pivotal role in the pathogenesis of psoriasis (31).
The purpose of this study was to investigate the unusual expression of USP16 in psoriatic KCs and its potential effect on treating psoriasis. The targeted knockdown of USP16 specifically in KCs in vivo resulted in reduced psoriasis phenotype. Mechanistically, lowering USP16 expression in KCs hinders cell cycle progression and suppresses cell proliferation and inflammation. Our results are significant, as they demonstrate that USP16 modulates the activity of NLRP3 signaling, which plays a role in promoting the development of psoriasis. Furthermore, we observed that USP16 interacts with NLRP3, leading to the removal of K48 link ubiquitination from NLRP3 and thereby stabilizing it. Finally, we confirmed the effect of NLRP3 on USP16 in vitro and vivo using an NLRP3 activator and KC-specific NLRP3 overexpression adenovirus. These findings indicate that USP16 is crucial in the abnormal proliferation and inflammation of KCs in psoriasis and could be a promising target for treating the disease.
USP16, a critical member of the ubiquitin-specific protease family, governs essential biological functions such as gene transcription, cell proliferation, differentiation, cell cycle regulation, and stem cell self-renewal (32). Notably, research indicates that histone H2A deubiquitinase USP16 is involved in regulating hematopoietic processes and hematopoietic stem cell function (33). It can also interact with HERC2 to modulate cellular responses to DNA damage, regulate gene expression, and influence lineage commitment of embryonic stem cells (34). In addition, it has been shown to regulate autoimmune immunoinflammation through its substrate IKKβ (8). Furthermore, USP16 has been implicated as a carcinogenic factor in KRAS-driven lung tumorigenesis (35). In our experimental findings, initial screening of extensive clinical sample data uncovered a notable elevation of USP16 levels in psoriatic tissues compared with normal skin tissue. Subsequent targeted knockdown of USP16 in KCs using adeno-associated virus demonstrated a substantial alleviation of psoriasis phenotype following USP16 suppression. Additionally, RNA-seq analysis of KCs with altered USP16 expression unveiled a significant regulatory influence of USP16 on cell cycle progression and cellular proliferation capacity. Follow-up experiments were conducted to validate these observations.
The inflammasome, comprising a collection of cytoplasmic protein complexes, plays a vital role in the innate immune response, aiding the host in defending against pathogens and initiating the inflammatory repair process (36). Found in various cell types including macrophages, dendritic cells, neutrophils, and KCs, NLRP3 has significant implications (37). Studies have shown that ginsenoside Rg1 can ameliorate psoriatic-like skin lesions by inhibiting KC proliferation and the NLRP3 inflammasome (38). Cycloastragenol-mediated inhibition of NLRP3 inflammasome–induced pyroptosis in macrophages has been found to alleviate psoriasis-like skin inflammation triggered by IMQ in mice (39). Furthermore, literature has reported the involvement of NLRP3 in cell proliferation dynamics. Specifically, NLRP3 has been linked to promoting tumor growth and metastasis in human oral squamous cell carcinoma (40), as well as participating in phenotypic alterations and proliferation of hypertensive vascular smooth muscle cells (41). In our experimental findings, upon conducting RNA-seq analysis on KCs with modified USP16 expression, we discovered a significant regulatory role of USP16 in the NLRP3 signaling pathway. Further validation confirmed the positive regulation of NLRP3 by USP16. Additionally, our observations revealed that USP16 can directly modulate NLRP3 by eliminating its K48 ubiquitination modification, thereby enhancing the protein stability of NLRP3. In vitro and vivo rescue experiments further supported the essential role of NLRP3 in USP16-mediated regulation of the psoriasis process.
In summary, our research highlights USP16 as a deubiquitination mediator capable of enhancing the stability of NLRP3 in psoriatic KCs. This mechanism leads to increased cell proliferation and instigation of inflammatory responses, thereby expediting the progression of psoriasis. The elevated levels of USP16 in psoriatic KCs underscore its pivotal role as a significant contributor to psoriasis development, offering a promising new target for potential therapeutic interventions in psoriasis.
-
Methods
Sex as a biological variable. For experiments in animals, only male mice were used based on practical considerations. This approach reduces variability from the female estrous cycle, allowing us to focus resources on delineating the core mechanism.
Human tissue. Human clinical skin tissue samples were obtained from patients with psoriasis and healthy volunteers and matched for sex and age in the Second Affiliated Hospital of Wenzhou Medical University. All the patients were assessed according to the PASI.
Animals. Eight- to 10-week-old male C57BL/6 mice were obtained from Model Animal Research Center of Nanjing University (Nanjing, China).
Adeno-associated virus production. AAV expressing GFP under the control of the skin epidermal KC-specific Krt14 promoter (Genechem, GOSV0207037_1): AAV9-Krt14 GFP (AAV-GFP) or AAV9-Krt14-Cre (AAV-Cre) was injected s.c. using 35-gauge needle around the specific region of dorsal skin. ADV expressing Flag under the control of the skin epidermal KC-specific Krt14 promoter (Genechem, GOSV0207037_1): ADV-Flag or ADV-NLRP3 was injected s.c. using 35-gauge needle around the specific region of dorsal skin. Five injections of approximately 10 μL at a dose of 1.8 × 1013 viral genomes were administered. Three weeks after injection, AAV-mediated gene knockdown was confirmed in skin by immunofluorescence.
Psoriasis mouse models. The psoriasis mouse models used in our research study were IMQ-induced and IL-23–induced psoriasis mouse models.
IMQ-induced psoriasis mouse models. The day before induction, the backs of the mice were shaved and then treated with Aldara cream (Sichuan MingXin Pharmaceutical Co., LTD., H20030129) containing 5% IMQ (55 mg) once daily for 7 days.
IL-23–induced psoriasis mouse model. We performed intradermal injection of 20 μL PBS, either alone or containing 500 ng recombinant mouse IL-23 (eBiosience), into the dorsal skin of anesthetized mice using a 35-gauge needle every other day for 16 days.
The mice treated with BMS-986299 were i.p. injected with BMS-986299 (10 mg/kg) 1 hour before IMQ or IL-23 application.
To measure the severity of inflammation on the back, a scoring system considering skin thickness, scaling, and erythema (not taking into account the area which was determined by the experimenter), similar to the human PASI score was used (cumulative IMQ score) (22).
Cal/BDP treats psoriasis. On the second day of successful IMQ-induced psoriasis in mouse model, Calciumtriol and Betamethasone dipropionate (Cal/BDP) gel was applied to the dorsal skin of mice every day for 6 consecutive days, and the skin of mice returned to normal level.
Reagents. CQ (MCE, HY-17589A), CHX (Selleck, S7418), MG132 (Selleck, 2619), BMS-986299 (MCE, HY-139396), Cal/BDP (LEO Pharma) were used.
Cell culture. The KCs cell line HaCaT was obtained from Procell, (Wuhan, China, RRID: CVCL_0038), while human embryonic kidney cells (HEK293T) were purchased from the ATCC (CRL-1573). The HaCaT cells were cultured in MEM medium (Procell, PM150478) and HEK293T cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco) supplemented with 10% (vol./vol.) fetal bovine serum (FBS; Thermo Fisher Scientific, 10099141), 100 U/mL penicillin G and 0.1 mg/mL streptomycin sulfate (Thermo Fisher Scientific, 15140122). The calcium concentration for cell culture is ~1.80 mmol/L. The cell culture conditions were 37°C and 5% CO2. Primary Normal Human Epidermal KCs (NHEKs; Lifeline Cell Technology cat. FC-0007) were cultured in the serum-free basal medium with growth factors (Lifeline Cell Technology cat. LL-0007). The medium was refreshed every 2 days, and the cells were subcultured according to the cell fusion.
In vitro psoriasis model. M5, a cytokine cocktail containing IL-1α, TNF-α, IL-17A, IL-22, and oncostatin M (PeproTech) was used to induce psoriatic inflammation in KCs at a concentration of 10 ng/mL for 24 hours.
RNA interference. The experimental analysis was carried out after the siRNA transfection. For the negative control, siRNA obtained from Santa Cruz Biotechnology was used. Additionally, USP16 siRNA (Santa Cruz Biotechnology, sc-41687) and IKKβ siRNA (Santa Cruz Biotechnology, sc-35644) were used. The cells were seeded 12 hours prior to transfection and allowed to reach a confluence of 30%–50% by day 0. Then, 30 nM of the siRNA was transfected into the cells using Lipofectamine 3000 (Invitrogen, L3000-001) and Opti-MEM I Reduced Serum Medium (Gibco, 31985088), following the instructions provided by the manufacturers. On day 1, when the cells reached confluence (24 hours after transfection), the siRNA solution was replaced with full growth medium. Subsequently, the experimental analysis was conducted.
Histological analysis. Paraffin-embedded tissue sections were stained with the H&E kit (Solarbio) to evaluate the thickness of the skin epidermis. The sections were observed using a Nikon camera (Nikon).
Immunofluorescence staining assay. For skin sections that were 5 μm thick, the process of deparaffinization and rehydration was conducted. Subsequently, antigen retrieval was performed by heating the slides in 10 mM Citrate buffer (pH 6.0) at 95°C for 10 minutes. As for HaCaT samples, the cells were fixed in 4% paraformaldehyde for 15 minutes and permeabilized with 0.5% Triton X-100 for 15 minutes at room temperature. Following fixation and permeabilization, the samples were blocked with 5% (vol./vol.) bovine serum albumin (BSA, Sigma-Aldrich, B2064) for 1 hour at room temperature. Next, the samples were incubated overnight at 4°C with primary antibodies against Krt14 (Abcam, ab181595; 1:200 dilution), K6 (proteintech, 10590-1-AP; 1:400 dilution), NLRP3 (Abcam, ab283819; 1:50 dilution), C-Caspase-1 (Proteintech, 22915-1-AP; 1:100 dilution), and GFP (Abcam, ab183734; 1:500 dilution) followed by washing and incubation with secondary antibodies. Finally, DAPI was used to label the cell nuclei. Images were acquired using a Leica SP8 confocal microscope (Leica).
IHC staining. For IHC, the following was used: USP16 (1:400, 14055-1-AP, Proteintech), Ki-67 (1:1000, ab15580). Then it was incubated with the secondary antibody for 60 minutes at room temperature. After washing with PBS, staining with diaminobenzidine (Rabbit-specific HRP/DAB Detection IHC Kit, ab64261, Abcam) was performed for 2 minutes. This was followed by counterstaining with hematoxylin, dehydration, and observation under a microscope.
The collection of human psoriasis skin specimens (60 samples) was conducted in accordance with an approved protocol by the IRB at the Second Affiliated Hospital of Wenzhou Medical University. These specimens were obtained and archived in an anonymous manner. In simple terms, percentages are defined as follows: 0 means no cell staining, 1 means 0%–25% cell staining, 2 means 25%–50% cell staining, 3 means 50%–75% cell staining, and 4 means more than 75% cell staining. The staining intensity score is defined as follows: 0 is unstained, 1 is weakly stained, 2 is moderately stained, and 3 is strongly stained. The total score is obtained by multiplying the percentage score by the intensity score. Two people, neither of whom knew about the slide exam, rated each sample. The final score for each sample is the average of its 2 scores.
Western blot. RIPA lysis buffer (Thermo Fisher Scientific) containing Protease and Phosphatase Inhibitor Cocktail (Abcam, ab65621) was utilized to extract total proteins from fresh skin tissues and cell samples. The protein concentrations were determined using the Pierce BCA Protein Assay Kit (Thermo Fisher Scientific, 23228). The proteins were then separated by SDS-PAGE and transferred onto a PVDF membrane (Millipore, IPVH00010). To prevent nonspecific binding, the membrane was blocked with 5% nonfat milk (BD Biosciences, 232100). Subsequently, the membrane was incubated with primary antibodies followed by secondary antibodies. Finally, the blots were developed using the ECL reagent (Millipore, WBLUF0500) and captured using the Amersham Image 680 system (GE Healthcare Life Sciences). The antibodies used in the Western blot analysis are listed in Supplemental Table 1.
RNA isolation and qPCR. Total RNA was extracted from cells using TRIzol Reagent (Invitrogen). Next, total RNA (2 μg) was reverse transcribed into cDNA using GoScript Reverse Transcription Kit (Promega). The cDNA was then subjected to qPCR analysis, and gene expression was quantified as previously described. The mRNA levels of target genes were normalized against that of β-actin. The gene-specific primer sequences used for RT-qPCR are listed in Supplemental Table 2.
EdU-assay. Cell proliferation was detected using a cytochemical method according to the instructions provided by the manufacturer (Beyotime, C0075L). HaCaT cells were incubated with the EdU staining buffer for 2 hours, followed by fixation with 4% paraformaldehyde. The cell nuclei were stained with Hoechst 33342. Subsequently, the stained cells were examined and imaged under a microscope.
Cell cycle flow cytometry. HaCaT cells were seeded in 6-well plates with 1.5 × 105 cells per well. After 24 hours, the cells were treated differently based on the experimental conditions. Cells were collected and then resuspended in 500 μL of cold PBS solution. Subsequently, the cells were fixed with 70% ethanol at –20°C for 2 days. After washing with PBS, the cells were resuspended in a PBS solution containing PI (BD Biosciences, 550825) at a concentration of 20 μg/mL and ribonuclease A at a concentration of 10 μg/mL. The cells were incubated in the dark for 30 minutes. Flow cytometry analysis was performed using a BD FACS Calibur instrument, capturing at least 10,000 events. The data obtained were analyzed using FlowJo software.
Immunoprecipitation. Cells were lysed using ice-cold IP buffer (Beyotime, P10013J) with PMSF. The primary antibody was covalently immobilized on Protein A/G immunoprecipitation magnetic beads following the manufacturer’s instructions (Thermo Fisher Scientific, 26147). The cell lysates were incubated with the immobilized antibody beads for at least 4 hours at 4°C for immunoprecipitation. After immunoprecipitation, the samples were washed 5 times with Tris-buffered saline. Subsequently, the immunoprecipitates were eluted with glycine-HCl (0.1 M, pH 3.5), and the eluted proteins were subjected to immunoblotting using specific primary antibodies.
GST-pulldown assay. Flag-USP16 plasmids were transfected into HEK293T cells, and subsequently, the cells were lysed using IP lysates. Flag-USP16 was then purified using the kit (P9801, Beyotime). GST and GST-NLRP3 were transfected into Expi293 cells and purified with GST beads (P2138, Beyotime) at 4°C. The purified proteins, GST and GST-NLRP3, were incubated with the purified Flag-USP16 proteins overnight at 4°C. Following this, the beads were washed 3 times with IP lysates. The complexes were boiled in 5× SDS loading buffer at 95°C for 10 minutes, after which Western blotting analyses were conducted.
Plasmid transfection. The constucts of USP16WT, USP161-142, USP16184-823 in pEGFP-C1 have been previously described. Briefly, the constructs were generated by PCR amplification and cloned into pEGFP-C1 vector. Plasmid harboring HA-ubiquitin, HA-ubiquitinK48 and HA-NLRP3 were purchased from Limbio Biotechnology. HEK293T cells were seeded into a 6-well plate 12 hours before transfection. After reaching 30%–50% confluence (day 0), 1 μg of the plasmid was transfected using Lipofectamine 3000 and Opti-MEM I Reduced Serum Medium (Gibco) according to the instructions of the manufacturers. On day 1 (24 hours after transfection), when cells reached confluence, the transfection medium was replaced with complete growth medium, and cells were subsequently harvested for analysis.
Ubiquitination assay. HEK293T cells were lysed with IP buffer and subsequently boiled for 10 minutes. For immunoprecipitation, 500 μL of the cell lysate was incubated with anti-NLRP3 antibody (antibody to cell lysate ratio of 1 μg/mg) for 12 hours. Afterward, 30 μL of Protein A/G immunoprecipitation magnetic beads were added and the mixture was incubated at 4°C for 12 hours. The ubiquitination of NLRP3 were then analyzed using anti-Ub antibody Western blotting.
Protein-protein docking studies. Given the protein sequence files provided, we employed the AlphaFold 2 software to model their 3-dimensional structures (42). Subsequently, we utilized the HDOCK software for protein-protein docking, yielding a variety of complex structures (43). These structures were meticulously sorted and screened based on their respective confidence scores. The complex structure with the highest confidence score was identified as the target for further analysis. We then employed PyMOL software to delve into the binding site of the protein-protein complex.
Generally speaking, a confidence score above 0.7 indicates a high likelihood of binding between the 2 molecules; a score between 0.5 and 0.7 suggests a possible binding interaction; and a score below 0.5 implies a low probability of binding.
RNA-seq analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis. RNA-seq was performed at Guangzhou Epibiotek Co. Ltd. Total RNA was isolated using Trizol reagent (Invitrogen). VAHTS Stranded mRNA-seq Library Prep Kit for Illumina V2 (Vazyme Biotech, NR612-02) was used for library preparation according to the instructions. Reads were aligned to the human Ensemble genome GRCh38 using Hisat2 aligner (v2.1.0) under parameters: “--rna-strandness RF”. The reads mapped the genome were calculated using feature Counts (v1.6.3). Differential gene expression analysis was performed using the DESeq2 R-package. KEGG pathway and GO terms enrichment analysis of differentially expressed genes (DEGs) was carried out by cluster Profiler R Bioconductor package with P < 0.05 as a statistically significant cutoff.
GSEA. For the functional enrichment analysis of gene sets, GO annotations of genes in the R software package org. hs. eg. db (version 3.1.0) and KEGG pathway gene annotations obtained from the KEGG rest API were used as background sets. DEGs were then mapped to background sets. The R software package Cluster Profiler (version 3.14.3) was used for the enrichment analysis. Statistical significance was set at P < 0.05.
Statistics. All data were obtained from at least 3 independent experiments and presented as mean ± SEM. Data between 2 groups were determined using the unpaired, 2-tailed Student’s t test, and comparisons among multiple groups were determined using 1-way ANOVA. P < 0.05 was considered statistically significant.
Study approval. Written informed patient consent was obtained from volunteer patients with psoriasis and healthy controls. The study was performed in accordance with the principles of the Declaration of Helsinki and approved by the Second Affiliated Hospital of Wenzhou Medical University. The collection of human psoriasis skin specimens (60 samples) was conducted in accordance with an approved protocol by the IRB at the Second Affiliated Hospital of Wenzhou Medical University.
All animal studies were performed according with the recommendations described in the Guide for the Care and Use of Laboratory Animals (National Academies Press, 2011). Treatment of the animals were approved by the animal welfare committee Wenzhou Medical University. All operations were performed under anesthesia, and all efforts were made to minimize suffering.
Data availability. Raw data have been deposited to National Center for Biotechnology Information (NCBI) under the BioProject no. PRJNA1393932. All underlying source data are available in the Supporting Data Values file.
-
Author contributions
NW, FG, YL, BS, JD, and XC carried out the experiments and analyzed the data. NW, WC, LJ, and ZZ supervised the project and designed the experiments. NW, XX, and WT wrote the manuscript. NW, YR, WG, XH, WC, and ZZ helped with the experiments and revised the manuscript. All authors have read and approved the final manuscript.
-
Funding support
- Natural Science Foundation of Jilin Province (YDZJ202201ZYTS286).
- National Nature Science Foundation of China (82272284, 82273560, 8220069 and 82504271).
-
Acknowledgments
The authors thank Scientific Research Center of Wenzhou Medical University for consultation and instrument availability that supported this work.
Address correspondence to: Zhongxin Zhu, or Weitao Cong, School of Pharmaceutical Science, Wenzhou Medical University, Wenzhou, China. Email: zhongxinzhu2010@163.com (ZZ). Email: cwt97126@126.com (WC).
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2026, Wang et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(5):e193017.https://doi.org/10.1172/jci.insight.193017.
-
References
- Griffiths CEM, et al. Psoriasis. Lancet. 2021;397(10281):1301–1315.
- Yamanaka K, et al. Pathophysiology of psoriasis: a review. J Dermatol. 2021;48(6):722–731.
- Su R, et al. Metabolic influences on T cell in psoriasis: a literature review. Front Immunol. 2023;14:1279846.
- Chen HL, et al. Galectin-7 downregulation in lesional keratinocytes contributes to enhanced IL-17A signaling and skin pathology in psoriasis. J Clin Invest. 2021;131(1):e130740.
- Xiong Y, et al. Ubiquitin-proteasome system-regulated protein degradation in spermatogenesis. Cells. 2022;11(6):1058.
- Schneider SM, et al. Viral entry and the ubiquitin-proteasome system. Cell Microbiol. 2021;23(2):e13276.
- Asano Y, et al. E3-ubiquitin ligases and recent progress in osteoimmunology. Front Immunol. 2023;14:1120710.
- Yu JS, et al. Substrate-specific recognition of IKKs mediated by USP16 facilitates autoimmune inflammation. Sci Adv. 2021;7(3):eabc4009.
- Montellese C, et al. USP16 counteracts mono-ubiquitination of RPS27a and promotes maturation of the 40S ribosomal subunit. Elife. 2020;9:e54435.
- Zhang L, et al. Endothelial DGKG promotes tumor angiogenesis and immune evasion in hepatocellular carcinoma. J Hepatol. 2024;80(1):82–98.
- Zhang Y, et al. USP16-mediated deubiquitination of calcineurin A controls peripheral T cell maintenance. J Clin Invest. 2019;129(7):2856–2871.
- Reinitz F, et al. Inhibiting USP16 rescues stem cell aging and memory in an Alzheimer’s model. Elife. 2022;11:e66037.
- Tastan B, et al. Targeting NLRP3 inflammasome with Nrf2 inducers in central nervous system disorders. Front Immunol. 2022;13:865772.
- Seyedsadr M, et al. IL-11 induces NLRP3 inflammasome activation in monocytes and inflammatory cell migration to the central nervous system. Proc Natl Acad Sci U S A. 2023;120(26):e2221007120.
- Huang Y, et al. NLRP3 inflammasome activation and cell death. Cell Mol Immunol. 2021;18(9):2114–2127.
- Fu J, Wu H. Structural mechanisms of NLRP3 inflammasome assembly and activation. Annu Rev Immunol. 2023;41:301–316.
- Shao BZ, et al. NLRP3 inflammasome and its inhibitors: a review. Front Pharmacol. 2015;6:262.
- Nakamura Y, Kambe N. Linkage of bacterial colonization of skin and the urticaria-like rash of NLRP3-mediated autoinflammatory syndromes through mast cell-derived TNF-α. J Dermatol Sci. 2013;71(2):83–88.
- Fang H, et al. Increased expression of NLRP3 inflammasome components and interleukin-18 in patients with bullous pemphigoid. J Dermatol Sci. 2016;83(2):116–123.
- Wang H, et al. NLRP3 promotes tumor growth and metastasis in human oral squamous cell carcinoma. BMC Cancer. 2018;18(1):500.
- Sun Z, et al. Celastrol attenuates ox-LDL-induced mesangial cell proliferation via suppressing NLRP3 inflammasome activation. Cell Death Discov. 2019;5:114.
- Wang Z, et al. Autophagy-based unconventional secretion of HMGB1 by keratinocytes plays a pivotal role in psoriatic skin inflammation. Autophagy. 2021;17(2):529–552.
- Liu M, et al. FOXE1 contributes to the development of psoriasis by regulating WNT5A. J Invest Dermatol. 2023;143(12):2366–2377.
- Schmacke NA, et al. IKKβ primes inflammasome formation by recruiting NLRP3 to the trans-Golgi network. Immunity. 2022;55(12):2271–2284.
- Chen X, et al. HET0016 inhibits neuronal pyroptosis in the immature brain post-TBI via the p38 MAPK signaling pathway. Neuropharmacology. 2023;239:109687.
- Lipovsky A, et al. ACT1 is required for murine IL-23–Induced psoriasiform inflammation potentially independent of E3 ligase activity. J Invest Dermatol. 2021;141(7):1772–1779.
- Lian N, et al. Gasdermin D-mediated keratinocyte pyroptosis as a key step in psoriasis pathogenesis. Cell Death Dis. 2023;14(9):595.
- Lovato P, et al. Calcipotriol and betamethasone dipropionate exert additive inhibitory effects on the cytokine expression of inflammatory dendritic cell-Th17 cell axis in psoriasis. J Dermatol Sci. 2016;81(3):153–164.
- Satake K, et al. Calcipotriol and betamethasone dipropionate synergistically enhances the balance between regulatory and proinflammatory T cells in a murine psoriasis model. Sci Rep. 2019;9(1):16322.
- Lowes MA, et al. Immunology of psoriasis. Annu Rev Immunol. 2014;32:227–255.
- Yang W, et al. FXYD3 enhances IL-17A signaling to promote psoriasis by competitively binding TRAF3 in keratinocytes. Cell Mol Immunol. 2023;20(3):292–304.
- Zheng J, et al. The pleiotropic ubiquitin-specific peptidase 16 and its many substrates. Cells. 2023;12(6):886.
- Gu Y, et al. The histone H2A deubiquitinase Usp16 regulates hematopoiesis and hematopoietic stem cell function. Proc Natl Acad Sci U S A. 2016;113(1):E51–E60.
- Zhang Z, et al. The histone H2A deubiquitinase USP16 interacts with HERC2 and fine-tunes cellular response to DNA damage. J Biol Chem. 2014;289(47):32883–32894.
- Xu G, et al. The deubiquitinase USP16 functions as an oncogenic factor in K-RAS-driven lung tumorigenesis. Oncogene. 2021;40(36):5482–5494.
- Mangan MSJ, et al. Targeting the NLRP3 inflammasome in inflammatory diseases. Nat Rev Drug Discov. 2018;17(8):588–606.
- Honda TSB, et al. Cell type-specific roles of NLRP3, inflammasome-dependent and -independent, in host defense, sterile necroinflammation, tissue repair, and fibrosis. Front Immunol. 2023;14:1214289.
- Mao J, et al. Ginsenoside Rg1 ameliorates psoriasis-like skin lesions by suppressing proliferation and NLRP3 inflammasomes in keratinocytes. J Food Biochem. 2022;46(5):e14053.
- Deng G, et al. Inhibition of NLRP3 inflammasome–mediated pyroptosis in macrophage by cycloastragenol contributes to amelioration of imiquimod-induced psoriasis-like skin inflammation in mice. Int Immunopharmacol. 2019;74:105682.
- Scuderi SA, et al. NLRP3 inflammasome inhibitor BAY-117082 reduces oral squamous cell carcinoma progression. Int J Mol Sci. 2021;22(20):11108.
- Burger F, et al. NLRP3 inflammasome activation controls vascular smooth muscle cells phenotypic switch in atherosclerosis. Int J Mol Sci. 2021;23(1):340.
- Jumper J, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596(7873):583–589.
- Yang Z, et al. AlphaFold2 and its applications in the fields of biology and medicine. Signal Transduct Target Ther. 2023;8(1):115.
-
Version history
- Version 1 (January 27, 2026): In-Press Preview
- Version 2 (March 9, 2026): Electronic publication












