Research ArticleImmunology Free access | 10.1172/jci.insight.123568
Functional heterogeneity of human tissue-resident memory T cells based on dye efflux capacities
Brahma V. Kumar,1 Radomir Kratchmarov,2 Michelle Miron,1,2 Dustin J. Carpenter,1,3 Takashi Senda,1 Harvey Lerner,4 Amy Friedman,4 Steven L. Reiner,2,5 and Donna L. Farber1,2,3
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Kumar, B. in: PubMed | Google Scholar
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Kratchmarov, R. in: PubMed | Google Scholar
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Miron, M. in: PubMed | Google Scholar
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Carpenter, D. in: PubMed | Google Scholar
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Senda, T. in: PubMed | Google Scholar
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Lerner, H. in: PubMed | Google Scholar
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Friedman, A. in: PubMed | Google Scholar
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Reiner, S. in: PubMed | Google Scholar
1Columbia Center for Translational Immunology,
2Department of Microbiology and Immunology, and
3Department of Surgery, Columbia University Medical Center, New York, New York, USA.
4LiveOnNY, New York, New York, USA.
5Department of Pediatrics, Columbia University Medical Center, New York, New York, USA.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
Authorship note: BVK and RK contributed equally to this work.
Find articles by Farber, D. in: PubMed | Google Scholar
Authorship note: BVK and RK contributed equally to this work.
Published November 15, 2018 - More info
JCI Insight. 2018;3(22):e123568. https://doi.org/10.1172/jci.insight.123568.
Copyright © 2018, American Society for Clinical Investigation
Received: July 13, 2018; Accepted: October 16, 2018
-
Abstract
Tissue-resident memory T cells (TRMs) accelerate pathogen clearance through rapid and enhanced functional responses in situ. TRMs are prevalent in diverse anatomic sites throughout the human lifespan, yet their phenotypic and functional diversity has not been fully described. Here, we identify subpopulations of human TRMs based on the ability to efflux fluorescent dyes [efflux(+) TRMs] located within mucosal and lymphoid sites with distinct transcriptional profiles, turnover, and functional capacities. Compared with efflux(–) TRMs, efflux(+) TRMs showed transcriptional and phenotypic features of quiescence including reduced turnover, decreased expression of exhaustion markers, and increased proliferative capacity and signaling in response to homeostatic cytokines. Moreover, upon activation, efflux(+) TRMs secreted lower levels of inflammatory cytokines such as IFN-γ and IL-2 and underwent reduced degranulation. Interestingly, analysis of TRM subsets following activation revealed that both efflux(+) and efflux(–) TRMs undergo extensive transcriptional changes following TCR ligation but retain core TRM transcriptional properties including retention markers, suggesting that TRMs carry out effector function in situ. Overall, our results suggest a model for tissue-resident immunity wherein heterogeneous subsets have differential capacities for longevity and effector function.
-
Introduction
Memory CD8+ T cells provide long-lived protection and exist as heterogeneous subsets differing in tissue homing and self-renewal properties (1). Tissue-resident memory T cells (TRMs) are a noncirculating subset maintained in peripheral tissues that mediate optimal in situ protection against invading pathogens (2–4). In mouse models, TRMs are distinguished from circulating memory subsets by expression of the early T cell activation marker CD69, along with the integrin CD103 for CD8+ TRMs (for a review, see ref. 4). Protective TRMs in mice can be generated by diverse viral and bacterial pathogens and following site-directed vaccination (2, 3). However, TRMs can also direct pathogenic immune responses to allergens in the lung (5, 6) and have been implicated in diseases such as psoriasis and vitiligo in skin (7). Given their potential to participate in protective and pathogenic immune responses in tissues, TRMs are an important target for immunomodulation. However, recent studies have demonstrated that TRMs are highly heterogeneous, encompassing multiple unique subsets (8–12). Understanding the role of these different subsets in immune responses is therefore necessary before therapeutic modulation of TRMs can be achieved.
We previously demonstrated through transcriptome profiling that human TRM-phenotype cells share a core signature with key homology with mouse TRMs (13), including expression of specific homing/adhesion molecules (CD49a and CXCR6), negative regulators (PD-1 and CD101), and elevated production of IL-2 and IL-10 compared with circulating effector-memory T (TEM) cells (12). However, our study also revealed phenotypic heterogeneity within human TRMs, as has been found in other studies (9, 10). TRM subsets identified by markers such as CD49a and CD103 have been shown to be transcriptionally and developmentally distinct and can occupy nonoverlapping subanatomic niches (9–11); this suggests that the TRM compartment actually comprises multiple distinct subsets with tissue-retention properties.
Like memory T cells, hematopoietic stem cells (HSCs) exhibit long-term persistence and are largely maintained in tissues — specifically in bone marrow (BM) niches. HSCs are endowed with self-renewal capacities and are resistant to chemotherapeutic agents, associated with an enhanced expression of ATP-binding cassette (ABC) family multidrug transporters, which efflux proteins and small molecules, and maintain cellular homeostasis (14–16). Recently, subsets of memory T cells with the ability to efflux fluorescent dyes have been identified in human tissues including BM (17) and intestines (18). Another recent study identified effluxing TRM populations in human tissue sites and showed in a mouse model of LCMV infection that effluxing TRMs are associated with quiescence (19). Nonetheless, the distribution of effluxing T cells across human tissues, within TRMs, and their functional role in tissue immunity are unclear.
Here, we identify a population of memory CD8+ T cells with the ability to efflux fluorescent dyes [efflux (+)] in multiple lymphoid and nonlymphoid sites. Efflux(+) cells predominate within the TRM compartment, are enriched for TRM core signature markers, and maintain a TRM profile when stimulated, yet represent a functionally and transcriptionally distinct subpopulation. Notably, efflux(+) TRMs exhibit reduced turnover, transcriptional signatures associated with longevity and quiescence, and an increased capacity to proliferate compared with efflux(–) TRMs, which have a higher effector capacity. Together, these results demonstrate that dye efflux identifies a population of TRMs present in multiple tissues with a unique role in the tissue immune response.
-
Results
CD8+ TRMs efflux fluorescent dyes. We investigated the efflux capacity of memory CD8+ T cells in different tissues based on their ability to be labeled with fluorescent mitochondrial dyes, as done previously with HSCs (20). As the predominant phenotype of memory (CD45RO+) CD8+ T cells across human tissues is TEM (CD45RA–CCR7–) (21, 22), our studies are focused on this subset. T cells isolated from healthy human tissues obtained from organ donors (see Methods) were labeled with MitoTracker Green, a fluorescent dye that labels total mitochondrial mass (23), revealing cells with high and low levels of mitochondrial dye (Mitohi and Mitolo) (Figure 1A). Similar fractions of Mitohi and Mitolo cells were observed using either MitoTracker Green or CMXRos, a dye dependent on mitochondrial membrane potential (Figure 1B), suggesting that changes in mitochondrial state are not responsible for the observed staining patterns. To determine whether the Mitolo subset was due to dye efflux, we stained cells in the presence of increasing concentrations of cyclosporine A (CSA), a competitive inhibitor of efflux pumps (24). A single Mitohi population was observed when cells were labeled in the presence of CSA (Figure 1C). Similar results were obtained when using verapamil (Figure 1C), another competitive inhibitor of efflux pumps that has been shown to have no effect on mitochondrial mass (19). Moreover, the 2 inhibitors used and the mitochondrial dye staining did not have any effect on cell viability (Supplemental Figure 1A; supplemental material available online with this article; https://doi.org/10.1172/jci.insight.123568DS1). Thus, the Mitolo subset constitutes a population with dye efflux capacity and will be referred to hereafter as “efflux(+),” with the corresponding Mitohi subset referred to as “efflux(–).” Compared with efflux(–) cells, efflux(+) cells expressed higher levels of MDR1 (ABCB1) (Supplemental Figure 1B), a cell surface transporter that mediates efflux of fluorescent dyes and xenobiotics in HSCs (25), suggesting that ABC transporters may contribute to dye efflux.
 Figure 1
Figure 1A subset of memory CD8+ T cells across human tissues efflux fluorescent dyes. Human T cells from the indicated tissue sites were loaded with mitochondrial dyes and analyzed by flow cytometry. (A) MitoTracker fluorescence within memory (CD45RA–CCR7–) CD8+ T cells in tissues of a representative donor. Numbers within plots indicate percentage of cells that are Mitohi (right) or Mitolo (left). (B) Comparison of CMXRos (upper) and MitoTracker Green (lower) staining of memory CD8+ T cells from the spleen and lung of a representative donor. (C) Inhibition of dye efflux by cyclosporine A (CSA) and verapamil. CD8+ T cells from spleen were labeled with MitoTracker green as in panel A in the presence or absence of CSA (top row) or verapamil (bottom row) at the indicated concentrations. Results are representative of 3 different donors. (D) Fraction of memory CD8+ T cells that are efflux(+) from the indicated tissue sites. LN, lymph node. Bar graph shows mean + SEM with individual samples shown from 4–20 donors for each tissue. (E) A subset of CMV-specific CD8+ T cells across tissues efflux dyes. Upper: Detection of CMV-specific CD8+ T cells within memory CD8+ T cells from indicated tissue sites. Lower: Frequency of efflux(+) and efflux(–) cells within the CMV tetramer+ population. Results are representative of 2 donors.
Compiling data from several donors, we found that the frequency of efflux(+) compared with efflux(–) populations differed between tissue sites, with the lowest frequency of efflux(+) memory T cells observed in the blood (≤40%) and the highest frequency observed in spleen and lung (>60%) (Figure 1D). Building on prior work in which we found that cytomegalovirus-specific (CMV-specific) T cells were maintained long-term in several tissue sites (26), we investigated whether efflux(+) T cells were present in CMV-specific populations. Indeed, efflux(+) cells were detected within CMV-specific memory T cells in multiple tissue sites in frequencies similar to the total efflux(+) frequency for that tissue (Figure 1E). Together, these results indicate that the proportion of efflux(+) cells is a feature of the tissue site and that efflux(+) cells can be generated following infection.
Tissue-resident phenotype of efflux(+) CD8+ T cells. Given the abundance of efflux(+) cells in tissues compared with blood, we investigated whether efflux(+) T cells in tissues exhibited features of human TRMs. We first assessed whether efflux(+) cells were differentially distributed within CD69+ and CD69– fractions, as CD69 is a phenotypic marker that distinguishes human TRMs from circulating memory T cells (9, 13, 18). In spleen, BM, and lung, efflux(+) cells were more highly represented among the CD69+ compared with the CD69– fraction of memory CD8+ T cells across multiple donors (Figure 2, A and B). Importantly, CD69+ memory CD8+ T cells were highly enriched for efflux(+) cells, comprising an average of 70% CD69+ T cells across multiple tissues and donors (Figure 2B). We also assessed whether the efflux(+) CD69+ T cells were mucosa-associated invariant T (MAIT) cells, which have previously been shown to efflux dyes (27, 28). We detected only a small fraction (<10%) of T cells within tissues expressing the canonical phenotype of MAIT cells (CD161+/Vα7.2+) among CD69+ T cells (Supplemental Figure 2), indicating that the majority of efflux(+) T cells across healthy human tissues are polyclonal memory CD8+ T cells.
 Figure 2
Figure 2Efflux(+) memory CD8+ T cells are enriched in the TRM fraction across human tissues. (A) Upper: CD69 expression by efflux(+) (Mitolo) and efflux(–) (Mitohi) subsets of memory CD8+ T cells in the indicated tissues. Lower: Compiled frequency of efflux(+) cells within the CD69+ (TRM-enriched) and CD69– subsets of memory CD8+ T cells in each tissue site, with each line connecting subsets from 1 individual donor (spleen, 17 donors; bone marrow, 7 donors; lung, 6 donors). (B) Frequency of TRMs that are efflux(+) in each tissue site, with bars showing mean ± SEM and individual donors represented by single dots. (C) Upper: CD103 expression by efflux(+) and efflux(–) TRMs in spleen and lung. Lower: Compiled frequency of CD103 expression among efflux(+) and efflux(–) TRMs in the spleen (n = 10) and lung (n = 6), with each line connecting subsets within individual donors. (D) Upper: Expression of CD49a and CD101 among efflux(+) and efflux(–) TRMs (CD69+) from the spleen of a representative donor. Lower: Compiled frequency of CD49a- and CD101-expressing cells within efflux(+) and efflux(–) splenic TRMs (n = 6). For all panels, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001 by paired t test.
We previously determined that human TRMs are enriched within the CD69+ fraction of tissue memory T cells and exhibit a core phenotypic and functional profile (12). We therefore investigated the expression of core TRM-associated markers by efflux(+) and efflux(–) subsets of CD69+ memory CD8+ T cells (TRMs). Strikingly, expression of CD103, a canonical CD8+ TRM marker (29), was enriched within the efflux(+) compared with efflux(–) fraction in both spleen and lung, with some interdonor variability for the lung (Figure 2C). Similarly, expression of the integrin CD49a and CD101, both part of the human TRM core signature (12), was increased within the efflux(+) compared with efflux(–) TRM subset in multiple donors (Figure 2D). Overall, these results suggest that efflux(+) cells predominate within the TRM compartment and express canonical TRM markers at high levels compared with their efflux(–) counterparts.
Efflux(+) TRMs but not circulating TEMs have a unique surface receptor phenotype. We next investigated the phenotype of efflux(+) and efflux(–) cells. Given the relative enrichment of efflux(+) cells within TRMs, we sought to determine if efflux status is associated with unique properties specifically for TRMs or more generally across memory subsets. We first measured CD127 (IL-7 receptor) expression, as IL-7 is critical for memory T cell homeostasis, and high CD127 expression is associated with long-lived memory CD8+ cells, while low CD127 expression can indicate chronic activation and exhaustion (30, 31). Efflux(+) TRMs had elevated CD127 expression compared with efflux(–) TRMs (Figure 3A), while no differences were observed between efflux(+) and efflux(–) TEMs. We then assessed the expression of the coreceptors CD27 and CD28, which are downregulated following TCR stimulation. Within the TRM compartment, efflux(+) TRMs had a significantly reduced frequency of CD27+ and CD28+ cells compared with efflux(–) TRMs (Figure 3, B and C). For circulatory TEMs, CD27 and CD28 expression was similar for both efflux(+) and efflux(–) subsets (Figure 3, B and C).
 Figure 3
Figure 3Efflux(+) TRMs have a unique phenotype. (A) Left: CD127 expression by efflux(+) and efflux(–) TRMs and TRMs from the spleen of 1 representative donor. Right: Compiled frequencies of CD127+ efflux(+) and efflux(–) TRMs and TEM subsets from spleen (n = 5) and bone marrow (BM, n = 7). (B and C) Left: CD27 (B) and CD28 (C) expression by efflux(+) and efflux(–) TRMs and TRMs from the spleen of 1 representative donor. Right: Compiled frequencies of CD27+ (B) and CD28+ (C) efflux(+) and efflux(–) TRMs and TEM subsets from spleen (n = 7). (D) Left: Expression of PD-1 by efflux(+) and efflux(–) TRM and TEM subsets from the spleen of a representative donor. Right: Compiled expression of PD-1 by efflux(+) and efflux(–) TRMs from spleen (n = 8), lung (n = 7), and BM (n = 6). (E) Left: Histograms of CD57 expression by efflux(+) and efflux(–) TRMs and TEM subsets from the spleen of 1 representative donor. Right: Compiled frequencies of CD57+ TRMs and TEMs from spleen (n = 7) and BM (n = 3). (F) Left: Plots show CD39 expression by efflux(+) and efflux(–) TRM and TEM subsets from the spleen of 1 representative donor. Right: Compiled frequencies of CD39+ TRMs and TEMs from spleen (n = 6). For all panels, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001 by paired t test. ns and n.s., not significant.
Expression of PD-1, which inhibits T cell responses and is associated with exhaustion and chronic activation (32), was reduced in efflux(+) compared with efflux(–) TRMs (CD69+) (Figure 3D). Within TEMs, some differences were seen within spleen and BM; however, the differences were of reduced magnitude as compared with the TRM fraction. Additionally, efflux(+) TRMs had lower expression of CD57, a marker of replicative senescence and cytotoxicity (33), compared with efflux(–) TRMs, while no significant differences were seen in circulatory TEMs (Figure 3E). Further supporting a core tissue-resident phenotype, efflux(+) cells also expressed higher levels of CD39 (Figure 3F), a marker of liver-resident TRMs (34) that has also been proposed to mark functionally distinct CD8+ T cells with immunomodulatory function (35). Overall, these results suggest that efflux status is associated with unique phenotypic properties within the TRM compartment but not for circulatory memory T cells. Efflux(+) TRMs appear to have a distinct activation history and potentially enhanced capacity for cytokine responses and longevity.
TRM subsets differing in efflux capacities exhibit a distinct transcriptional profile. Given that efflux status was associated with a unique phenotype primarily within TRMs, we performed whole transcriptome profiling of efflux(+) and efflux(–) CD69+ memory CD8+ T cells from 3 organ donors by RNA-sequencing (RNA-Seq) to further characterize these subsets (Supplemental Table 1). Principal component analysis (PCA) revealed that the efflux(+) subset is transcriptionally distinct from the efflux(–) subset across all donors, based on the second principal component accounting for 21% of the variation in gene expression between these subsets (Figure 4A). We used DAVID online functional annotation analysis on the top 500 differentially expressed genes to interpret their biological significance (36, 37). This analysis reveals enriched pathways that contain genes differentially expressed within our data set. These results indicate fold enrichment and significance but do not give information about the directionality of the pathways. The pathway with the greatest enrichment within our gene set was phospholipid translocation, and significant enrichment for pathways controlling T cell costimulation, signaling, cytokine responses, adhesion, and migration was also observed (Figure 4B).
 Figure 4
Figure 4Efflux(+) cells are a transcriptionally distinct subset of TRMs. Whole transcriptome profiling by RNA sequencing was performed on efflux(+) and efflux(–) CD69+ (TRM) memory CD8+ T cells from the spleen of 3 donors. (A) PCA of efflux(+) and efflux(–) TRM samples, based on the global transcriptome. (B) Functional annotation analysis by DAVID software. Select gene ontology (GO) terms with significant adjusted P values (Adj. p) are displayed, along with fold enrichment. (C) Heatmap of normalized expression levels of all genes with significant differential expression between the 2 groups, defined as FDR ≤ 0.05 and absolute value of log2 fold change ≥ 1. (D) Select significantly differentially expressed genes that are upregulated [“up in efflux(+)”]or downregulated [“down in efflux(+)”] grouped by category. Shown are the log2 fold changes (log2FC) of select genes between efflux(+) and efflux(–) cells for each donor, designated by a unique shape (see legend in bottom left panel). Genes marked with a “*” did not meet FDR criteria (0.05), but had log2FC ≥ 1 and significant P values and were included for potential biological relevance.
We identified 133 genes differentially expressed between efflux(+) and efflux(–) TRMs by DESEQ analysis applying criteria for significance (false discovery rate [FDR] ≤ 0.05 and absolute value of log2 fold change > 1; see Methods). Notably, the number of genes distinguishing efflux(+) and efflux(–) TRMs was much lower than the 300–400 differentially expressed genes we previously reported when comparing TRM-enriched (CD69+) to circulating CD69– memory CD8+ T cells in human blood and tissues (12). Expression levels of these 133 genes were consistent across all 3 donors (Figure 4C) and comprised several major pathways (Figure 4D). Genes associated with nutrient, ion, and xenobiotic transporters including ABCA1, which encodes the protein MDR1, were expressed at higher levels in efflux(+) TRMs, as were those involved in cell adhesion such as ITGAE (encoding CD103), ITGA1 (encoding CD49a), NCAM1, MCAM1, CDH4, and ANK1 (Figure 4D); efflux(+) TRMs additionally upregulated CCR9, CCR1, and CCR6, and downregulated CCR4 and CCR8, receptors involved in chemotaxis (Figure 4D).
Expression of key transcription factors regulating differentiation and function differed between effluxing subsets (Figure 4D), with efflux(+) TRMs exhibiting increased expression of TLE1, a transcriptional regulator associated with Notch/RBPJ signaling (38), which regulates TRM differentiation (39). Moreover, efflux(+) cells had elevated RORC and RORA expression, two transcription factors that drive Tc17-type responses in CD8+ T cells (40), and expressed high levels of IL17A, and genes encoding IL-23 and IL-17 receptors (Figure 4D), all associated with type 17 responses. Notably, levels of the nuclear receptor NR4A1, which is upregulated in blood efflux(+) cells (19), did not differ significantly between efflux(+) and efflux(–) TRM subsets. Analysis of genes associated with cell cycle and apoptosis control showed elevated levels of CD101, SPRY1, and SPRY2, genes with documented roles in suppressing T cell proliferation and TCR-mediated calcium signaling (41, 42), coupled with reduced levels of cyclin B2 (CCNB2). Overall, these data indicate that efflux(+) TRMs have a unique transcriptional program composed of genes controlling adhesion and migration, as well as T cell activation, function, and proliferation.
Distinct functional profile and proliferative capacity of efflux(+) TRMs. Differential expression of key genes involved in cell cycle, quiescence, and T cell function suggested distinct proliferative and functional capacities of efflux(+) and efflux(–) TRMs. Before examining the function of these TRM subsets, we chose to investigate the stability of these subsets after stimulation. After 48 hours of stimulation, the majority of efflux(+) TRMs retained their ability to efflux dyes, and similarly the majority of efflux(–) TRMs remained efflux(–) (Figure 5A). However, a small fraction of each subset did convert to the opposite subset (Figure 5A), suggesting some plasticity following stimulation. We then measured the ability of these TRM subsets to produce cytokines following 48 hours of stimulation. Efflux(–) TRMs produced higher levels of multiple cytokines compared with efflux(+) TRMs, including proinflammatory mediators TNF-α (1.7-fold difference) and IFN-γ (1.4-fold difference), IL-2 (2.2-fold difference), and the Th2-associated cytokine IL-4 (8.7-fold difference) (Figure 5B). By contrast, efflux(+) TRMs produced increased levels of IL-17 compared with efflux(–) TRMs (Figure 5B), consistent with transcriptome profiling results. Interestingly, levels of IL-10 production were comparable between the 2 subsets. Along with increased effector cytokine production, efflux(–) TRMs exhibited markedly higher degranulation measured by CD107a (LAMP1) expression compared with efflux(+) TRMs (Figure 5C).
 Figure 5
Figure 5Efflux(+) TRM subset stability and function following stimulation. (A) Sorted efflux(+) and efflux(–) TRMs were stimulated with anti–CD3/CD28/CD2 beads, and efflux capacity was reassessed after 48 hours along with CD25 expression. Plots show MitoTracker and CD25 expression from the spleen of 1 donor for the indicated subsets. (B) Cytokine production following TCR stimulation. Efflux(+) and efflux(–) TRMs were sorted and stimulated with anti–CD3/CD28/CD2 beads for 72 hours, and cytokines in supernatant were quantified using cytometric bead array (see Methods). Graphs show levels of indicated cytokines in supernatants compiled from 8 donors. (C) Degranulation of efflux(+) and efflux(–) TRM subsets. Sorted cells were pulse labeled with CD107a antibody followed by PMA/ionomycin stimulation. Representative plots and quantification of CD107a+ from splenic efflux(+) and efflux(–) TRM subsets from 4 donors. For all panels, *P ≤ 0.05. **P ≤ 0.01 ****P ≤ 0.0001 by paired t test. ns, not significant.
We recently showed that CD8+ TRMs can vary in proliferative capacity (43). When fractionated based on efflux capacity, efflux(+) memory cells exhibited increased proliferation compared with efflux(–) cells following stimulation (Figure 6A). TCR stimulation induces metabolic reprogramming and expression of key transcription factors that regulate proliferation and effector cell differentiation such as IRF4 (44–47). While proliferating efflux(+) and efflux(–) TRMs expressed increased levels of IRF4 compared with nondividing cells, IRF4 levels in proliferating efflux(–) cells were higher than those in efflux(+) cells across multiple divisions and in multiple donors (Figure 6B). As IRF4 is associated with effector differentiation, these results are consistent with the findings above showing that efflux(–) TRMs produce higher levels of TNF and IFN-γ compared with efflux(+) TRMs (Figure 5B).
 Figure 6
Figure 6Efflux(+) TRMs have enhanced proliferative capacity and responses to homeostatic cytokines. (A and B) Proliferative capacity of TRM subsets. Sorted efflux(+) and efflux(–) TRMs were labeled with cell proliferation dye and stimulated with anti–CD3/CD28/CD2 beads. Proliferation was assessed on day 4 of culture. (A) Left: Histogram showing cell proliferation dye dilution at day 4. Results are representative of 4 independent experiments. Middle: Quantification of the percentage of proliferating cells. Right: Quantification of the percentage of proliferating cells at each division number. Significance assessed by 2-way ANOVA with Sidak’s multiple comparisons test. (B) Differential IRF4 induction in efflux(+) and efflux(–) TRMs following TCR stimulation. Left: IRF4 expression as a function of cell division at day 4 after stimulation; plots are representative of 3 independent experiments. Rectangular gate identifies proliferating cells that have upregulated IRF4. Middle: IRF4 expression within proliferating cells from efflux(+) and efflux(–) subsets. Right: Quantification of IRF4 mean fluorescence intensity within activated proliferating cells from 3 donors. (C) Left: Expression of Ki67 within TRM subsets from the spleen of 1 representative donor. Right: Frequency of effluxing cells in Ki67+ and Ki67– TRMs. (D) Efflux(+) TRMs exhibit increased responses to IL-7. Left: STAT5 phosphorylation following IL-7 stimulation ex vivo. Right: Quantification of pSTAT5+ cell percentage within efflux(+) and efflux(–) TRMs. For all panels, *P ≤ 0.05. **P ≤ 0.01 ****P ≤ 0.0001 by paired t test. n.s., not significant.
The reduced effector function and increased proliferative capacity of efflux(+) compared with efflux(–) TRMs suggested that these subsets may likewise differ in their homeostatic maintenance. Consistent with this hypothesis, efflux(+) TRMs expressed lower levels of Ki67 (Figure 6C) compared with efflux(–) TRMs, suggesting that efflux(+) TRMs persist in a more quiescent state. Efflux(+) TRMs also exhibited increased IL-7 signaling following IL-7 stimulation ex vivo as shown by enhanced STAT5 phosphorylation (Figure 6D), a key effector of IL-7 signals (31), consistent with our finding of increased CD127 expression by efflux(+) compared with efflux(–) TRMs (Figure 3). These results indicate that efflux(+) TRMs have an increased capacity to respond to homeostatic cytokines important for memory cell longevity. Overall, these results demonstrate that while efflux(+) TRMs have increased capacity to proliferate and produce IL-17, efflux(–) TRMs exhibit high effector and cytotoxic potential and reduced proliferation.
The transcriptional response of TRM subsets to TCR stimulation. To investigate the bases for the different functional response of efflux(+) and efflux(–) TRM subsets to TCR/CD3 stimulation, we examined their activation-induced transcriptional profile. We performed RNA-Seq on efflux(+) and efflux(–) TRM subsets as in Figure 4 following 12-hour TCR stimulation. Applying the criteria for significance as above, we found 865 and 487 genes to be differentially expressed between stimulated and unstimulated efflux(+) and efflux(–) TRMs, respectively (Figure 7A). Interestingly, more genes were downregulated than upregulated following stimulation for both efflux(+) and efflux(–) cells (Figure 7A). Further, the number of genes differentially expressed by either subset was substantially higher than the number of genes differentially expressed between memory and naive T cell subsets (48) or between TRMs and TEMs in humans (12). These data suggest that TRMs are poised for a robust and rapid transcriptional response to stimulation.
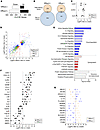 Figure 7
Figure 7The transcriptional response of efflux(+) and efflux(–) TRMs to stimulation. RNA-Seq analysis was performed on splenic efflux(+) and efflux(–) TRM subsets isolated as in Figure 3 following stimulation with anti–CD3/CD28 for 12 hours. (A) Differential expression was assessed using DESEQ2, and plots display the number of significantly upregulated and downregulated genes following stimulation in both efflux(+) and efflux(–) subsets. (B) Both efflux(+) and efflux(–) TRMs preserve TRM-like characteristics after stimulation. Venn diagrams show overlap between genes that are differentially expressed when comparing stimulated versus unstimulated TRMs (current data set) and genes that are differentially expressed when comparing human spleen CD8+ TRMs and TEMs (“TRM core genes”; from ref. 12). (C) TRMs downregulate egress receptors after stimulation. Plot shows log2 fold changes (log2FC) of CCR7, S1PR1, and KLF2 when comparing stimulated versus unstimulated efflux(+) and efflux(–) TRM samples. (D) Scatterplot displays all genes found to have significant differential expression in either efflux(+) or efflux(–) stimulated versus unstimulated samples as in A. Value on the x axis represents the log2FC of the gene between stimulated versus unstimulated efflux(+) samples, and the y axis represents the log2FC of the same gene between stimulated versus unstimulated efflux(–) samples. Samples are color coded by whether the differential expression was significant in only efflux(+) TRMs, only efflux(–) TRMs, or both groups. (E) Ingenuity Pathway analysis (IPA) analysis. Select pathways that had significant P values (≤0.01) when comparing stimulated versus unstimulated samples in both efflux(+) and efflux(–) TRMs are displayed. Direction of enrichment in stimulated samples is proportional to the color intensity of each bar. (F) Plot shows log2FC of select genes related to T cell function when comparing stimulated versus unstimulated TRM samples for both efflux(+) and efflux(–) subsets. (G) Expression of genes (log2FC) from C that have opposite-direction changes for efflux(+) and efflux(–) samples after stimulation across all 3 donors. Plot shows log2FC of the selected genes, comparing expression by stimulated versus unstimulated samples for efflux(+) (orange triangles) and efflux(–) (blue circles) TRMs.
The magnitude of the transcriptional response to stimulation raised the question of whether efflux(+) or efflux(–) cells lose TRM-like properties after stimulation, particularly those associated with tissue retention. To address this, we compared the genes that were differentially expressed following stimulations (from our current data set) to the genes that are differentially expressed between TRMs and TEMs (using our previously published data set in ref. 12). Interestingly, very few genes overlapped between these 2 data sets (Figure 7B), implying that transcriptional changes following stimulation in either efflux(+) or efflux(–) cells are unrelated to the genes that define the TRM subset. TRMs are characterized by low expression of the homing receptors S1PR1 and CCR7 and reduced expression of the associated transcription factor KLF2, which together help TRMs avoid egress cues (4, 49). Following stimulation, both efflux(+) or efflux(–) TRMs further downregulated these genes (Figure 7C), implying that the migratory program of TRMs is reinforced to allow these cells to carry out functions in situ. Taken together, these results suggest that while TRMs undergo large transcriptional changes upon activation, these cells remain TRM and do not lose defining features of the subset.
Comparison of the transcriptional response to stimulation between efflux(+) and efflux(–) TRMs revealed a similar set of genes that was differentially regulated (Figure 7D). Following stimulation, there was significant downregulation of genes involved in pathways for cytokine signaling and TCR-coupled signaling, with the sirtuin pathway, which controls metabolism and cell cycle progression/apoptosis (50), emerging as the top result for both subsets (Figure 7E). Other pathways included cell cycle regulation and metabolism (Figure 7E). Genes related to T cell function and cytokine signaling were also differentially expressed, including proinflammatory cytokine genes (IL17F, LTA [lymphotoxin α], and IL13), the cytotoxic gene GZMB (encoding granzyme B), and a number of chemokines (Figure 7F).
A limited number of genes exhibited divergent upregulation/downregulation in activated efflux(+) compared to efflux(–) cells (Figure 7G). Of note, efflux(+), but not efflux(–), TRMs upregulated expression of ITGB8, an integrin critical for local activation of latent TGF-β complexes on Tregs (51). Efflux(+) TRMs also upregulated expression of TACR1, encoding neurokinin-1-R, a receptor with a role in type 17 responses that is potentiated by TGF-β signaling (52). Taken together, these results suggest that TGF-β signaling and type 17 responses may be differentially regulated by efflux(+)/efflux(–) subsets. Finally, efflux(+) cells downregulated ABCA1 upon activation (Figure 7G). As this transporter may mediate dye efflux (Supplemental Figure 1 and ref. 25), this suggests that at a least a fraction of efflux(+) cells give rise to efflux(–) progeny.
-
Discussion
It has become increasingly clear that long-term immunity is regulated locally at the tissue level through the establishment and persistence of TRMs. Using healthy primary human tissues, we provide evidence for functional heterogeneity within the CD8+ TRM compartment based on differential capacity to efflux fluorescent dyes. Critically, efflux(+) and efflux(–) TRM subsets were transcriptionally and functionally distinct, both at steady state and following TCR stimulation. Efflux(+) cells differentially express transcription factors related to type 1 and type 17 inflammatory responses, indicating that key regulators of lymphocyte cell fate decisions may also program distinct subsets of TRMs. These TRM subsets also exhibited a differential capacity for IFN-γ/TNF-α and IL-17 production as well as a differential propensity toward degranulation. This stratification of function suggests that different TRM subsets may preferentially mediate specific functions.
Previous studies have found that effluxing memory T cells were enriched among certain TRM populations in specific sites (19). Here, we show through an extended functional and transcriptional analysis of TRMs in multiple human tissues that TRMs comprise both effluxing and non-effluxing populations, with each playing distinct roles. Efflux(+) cells retain a higher proliferative potential following TCR stimulation and may constitute a resting pool of cells that repopulates the more effector-like efflux(–) subset to promote type 1 inflammatory responses. Differences in the expression of genes controlling cell cycle progression and proliferation, as well as different levels of inhibitory receptor expression, may together explain the observed differences in proliferative capacities. Additionally, efflux(+) TRMs exhibited increased responses to IL-7, a cytokine associated with memory maintenance. These results demonstrate that a portion of TRMs are not fully terminally differentiated and can undergo substantial proliferation during re-activation. This proliferation of human TRMs identified here is consistent with 2 recent reports showing that mouse TRMs could proliferate in situ to antigenic peptide and pathogen challenge (53, 54).
Our analysis of transcriptional responses to TCR stimulation has implications for TRM biology as a whole. We found that TRMs preserve their core profile of downmodulated tissue egress molecules (e.g., S1PR1, CCR7, and KLF2) after TCR stimulation, while undergoing a rapid and extensive transcriptional response. Notably, TRM stimulation preserved a tissue-retentive profile and was accompanied by extensive downmodulation of genes involved in cytokine and TCR signaling, with a focused upregulation of specific cytokines and chemokines. We propose that this narrowing of transcriptional response in activated TRMs enables them to promote effective and specific responses in situ, without triggering overt tissue damage and inflammation.
The stratification of function between TRM subsets suggests a cooperative model for TRM maintenance and functional responses. Specifically, efflux(–) TRMs may contribute more to IFN-γ/TNF-α responses and cytotoxic functions, while efflux(+) TRMs may mediate type 17 inflammation to a greater extent while serving as a proliferative reservoir to replenish the TRM compartment. However, transcriptional data also indicate substantial overlapping functions, suggesting a model in which certain functions are universal properties of TRMs, while others are primarily mediated by a specific TRM subset. The division of labor between TRM subsets may also be driven by anatomic differences, as suggested by distinct adhesion molecule and migratory receptor expression.
While we identified differences in function between these TRM subsets, we have no evidence that efflux capacity, per se, has direct effects on their function. Rather, it may reflect the physiological state of the cells and their ability to survive in diverse environments. Efflux pumps expel toxic xenobiotics and have been implicated in the persistence of human lymphocytes during chemotherapy (28). TRMs persist in peripheral tissue sites for years or even decades, where they are exposed to a range of foreign agents, particularly in sites such as skin and lung. Given our data that efflux(+) TRMs show evidence of increased quiescence and longevity, heightened expression of efflux pumps may contribute to TRM survival and homeostasis in peripheral tissues. Efflux capacity may also mediate differential susceptibility to chemotherapy and drug treatments (55).
Given that efflux(+) TRMs exhibit increased IL-17 production as well as Th17-associated signaling, specifically targeting efflux(+) cells in IL-17–mediated inflammatory diseases such as psoriasis might be an optimal therapy that spares protective TRMs while eliminating pathogenic TRMs. Previously, there has been interest in the possibility of targeted immune therapies that specifically modulate either TRMs or circulatory T cells while leaving other aspects of the immune system undisturbed (7). Our data suggest that this specificity can be extended to target specific TRM subsets. Overall, the identification of these distinct subsets could be leveraged toward next-generation therapies for infection, cancer, and autoimmunity.
-
Methods
Acquisition of tissue from human organ donors. Human tissues were obtained from deceased organ donors at the time of organ acquisition for clinical transplantation through an approved research protocol and MTA with LiveOnNY, the organ procurement organization for the New York metropolitan area. All donors were free of chronic disease and cancer, hepatitis B, C, and HIV negative. A list of donors from which tissues were obtained and used in this study is presented in Supplemental Table 1.
Cell isolation from human lymphoid and nonlymphoid tissues. Tissue samples were maintained in cold saline and brought to the laboratory within 2–4 hours of organ procurement. Spleen, lung, and BM samples were processed using enzymatic and mechanical digestion resulting in high yields of live leukocytes, as described previously (21, 22). Mononuclear cells were isolated from peripheral blood using centrifugation through lymphocyte separation medium (Corning). Non-enzymatic isolation was also used for spleen tissue using the Bullet Blender Tissue Homogenizer (Next Advance). Tissue samples were chopped into small pieces (≤5 mm) using scissors and 4–5 g of tissue was placed in a 50-ml conical tube and complete RPMI was added to a total volume of 10 ml, followed by addition of 7 or 8 4.8-mm stainless steel beads (product SSB48). Tissues were homogenized in the bullet blender for 2 minutes at speed setting 3–4. Following homogenization, the mixture was filtered through a 70-μm Cell Strainer (Corning). ACK buffer was used for RBC lysis, followed by an additional filtration through a 70-μm Cell Strainer.
Efflux dye labeling. T cells were labeled with MitoTracker Green FM (50 nM) or CMXRos (25 nM) (Thermo Fischer Scientific) for 15 minutes in complete media (10% FBS in RPMI) according to the manufacturer’s instructions. Efflux was blocked by performing fluorescent labeling in the presence of 25–50 μM CSA (24) or in the presence of 25-50 μM verapamil.
Functional assays. For quantification of cytokine production by different subsets, sorted cells were plated in 96-well round-bottom plates at 105 cells/well in complete RPMI medium and stimulated using anti–CD3/CD28/CD2 beads for 72 hours (T Cell Activation/Expansion Kit, Miltenyi Biotec). Cytokine levels in cell supernatants were measured using a BD Cytometric Bead Array (Human Th1/Th2/Th17 Cytokine Kit). To quantify degranulation, sorted cells were labeled with fluorochrome-conjugated anti-CD107a antibody before plating in complete media and subsequently stimulated with PMA/ionomycin for 3 hours. For assays of responses to cytokines, sorted cells were rested in RPMI medium without serum for 2 hours. Cells were then transferred to complete media and stimulated for 20 minutes with 50 ng/ml IL-7 (PeproTech) before flow cytometry analysis. For proliferation assays, cells were labeled with Cell Trace Violet (Thermo Fisher Scientific) according to the manufacturer’s instructions before stimulation with anti–CD3/CD28/CD2 beads for 4 days.
Flow cytometry. Cells were stained according to standard flow cytometry protocols using antibodies listed in Supplemental Table 2. For detection of transcription factors, cells were fixed and permeabilized using the eBioscience Foxp3 kit. For phospho–flow cytometry, cells were first fixed with 4% paraformaldehyde for 15 minutes followed by permeabilization with ice-cold methanol. All samples were acquired on a BD Fortessa or LSRII flow cytometer. For isolation of subsets by fluorescence-activated cell sorting (FACS), lymphocyte suspensions were enriched for T cells using the MojoSort Human CD3 T cell Isolation Kit (Biolegend), stained for surface markers as indicated, and sorted using a BD Influx high-speed sorter or an Aria IIu sorter (BD Biosciences). For identification of CMV-specific T cells, samples from HLA-A2 haplotype donors were labeled with IE1-VLE peptide pentamers (Proimmune) (26). Data were analyzed using FlowJo V.10 software (Tree Star).
RNA-Seq and analysis. FACS was used to isolate CD3+CD8+CD69+ memory cells (CD45RA–CCR7–) that were either efflux(–) or efflux(+). Cells were sorted from the spleen of 3 individual donors (aged 32, 32, and 59 years); some cells were stimulated with anti–CD3/CD28/CD2 beads for 12 hours prior to RNA isolation. RNA was isolated from cell pellets using the RNeasy Mini Kit (Qiagen) and quantified using an Agilent 2100 Bioanalyzer (Agilent Technologies). RNA-Seq libraries were prepared using standard TruSeq with poly(A) pulldown and were sequenced on the Illumina HiSeq 2500 with 101-bp single-end reads at the Columbia Genome Center. RNA-Seq reads were mapped using TopHat (56) with default parameters to the human reference genome build hg19, data quality control was performed using RNA-SeQC (57), read counts computed using HTSeq (58), and per-gene fragment per kilobases per million reads (FPKM) estimated using Cufflinks (59). Differential gene expression analysis was performed with DESeq2 (60), and pathway analysis with Ingenuity Pathway Analysis software (IPA, Qiagen). We considered genes to be significantly differentially expressed between 2 groups if FDR ≤ 0.05 and absolute value of log2 fold change > 1. Data were deposited in the NCBI’s Gene Expression Omnibus database (GEO GSE109841).
Statistics. Statistical analyses were performed in GraphPad Prism (v.7). A P value of ≤ 0.05 was considered significant; statistical tests are specified in each figure legend. All t tests were 2 tailed.
Study approval. Isolation of blood and tissues from organ donors does not qualify as human subjects research and does not require informed consent, as confirmed by the Columbia University Institutional Review Board (IRB, Columbia University, New York).
-
Author contributions
RK and BVK designed and performed experiments, processed tissues, analyzed data, and wrote the manuscript. MM performed experiments, processed tissues, and analyzed data. DJC and TS obtained donor tissues. SLR planned experiments, analyzed data, and wrote the manuscript. HL and AF coordinated tissue acquisition from organ donors. DLF planned experiments, coordinated tissue and data acquisition, analyzed data, and wrote the manuscript.
-
Acknowledgments
This work was supported by NIH grants AI06697 and HL116136 (to DLF) and T32GM007367 (to BVK). These studies were performed in the CCTI Flow Cytometry Core funded in part through the S10 Shared Instrumentation Grants, 1S10RR027050 (LSRII) and S10OD020056 (Influx), and 5P30DK063608. We wish to gratefully acknowledge the generosity of the donor families and the outstanding efforts of LiveOnNY transplant coordinators and staff for making this study possible.
Address correspondence to: Donna L. Farber, 650 West 168th Street, BB1701G, New York, New York 10032, USA. Phone: 212.305.6030; Email: df2396@cumc.columbia.edu.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
License: Copyright 2018, American Society for Clinical Investigation.
Reference information: JCI Insight. 2018;3(22):e123568. https://doi.org/10.1172/jci.insight.123568.
-
References
- Farber DL, Yudanin NA, Restifo NP. Human memory T cells: generation, compartmentalization and homeostasis. Nat Rev Immunol. 2014;14(1):24–35.
- Shin H, Iwasaki A. A vaccine strategy that protects against genital herpes by establishing local memory T cells. Nature. 2012;491(7424):463–467.
- Zens KD, Chen JK, Farber DL. Vaccine-generated lung tissue-resident memory T cells provide heterosubtypic protection to influenza infection. JCI Insight. 2016;1(10):e85832.
- Mueller SN, Mackay LK. Tissue-resident memory T cells: local specialists in immune defence. Nat Rev Immunol. 2016;16(2):79–89.
- Hondowicz BD, et al. Interleukin-2-dependent allergen-specific tissue-resident memory cells drive asthma. Immunity. 2016;44(1):155–166.
- Turner DL, et al. Biased generation and in situ activation of lung tissue-resident memory CD4 T cells in the pathogenesis of allergic asthma. J Immunol. 2018;200(5):1561–1569.View this article via: PubMed Google Scholar
- Clark RA. Resident memory T cells in human health and disease. Sci Transl Med. 2015;7(269):269rv1.
- Wong MT, et al. A high-dimensional atlas of human T cell diversity reveals tissue-specific trafficking and cytokine signatures. Immunity. 2016;45(2):442–456.
- Watanabe R, et al. Human skin is protected by four functionally and phenotypically discrete populations of resident and recirculating memory T cells. Sci Transl Med. 2015;7(279):279ra39.
- Cheuk S, et al. CD49a expression defines tissue-resident CD8+ T cells poised for cytotoxic function in human skin. Immunity. 2017;46(2):287–300.
- Bergsbaken T, Bevan MJ. Proinflammatory microenvironments within the intestine regulate the differentiation of tissue-resident CD8+ T cells responding to infection. Nat Immunol. 2015;16(4):406–414.
- Kumar BV, et al. Human tissue-resident memory T cells are defined by core transcriptional and functional signatures in lymphoid and mucosal sites. Cell Rep. 2017;20(12):2921–2934.
- Mackay LK, et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science. 2016;352(6284):459–463.
- Goodell MA, Brose K, Paradis G, Conner AS, Mulligan RC. Isolation and functional properties of murine hematopoietic stem cells that are replicating in vivo. J Exp Med. 1996;183(4):1797–1806.
- Goodell MA, et al. Dye efflux studies suggest that hematopoietic stem cells expressing low or undetectable levels of CD34 antigen exist in multiple species. Nat Med. 1997;3(12):1337–1345.
- Chaudhary PM, Roninson IB. Expression and activity of P-glycoprotein, a multidrug efflux pump, in human hematopoietic stem cells. Cell. 1991;66(1):85–94.
- Kudernatsch RF, et al. Human bone marrow contains a subset of quiescent early memory CD8(+) T cells characterized by high CD127 expression and efflux capacity. Eur J Immunol. 2014;44(12):3532–3542.
- Fergusson JR, et al. CD161(int)CD8+ T cells: a novel population of highly functional, memory CD8+ T cells enriched within the gut. Mucosal Immunol. 2016;9(2):401–413.
- Boddupalli CS, et al. ABC transporters and NR4A1 identify a quiescent subset of tissue-resident memory T cells. J Clin Invest. 2016;126(10):3905–3916.
- de Almeida MJ, Luchsinger LL, Corrigan DJ, Williams LJ, Snoeck HW. Dye-independent methods reveal elevated mitochondrial mass in hematopoietic stem cells. Cell Stem Cell. 2017;21(6):725–729.e4.
- Thome JJ, et al. Spatial map of human T cell compartmentalization and maintenance over decades of life. Cell. 2014;159(4):814–828.
- Sathaliyawala T, et al. Distribution and compartmentalization of human circulating and tissue-resident memory T cell subsets. Immunity. 2013;38(1):187–197.
- Pendergrass W, Wolf N, Poot M. Efficacy of MitoTracker Green and CMXrosamine to measure changes in mitochondrial membrane potentials in living cells and tissues. Cytometry A. 2004;61(2):162–169.View this article via: PubMed Google Scholar
- Takeguchi N, Ichimura K, Koike M, Matsui W, Kashiwagura T, Kawahara K. Inhibition of the multidrug efflux pump in isolated hepatocyte couplets by immunosuppressants FK506 and cyclosporine. Transplantation. 1993;55(3):646–650.
- Zhou S, et al. The ABC transporter Bcrp1/ABCG2 is expressed in a wide variety of stem cells and is a molecular determinant of the side-population phenotype. Nat Med. 2001;7(9):1028–1034.
- Gordon CL, et al. Tissue reservoirs of antiviral T cell immunity in persistent human CMV infection. J Exp Med. 2017;214(3):651–667.View this article via: PubMed Google Scholar
- Dusseaux M, et al. Human MAIT cells are xenobiotic-resistant, tissue-targeted, CD161hi IL-17-secreting T cells. Blood. 2011;117(4):1250–1259.
- Turtle CJ, Swanson HM, Fujii N, Estey EH, Riddell SR. A distinct subset of self-renewing human memory CD8+ T cells survives cytotoxic chemotherapy. Immunity. 2009;31(5):834–844.
- Schenkel JM, Masopust D. Tissue-resident memory T cells. Immunity. 2014;41(6):886–897.
- Kaech SM, Tan JT, Wherry EJ, Konieczny BT, Surh CD, Ahmed R. Selective expression of the interleukin 7 receptor identifies effector CD8 T cells that give rise to long-lived memory cells. Nat Immunol. 2003;4(12):1191–1198.
- Mazzucchelli R, Durum SK. Interleukin-7 receptor expression: intelligent design. Nat Rev Immunol. 2007;7(2):144–154.
- Sharpe AH, Pauken KE. The diverse functions of the PD1 inhibitory pathway. Nat Rev Immunol. 2018;18(3):153–167.View this article via: PubMed Google Scholar
- Brenchley JM, et al. Expression of CD57 defines replicative senescence and antigen-induced apoptotic death of CD8+ T cells. Blood. 2003;101(7):2711–2720.
- Pallett LJ, et al. IL-2high tissue-resident T cells in the human liver: Sentinels for hepatotropic infection. J Exp Med. 2017;214(6):1567–1580.
- Bai A, et al. NADH oxidase-dependent CD39 expression by CD8(+) T cells modulates interferon gamma responses via generation of adenosine. Nat Commun. 2015;6:8819. View this article via: PubMed Google Scholar
- Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57.
- Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37(1):1–13.
- Larabee JL, Shakir SM, Barua S, Ballard JD. Increased cAMP in monocytes augments Notch signaling mechanisms by elevating RBP-J and transducin-like enhancer of Split (TLE). J Biol Chem. 2013;288(30):21526–21536.
- Hombrink P, et al. Programs for the persistence, vigilance and control of human CD8+ lung-resident memory T cells. Nat Immunol. 2016;17(12):1467–1478.
- Curtis MM, Way SS, Wilson CB. IL-23 promotes the production of IL-17 by antigen-specific CD8 T cells in the absence of IL-12 and type-I interferons. J Immunol. 2009;183(1):381–387.
- Akbulut S, et al. Sprouty proteins inhibit receptor-mediated activation of phosphatidylinositol-specific phospholipase C. Mol Biol Cell. 2010;21(19):3487–3496.
- Soares LR, Tsavaler L, Rivas A, Engleman EG. V7 (CD101) ligation inhibits TCR/CD3-induced IL-2 production by blocking Ca2+ flux and nuclear factor of activated T cell nuclear translocation. J Immunol. 1998;161(1):209–217.View this article via: PubMed Google Scholar
- Miron M, et al. Human lymph nodes maintain TCF-1hi memory T cells with high functional potential and clonal diversity throughout life. J Immunol. 2018;201(7):2132–2140.View this article via: PubMed Google Scholar
- Yao S, et al. Interferon regulatory factor 4 sustains CD8(+) T cell expansion and effector differentiation. Immunity. 2013;39(5):833–845.
- Man K, et al. The transcription factor IRF4 is essential for TCR affinity-mediated metabolic programming and clonal expansion of T cells. Nat Immunol. 2013;14(11):1155–1165.
- Raczkowski F, et al. The transcription factor interferon regulatory factor 4 is required for the generation of protective effector CD8+ T cells. Proc Natl Acad Sci USA. 2013;110(37):15019–15024.
- Pollizzi KN, Powell JD. Integrating canonical and metabolic signalling programmes in the regulation of T cell responses. Nat Rev Immunol. 2014;14(7):435–446.
- Weng NP, Araki Y, Subedi K. The molecular basis of the memory T cell response: differential gene expression and its epigenetic regulation. Nat Rev Immunol. 2012;12(4):306–315.
- Skon CN, Lee JY, Anderson KG, Masopust D, Hogquist KA, Jameson SC. Transcriptional downregulation of S1pr1 is required for the establishment of resident memory CD8+ T cells. Nat Immunol. 2013;14(12):1285–1293.
- Vachharajani VT, Liu T, Wang X, Hoth JJ, Yoza BK, McCall CE. Sirtuins link inflammation and metabolism. J Immunol Res. 2016;2016:8167273. View this article via: PubMed Google Scholar
- Kitamura H, et al. Mouse and human lung fibroblasts regulate dendritic cell trafficking, airway inflammation, and fibrosis through integrin αvβ8-mediated activation of TGF-β. J Clin Invest. 2011;121(7):2863–2875.
- Beinborn M, et al. TGF-beta regulates T-cell neurokinin-1 receptor internalization and function. Proc Natl Acad Sci USA. 2010;107(9):4293–4298.
- Park SL, et al. Local proliferation maintains a stable pool of tissue-resident memory T cells after antiviral recall responses. Nat Immunol. 2018;19(2):183–191.
- Beura LK, et al. Intravital mucosal imaging of CD8+ resident memory T cells shows tissue-autonomous recall responses that amplify secondary memory. Nat Immunol. 2018;19(2):173–182.
- Dimeloe S, et al. Human regulatory T cells lack the cyclophosphamide-extruding transporter ABCB1 and are more susceptible to cyclophosphamide-induced apoptosis. Eur J Immunol. 2014;44(12):3614–3620.
- Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25(9):1105–1111.
- Wang L, Wang S, Li W. RSeQC: quality control of RNA-seq experiments. Bioinformatics. 2012;28(16):2184–2185.
- Anders S, Pyl PT, Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31(2):166–169.
- Trapnell C, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28(5):511–515.
- Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550.
-
Version history
- Version 1 (November 15, 2018): Electronic publication










