Research ArticleHematologyOncology
Open Access |  10.1172/jci.insight.190833
10.1172/jci.insight.190833
Influence of oxidation resistance 1 on disease progression in chronic myeloid leukemia
Weiqi Huang,1,2,3 Bin Liu,1,2,3 Liping Hu,1,2 Chi-Hao Luan,4 Priyam Patel,1,2 Elizabeth Bartom,1,2 and Elizabeth A. Eklund1,2,3
1The Feinberg School of Medicine at Northwestern University, Chicago, Illinois, USA.
2The Lurie Cancer Center at Northwestern University, Chicago, Illinois, USA.
3Jesse Brown Veteran’s Hospital, Chicago, Illinois, USA.
4High Throughput Analysis Laboratory and Department of Molecular Biosciences, Northwestern University, Evanston, Illinois, USA.
Address correspondence to: E. Eklund, Northwestern University, 303 E. Superior Ave, Lurie 5-111, Chicago, Illinois 60611, USA. Phone: 213.503.3208; Email; e-eklund@northwestern.edu.
Find articles by Huang, W. in: PubMed | Google Scholar
1The Feinberg School of Medicine at Northwestern University, Chicago, Illinois, USA.
2The Lurie Cancer Center at Northwestern University, Chicago, Illinois, USA.
3Jesse Brown Veteran’s Hospital, Chicago, Illinois, USA.
4High Throughput Analysis Laboratory and Department of Molecular Biosciences, Northwestern University, Evanston, Illinois, USA.
Address correspondence to: E. Eklund, Northwestern University, 303 E. Superior Ave, Lurie 5-111, Chicago, Illinois 60611, USA. Phone: 213.503.3208; Email; e-eklund@northwestern.edu.
Find articles by Liu, B. in: PubMed | Google Scholar
1The Feinberg School of Medicine at Northwestern University, Chicago, Illinois, USA.
2The Lurie Cancer Center at Northwestern University, Chicago, Illinois, USA.
3Jesse Brown Veteran’s Hospital, Chicago, Illinois, USA.
4High Throughput Analysis Laboratory and Department of Molecular Biosciences, Northwestern University, Evanston, Illinois, USA.
Address correspondence to: E. Eklund, Northwestern University, 303 E. Superior Ave, Lurie 5-111, Chicago, Illinois 60611, USA. Phone: 213.503.3208; Email; e-eklund@northwestern.edu.
Find articles by Hu, L. in: PubMed | Google Scholar
1The Feinberg School of Medicine at Northwestern University, Chicago, Illinois, USA.
2The Lurie Cancer Center at Northwestern University, Chicago, Illinois, USA.
3Jesse Brown Veteran’s Hospital, Chicago, Illinois, USA.
4High Throughput Analysis Laboratory and Department of Molecular Biosciences, Northwestern University, Evanston, Illinois, USA.
Address correspondence to: E. Eklund, Northwestern University, 303 E. Superior Ave, Lurie 5-111, Chicago, Illinois 60611, USA. Phone: 213.503.3208; Email; e-eklund@northwestern.edu.
Find articles by Luan, C. in: PubMed | Google Scholar
1The Feinberg School of Medicine at Northwestern University, Chicago, Illinois, USA.
2The Lurie Cancer Center at Northwestern University, Chicago, Illinois, USA.
3Jesse Brown Veteran’s Hospital, Chicago, Illinois, USA.
4High Throughput Analysis Laboratory and Department of Molecular Biosciences, Northwestern University, Evanston, Illinois, USA.
Address correspondence to: E. Eklund, Northwestern University, 303 E. Superior Ave, Lurie 5-111, Chicago, Illinois 60611, USA. Phone: 213.503.3208; Email; e-eklund@northwestern.edu.
Find articles by Patel, P. in: PubMed | Google Scholar
1The Feinberg School of Medicine at Northwestern University, Chicago, Illinois, USA.
2The Lurie Cancer Center at Northwestern University, Chicago, Illinois, USA.
3Jesse Brown Veteran’s Hospital, Chicago, Illinois, USA.
4High Throughput Analysis Laboratory and Department of Molecular Biosciences, Northwestern University, Evanston, Illinois, USA.
Address correspondence to: E. Eklund, Northwestern University, 303 E. Superior Ave, Lurie 5-111, Chicago, Illinois 60611, USA. Phone: 213.503.3208; Email; e-eklund@northwestern.edu.
Find articles by Bartom, E. in: PubMed | Google Scholar
1The Feinberg School of Medicine at Northwestern University, Chicago, Illinois, USA.
2The Lurie Cancer Center at Northwestern University, Chicago, Illinois, USA.
3Jesse Brown Veteran’s Hospital, Chicago, Illinois, USA.
4High Throughput Analysis Laboratory and Department of Molecular Biosciences, Northwestern University, Evanston, Illinois, USA.
Address correspondence to: E. Eklund, Northwestern University, 303 E. Superior Ave, Lurie 5-111, Chicago, Illinois 60611, USA. Phone: 213.503.3208; Email; e-eklund@northwestern.edu.
Find articles by Eklund, E. in: PubMed | Google Scholar
Published November 25, 2025 - More info
JCI Insight. 2026;11(1):e190833. https://doi.org/10.1172/jci.insight.190833.
© 2026 Huang et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: January 2, 2025; Accepted: November 17, 2025
-
Abstract
Survival in chronic myeloid leukemia (CML) was dramatically improved by development of tyrosine kinase inhibitors (TKIs) directed to the BCR:ABL1 oncogene. Unfortunately, ~30% of patients with CML develop TKI resistance during prolonged treatment, with enhanced blast crisis risk. Oxidation Resistance 1 (Oxr1) regulates antioxidant pathways that detoxify reactive oxygen species (ROS) generated by the phagocyte-NADPH oxidase. In the current studies, we found that Oxr1 expression increased in hematopoietic stem and progenitor cells (HSPCs) from CML mice versus controls, decreased during TKI-induced remission, and rose during chronic phase relapse. Oxr1 has long and short isoforms, and we found increased short, but decreased long, Oxr1 in mice or humans during CML relapse. We determined that long Oxr1 prevents ROS accumulation in CML marrow, but short Oxr1 is a dominant negative. Previously, we found exaggerated and sustained emergency granulopoiesis in CML mice, with repeated episodes facilitating relapse during TKI remission. In the current studies, we found knocking down Oxr1 in murine marrow further accelerated CML progression during this physiologic stress. We found increased DNA-damage in HSPCs from these mice, including a BCR:ABL1 kinase-domain mutation found in TKI-resistant human CML. These studies suggest that long Oxr1 detoxifies ROS to decrease mutagenesis in CML, but aberrant short Oxr1 expression enhances progression.
-
Introduction
CML is defined by translocations of chromosomes 9 and 22 that generate BCR:ABL1: a tyrosine kinase oncogene (1). CML outcomes were revolutionized by BCR:ABL1-directed TKIs, with > 50% of patients achieving a 3 log BCR:ABL1 transcript reduction with these agents (2, 3). Unfortunately, ~30% develop TKI resistance during prolonged treatment, usually due to BCR:ABL1 kinase domain (KD) mutations (4). Many patients respond to switching TKI, but the risk of blast crisis (BC) increases 10× in those who do not. Conversely, 20% of patients with CML achieve a 4 log transcript reduction, half of whom can discontinue therapy without relapse. Molecular predictors of drug resistance or therapy-free remission (TFR) duration are unknown (5–7).
To define such molecular mechanisms, we transplanted mice with BCR:ABL1-transduced syngeneic bone marrow. Marrow from primary recipients in early chronic phase (CP) was transplanted into secondary recipients to provide homogenous cohorts for study. All secondary recipients achieved TKI-induced remission (≥3 log BCR:ABL1 transcript reduction), but ~25% relapsed in CP by 30 weeks (~20 human years) with half of these developing BC, similar to the clinical course of human CML (8, 9). To model therapy discontinuation, we transplanted tertiary recipients with marrow from secondary recipients with TKI-induced, 4 log transcript reduction (8, 9). Untreated, half of these mice relapsed in CP and half sustained TFR, also mimicking human CML.
In HSPCs from CML mice, we found constitutive activation of innate immune response pathways, with further increase in TKI-induced remission (9). During the innate immune response, neutrophils are rapidly produced via emergency granulopoiesis, and patients with CML with bacterial infections have profound neutrophilia (10, 11). Consistent with this, we found exaggerated and sustained emergency granulopoiesis in CML mice, with acceleration of CP relapse and BC during repeated challenges (9). Genotoxic stress is increased in emergency granulopoiesis due to S phase shortening, accelerated differentiation, and ROS production by accumulating marrow neutrophils (12, 13).
Expression of IRF8 (IFN regulatory factor 8) is less in CML-HSPCs compared with controls, with an increase during TKI-induced remission but a decrease in BC (14, 15). Irf8 reexpression delays progression in a murine CML model, suggesting it is a leukemia suppressor (14). Previously, we found that Irf8 is required to terminate emergency granulopoiesis via repression of genes encoding Fas-associated phosphatase 1 (Fap1) or Growth arrest specific 2 (Gas2) (12, 16–19). Consistent with this, decreased Irf8 in CML-HSPCs stabilized β-catenin and enhanced Survivin expression via Gsk3β inhibition by Fap1, or calpain inhibition by Gas2 (9, 20). Gas2 also stabilized Xiap1 in neutrophils (9, 20). Compared with TKI alone, relapse was delayed during therapy or after discontinuation in CML mice treated with TKI (Imatinib) plus Ym155 (Survivin inhibitor), but adding Embelin (Xiap1 inhibitor) to TKI enhances BC (9). In the current study, we identify Oxr1 as a differentially expressed gene in these treatment groups and explore mechanistic significance for CML outcomes.
Regulation of antioxidant pathways by OXR1 detoxifies mitochondrial ROS and is a major mechanism detoxifying ROS produced by the phagocyte-NADPH oxidase (21–23). Human and murine OXR1 have long and short isoforms that are expressed from different promoters and are hypothesized to have differential regulation and activity (24, 25). Neither isoform is an efficient-free radical scavenger, but long OXR1 contains domains that are not found in short OXR1 and may activate antioxidant genes and pathways. OXR1 has not been studied in hematopoiesis or leukemogenesis, but our current studies suggest that aberrant expression of OXR1 isoforms in CML affects pathogenesis.
-
Results
Oxr1 is increased in CML, normalizes in remission, but increases with CP-relapse. In a murine CML model, adding a Survivin inhibitor (Ym155) to TKI (Imatinib) prolonged remission duration during therapy or after discontinuation, but treatment with TKI plus an Xiap1 inhibitor (Embelin) increased BC (9). In the current studies, we explored mechanisms for these differences. We transplanted mice with syngeneic marrow transduced with a BCR:ABL1 expression vector (coexpressing green fluorescent protein [GFP]) and transplanted marrow from primary recipients in stable, early CP (>15,000 GFP+ neutrophils, <5% blasts) into secondary recipients to generate uniform cohorts for study (8, 9). Secondary recipients were treated as above, and GFP+Lin– marrow cells from mice in remission (≥3 log BCR:ABL1 transcript reduction) were isolated for RNA-Seq (n = 5 per cohort, individually analyzed, unpooled). Differences in pathway activities were suggested by gene ontology analysis of differential gene expression under various conditions (with an uncorrected P < 0.05, or FDR adjusted P value to less than 5% false positive).
We found the major differences GFP+Lin– marrow cells from mice treated with TKI plus Embelin compared with those treated with TKI plus Ym155 included increased expression of genes involved in positive regulation of the immune response, inflammation, or myeloid cell differentiation but downregulation of genes in pathways involved in negative regulation of the inflammatory response (30% of all differences) (Figure 1A, underlined in red).
 Figure 1
Figure 1Increased Oxr1 is associated with sustained CML remission. Secondary recipients of bone marrow from mice with CP-CML were treated with TKI alone, or TKI plus Ym155 or Embelin. RNA-Seq was performed on GFP+Lin– marrow cells during remission. (A) Gene ontology identified differential pathway activity in treatment groups. Pathways involved in regulation of the innate immune response or inflammation are underlined in red. For gene expression comparisons, an uncorrected P < 0.05 and FDR-adjusted P < 0.05 were considered to determine statistical significance. Pathway differences were indicated by Log10, n = 6 individual mice per cohort. (B) Addition of Ym155 or Embelin to TKI-treatment alters Oxr1, Kdr, and Calr expression in murine CML marrow. Venn diagram for treatment group transcriptomes. (C) Oxr1 is increased in CML and altered by TKI-treatment or relapse. Significant differences, determined by 1-way ANOVA with Tukey correction. ***P < 0.001, ****P < 0.0001. Data are shown as mean ± SD, n = 3.
Oxr1 was the only gene that decreased with the addition of Embelin but increased upon adding Ym155 compared with TKI alone (Figure 1B). Oxr1 expression was verified in independent experiments by qPCR with GFP+Lin– RNA from secondary recipients and primers common to all Oxr1 isoforms (Figure 1C). We found more abundant Oxr1 mRNA in untreated CML mice versus controls, with a decrease in TKI-induced remission or BC, and an increase in CP relapse (P < 0.0001, n = 3 by 1-way ANOVA with Tukey correction for each comparison). Oxr1 was also more abundant in mice with prolonged versus short TFR after therapy discontinuation (P = 0.0008, n = 3). Therefore, increased total Oxr1 was associated with both uncontrolled CML and sustained TFR. We performed additional studies to address this apparent contradiction.
Oxr1 knockdown shortens TFR after TKI discontinuation in CML and facilitates BC during emergency granulopoiesis. To assess the functional effect of Oxr1 on CML, primary recipients of BCR:ABL1-transduced marrow were sacrifice in CP and marrow was transduced with a combination of 3 Oxr1 specific-shRNA vectors (to knockdown both long and short isoforms versus scrambled controls) (Figure 2A). In recipients of this marrow, Oxr1 knockdown did not influence rates of TKI-induced remission, relapse in CP, or BC progression (> 10% circulating myeloid blasts). In non–TKI-treated recipients, Oxr1 knockdown did not hasten neutrophilia or BC. Oxr1 knockdown also did not alter survival in untreated CML mice, nor did it alter those in TKI-remission (P = 0.7 and P = 0.585, respectively; n = 8). As a control, we determined Oxr1-knockdown efficiency during the experiment. We compared mRNA abundance for total, short, or long Oxr1 in GFP+Lin– marrow cells from CML mice with versus without Oxr1 knockdown at 4, 12, and 20 weeks after transplant. Over these time points, Oxr1-knockdown averaged 43.2% ± 6.4% for total, 43.5% ± 5.5% for long, and 42.4% + 7.3% for short Oxr1 (P = 0.06, P = 0.38, and P = 0.27, respectively, comparing knockdown of the isoforms at the 3 timepoints [n = 3] by 1-way ANOVA with Tukey correction).
 Figure 2
Figure 2Oxr1-knockdown in murine CML marrow enhances TFR after therapy discontinuation. Recipients of BCR:ABL1-transduced bone marrow were sacrificed in CP. Marrow was harvested and transduced with vectors to express Oxr1-specific shRNAs or scrambled control shRNAs for transplant into secondary recipients, and cohorts were treated with TKI. Marrow from some secondary recipients in remission was transplanted into tertiary recipients to observe without treatment. (A) Oxr1 knockdown in CML marrow did not delay remission, enhance CP-relapse or BC, or shorten survival. Differences were determined by log rank analysis. ****P < 0.0001. Data are shown as mean ± SD, n = 10. (B) Oxr1 knockdown in CML marrow increased BCR:ABL1 transcript abundance during TKI treatment and shortened TFR after discontinuation. Significant differences in transcript abundance by 1-way ANOVA with Tukey correction. *P < 0.05 or ****P < 0.0001. Data are shown as mean ± SD, n = 6. Relapse in tertiary recipients was compared by log rank analysis. *P < 0.05, n = 10.
To investigate the contribution of Oxr1 to relapse after TKI discontinuation, cohorts of secondary recipients were sacrificed after 24 weeks of TKI treatment, and CP marrow was transplanted into tertiary recipients to observe without further treatment (Figure 2B) (8, 9). All secondary donors had a ≥ 3 log reduction of BCR:ABL1 transcripts in the marrow. However, transcripts were higher in secondary recipients with Oxr1 knockdown versus without (3 versus 4 log reduction) (P = 0.015, n = 4), and time to CP relapse was shorter in their tertiary recipients (P = 0.028, n = 10, by log rank analysis) (8, 9).
Since emergency granulopoiesis is characterized by marrow neutrophilia, and Oxr1 detoxifies ROS generated by the phagocyte-NADPH oxidase, we studied the effect of this stress on secondary recipients in TKI-induced remission (Figure 3A) (8, 9). Episodes were induced at 4-week intervals to mimic ongoing infectious challenges in human CML but not in the protected animal facility environment. Emergency granulopoiesis was induced by Alum injection to activate the Nlrp3 inflammasome: a standard and widely accepted approach (versus saline as a steady state control) (10, 11). Alum injection induces IL-1β–dependent emergency granulopoiesis, characterized by 10× increase in G-CSF over steady state, and increased expression of inflammatory cytokines TNF-α and IL-6 (26). This cytokine profile is also seen in humans with activation of the NLRP3 inflammasome by damps or pamps (damage or pathogen activated molecular patterns) during the innate immune response (26).
 Figure 3
Figure 3Oxr1 knockdown in murine CML marrow increases BC and alters cytokine expression in the microenvironment. Recipients of BCR:ABL1-transduced bone marrow were sacrificed in CP and marrow transduced with vectors to express Oxr1-specific shRNAs or scrambled control shRNAs for transplant into secondary recipients, and cohorts were treated with TKI. (A) Oxr1 knockdown in CML marrow enhanced BC during episodes of emergency granulopoiesis. CML mice in TKI-induced remission were injected every 4 weeks with Alum to induce emergency granulopoiesis, or saline as steady state control. Relapse and survival were compared by log rank analysis. *P < 0.05, **P < 0.01, ****P < 0.0001, n = 10. (B) Oxr1 knockdown increases and sustains expression of cytokines involved in emergency granulopoiesis in CML marrow. CML mice in TKI remission, with or without Oxr1 knockdown, were injected with Alum and sacrificed at 0 hours, 12 hours, or 4 days. Marrow supernatants were analyzed for IL-1β, G-CSF, IL-6 or TNF-α by ELISA. Significant differences were found by 1-way ANOVA with Tukey correction. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Data are shown as mean ± SD, n = 4.
We found that episodes of emergency granulopoiesis hastened CP relapse during TKI treatment in CML mice without Oxr1 knockdown, as anticipated based on our prior work (P = 0.002, n = 8, by log rank analysis versus steady state) (Figure 3A) (9). TKI-treated CML mice with Oxr1 knockdown developed BC more rapidly during repeated Alum injections than at steady state (P = 0.0017, n = 8, by log rank analysis) and at higher rates than mice without Oxr1 knockdown (P = 0.0033, n = 8, by log rank analysis). Oxr1 knockdown shortened survival during emergency granulopoiesis episodes (P = 0.014, n = 8, versus without knockdown, by log rank analysis). To determined efficiency of Oxr1 knockdown, we compared Oxr1 expression pretreatment and week 1 and 4 after Alum injection. We found Oxr1 knockdown averaged 42.9% ± 5.8% for total, 42.5% ± 6.2% for long, and 41.8% ± 6.6% for short Oxr1 over these timepoints (P = 0.26, P = 0.87, and P = 0.92, respectively, comparing knockdown at the 4 time points [n = 3] by 1-way ANOVA with Tukey correction).
We next studied the effect of Oxr1 knockdown on the bone marrow microenvironment, with a focus on expression of emergency granulopoiesis-related cytokines (Figure 3B). For these studies, secondary recipient CML mice in TKI-induced remission were compared with or without Oxr1 knockdown. Bone marrow supernatants were studied at steady state versus 12 hours or 4 days after Alum injection. We found mice with Oxr1 knockdown had enhanced expression of these emergency granulopoiesis-related cytokines, with delayed resumption of steady state levels. Since Oxr1 detoxifies ROS generated via the neutrophil NADPH oxidase, this suggests that ROS levels sustain emergency granulopoiesis.
Oxr1 isoform switch occurs during emergency granulopoiesis or CP relapse in CML. These data suggest that Oxr1 confers protection from CML progression. However, we found that increased Oxr1 was associated with CP relapse during TKI treatment but also with long TFR after TKI discontinuation. To investigate this apparent contradiction, we quantified long versus short OXR1 isoforms under various conditions (Figure 4A) (24). These 2 isoforms are driven by different promoters and are thought to have differential regulation. We first analyzed peripheral blood CD34+CD38– cells from patients with CML in TKI-induced remission at the time of therapy discontinuation (with ≥ 4 log transcript reduction). We found more total OXR1 mRNA in patients with CML with a subsequent long TFR (P < 0.0001, n = 7, versus short TFR by Mann-Whitney rank sum test, but short TFR was associated with a relative increase in the short Oxr1 isoform (P = 0.015, n = 7, for short versus long TFR) (Figure 4B).
 Figure 4
Figure 4Expression of the short OXR1 isoform is associated with CML relapse. (A) Schema of OXR1 isoforms in humans and mice. Primers for amplification of short versus long isoforms indicated. (B) A relative increase in short OXR1 is associated with short TFR in human CML. Total and short OXR1 mRNA was quantified in CD34+CD38– peripheral blood cells from patients with CML at therapy discontinuation. Significant differences were found by 2-tailed t test. *P < 0.05 or ****P < 0.0001. Data are shown as mean ± SD, n = 7 unique patients. (C) In TKI-treated CML mice, we found an increase in short Oxr1 that occurred during emergency granulopoiesis, with IM + Embelin versus IM + Ym155 treatment, during CP relapse, or with short or long TFR. Recipients of BCR:ABL1-transduced marrow were sacrificed in CP, and marrow was transplanted into secondary recipients. Mice in TKI-induced remission were injected every 4 weeks with Alum to induce emergency granulopoiesis, or saline steady state control, and analyzed 2 weeks later. Other mice were treated with TKI plus Embelin or Ym155. Short (aqua) versus long (white) Oxr1 mRNA was quantified in GFP+Lin– marrow cells. Significant differences were found by 1-way ANOVA with Tukey correction. ***P < 0.001 or ****P < 0.0001, or #P < 0.05, ##P < 0.01, or ####P < 0.0001. Data are shown as mean ± SD, n = 6. (D) Oxr1 protein expression reflects mRNA abundance. Western blots of GFP+Lin– marrow cells were probed with antibodies to Oxr1 or GAPDH (loading control) and densitometry performed (the experiment was repeated 3 times and a representative blot shown). (E) The ratio of short to long Oxr1 isoforms is greater in CML versus non-CML stem cells and in stem cells versus progenitors. CML mice in TKI-induced remission were compared with control non-CML mice. Bone marrow Lin–Sca1+ckit+ cells (stem cells) were compared with Lin–Sca1–ckit+ cells (progenitors). Significant differences were found by 1-way ANOVA with Tukey correction. ****P < 0.0001. Data are shown as mean ± SD, n = 4.
We also quantified Oxr1 isoforms in GFP+Lin– marrow cells from CML mice prior to treatment, in TKI-induced remission, at CP relapse, or with long versus short TFR duration (Figure 4C). Lin– marrow cells from non-CML mice were controls. Total Oxr1 in untreated CML mice was greater than control non-CML mice (P < 0.0001, n = 5), but the ratio of long to short Oxr1 was comparable. Expression of the long Oxr1 isoform in CML mice in TKI remission was not different than control mice (P = 0.66, n = 5, by 1-way ANOVA with Tukey correction), but ~2× more short isoform was expressed in the former (P = 0.045, n = 5). In CP relapse, short Oxr1 mRNA was ~4× greater than in remission (P < 0.0001, n = 5), but long Oxr1 decreased 25% (P < 0.0001, n = 5). Therefore, more long versus short Oxr1 was expressed in remission, but more short versus long with CP relapse (P < 0.0001, n = 5, by 1-way ANOVA with Tukey correction). Consistent with human results, expression of short Oxr1 in mice in TKI remission with subsequent short TFR after therapy discontinuation was greater than with long TFR (P < 0.0001, n = 5).
We also quantified Oxr1 isoforms 2 weeks after Alum injection (Figure 4C). In non-CML mice, long Oxr1 increased during emergency granulopoiesis, but short Oxr1 did not (P < 0.0001, n = 5, by 1-way ANOVA with Tukey correction). Conversely, in CML mice in TKI-remission, short Oxr1 increased ~2.5× post Alum (P < 0.0001, n = 5), but long Oxr1 was unaltered. Therefore, during emergency granulopoiesis, the ratio of long/short Oxr1 increased in non-CML mice, but the ratio of short/long increased in CML mice in remission (Figure 4C). Abundance of Oxr1 isoform proteins correlated with mRNA (Figure 4D).
We also investigated the relative abundance of long versus short Oxr1 isoforms in stem cells versus differentiating progenitor populations. For these studies, we compared Lin–Sca1+ckit+ (LSK) cells to Lin–Sca1–ckit+ cells (Figure 4E). We found no difference in the ratio of short/long Oxr1 in progenitors from CML mice in TKI-induced remission compared with control, non-CML mice. In contrast, the ratio of short/long Oxr1 was greater in LSKs from CML mice in TKI-induced remission compared with control, non-CML mice (P < 0.0001, n = 4).
Knocking down long Oxr1 or overexpressing short Oxr1 increases intracellular ROS and DNA damage in CML. ROS in CML leukemia stem cells (LSCs) may be produced by mitochondria or by other marrow cells with diffusion into LSCs. Since ROS may cause TKI-resistance via point mutations of the BCR:ABL1-KD, we further studied the cohorts of secondary CP-CML recipients in TKI-induced remission. Bone marrow GFP+ LSK cells from these mice were compared with LSKs from non-CML mice at steady state or 2 weeks after Alum injection. Intracellular total versus mitochondrial ROS and DNA damage were determined by flow cytometry.
We first compared CML mice in TKI remission to non-CML mice or CML mice in CP relapse during TKI treatment (Figure 5, A and B). In LSKs from non-CML mice, total and mitochondrial ROS increased during emergency granulopoiesis (P = 0.0005 and P = 0.017, n = 3, respectively by 1-way ANOVA with Tukey correction) and DNA damage doubled (P = 0.0061, n = 3). In Alum-injected CML mice in TKI remission, we found total ROS, mitochondrial ROS, and DNA damage were also increased compared with steady state (P = 0.016, P < 0.0001, or P = 0.0035, n = 3, respectively, with versus without Alum injection by 1-way ANOVA with Tukey correction). Total ROS, mitochondrial ROS, and DNA damage were greater in these mice compared with non-CML, control mice (P < 0.0001, n = 3, for all comparisons). However, LSKs from CML mice with CP relapse during TKI treatment had more total and mitochondrial ROS, as well as DNA damage, compared with Alum-injected CML mice in TKI remission (P = 0.0005, P = 0.0015, P = 0.0016, n = 3, respectively) (Figure 5, A and B).
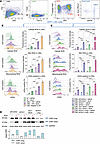 Figure 5
Figure 5Oxr1-knockdown in CML bone marrow enhances intracellular ROS and DNA damage. (A) Recipients of BCR:ABL1 transduced marrow were sacrificed in CP, and marrow transduced with vectors to express shRNAs to total Oxr1, the long Oxr1 isoform, scrambled controls, or to overexpress short Oxr1. Transduced marrow was transplanted into secondary recipients. Secondary recipients in TKI-induced remission were treated with Alum to induce emergency granulopoiesis, or saline steady state control, and sacrificed 2 weeks later. (B) CML mice have increased total cellular and mitochondrial ROS, and DNA damage during emergency granulopoiesis compared with non-CML mice, and this increases further with CP relapse. GFP+LSK cells were analyzed by flow cytometry. Significant differences were found by 1-way ANOVA with Tukey correction. *P < 0.05, **P < 0.01, ***P < 0.001, or ****P < 0.0001. Data are shown as mean ± SD, n = 3 and a representative histogram is shown. (C) Total Oxr1-knockdown, long Oxr1 knockdown, or short Oxr1 overexpression increases total intracellular and mitochondrial ROS and DNA damage during emergency granulopoiesis in CML mice, but long Oxr1 knockdown has the greatest effect. GFP+LSK cells were analyzed by flow cytometry. Significant differences were found by 1-way ANOVA. *P < 0.05, **P < 0.01 or ****P < 0.0001. Data are shown as mean ± SD, n = 3 and a representative histogram is shown. (D) Oxr1 long versus short isoform expression in these cells was verified. Western blots of GFP+Lin– marrow cells were probed with Oxr1 or GAPDH (loading control) antibody and densitometry performed (experiment was repeated 3 times and a representative blot shown).
We next studied the effect of knocking down just the long isoform in the bone marrow of CML mice. Vectors for long Oxr1 knockdown were specific to unique domains in this isoform compared with short Oxr1. To explore our hypothesis that short Oxr1 functions as a dominant negative, other CML bone marrow was transduced with a vector to overexpress this isoform. We did not knock down short Oxr1 alone since the 100 bp sequence unique to short Oxr1 did not permit design of shRNA of adequate specificity. These vectors were transduced into bone marrow from primary recipients with established CP CML prior to transplant into secondary recipients. Mice were TKI treated, as above, and studied at steady state or after Alum injection.
In CML mice in TKI-induced remission, knocking down total or long Oxr1, or overexpression of short Oxr1, did not alter total or mitochondrial ROS or DNA-damage during steady state (Figure 5, A and C). However, in Alum-injected CML mice, knockdown of all Oxr1 isoforms increased total and mitochondrial ROS as well as DNA damage (P < 0.0001, P < 0.0001, or P = 0.0009 [n = 3], respectively, versus without Oxr1 knockdown). Interestingly, total Oxr1 knockdown induced comparable levels of intracellular ROS in Alum-injected CML mice in TKI remission as found in LSKs from CML mice in CP relapse (P = 0.156, n = 3). We found specifically knocking down the long Oxr1 isoform induced a greater increase in total and mitochondrial ROS as well as DNA damage in Alum-injected CML mice (P < 0.0001, P < 0.0001, P = 0.031 [n = 3], respectively, for comparison of total versus long Oxr1 knockdown). This suggested knockdown of the short Oxr1 isoform by the total Oxr1 shRNA vector partly ameliorated effects of knocking down the long isoform. To directly address this, we performed similar studies with short Oxr1 overexpression. We found that overexpressing short Oxr1 in the marrow of Alum-injected CML mice in TKI-induced remission increased total and mitochondrial ROS, and DNA damage, compared with Alum-injected CML mice without short Oxr1 overexpression (P = 0.0005, P < 0.0001, P < 0.0001 [n = 3], respectively). Oxr1 isoform expression during emergency granulopoiesis, and efficiency of knockdown, were confirmed by Western blot (Figure 5D).
Short Oxr1 exerts dominant-negative effects via increased ROS. To determine functional contributions of Oxr1 isoforms to ROS handling and DNA damage, secondary recipients of CP-CML marrow in TKI-induced remission were injected with Alum to induce emergency granulopoiesis (versus saline as a steady state control). Marrow cells harvested 2 weeks after injection were transduced with vectors to express short or long Oxr1 (or empty vector) (Figure 6A). Oxr1 isoform overexpression was confirmed by Western blot (Figure 6B) and real-time PCR (Figure 6C) of GFP+Lin– marrow cells. In control GFP+Lin– cells, Alum injection increased short Oxr1 mRNA (P = 0.00021, n = 5, by 1-way ANOVA with Tukey correction) but not long Oxr1 mRNA. Overexpression of one isoform did not influence the other.
 Figure 6
Figure 6Short Oxr1 enhances, and long Oxr1 decreases, intracellular ROS and DNA damage in CML bone marrow. Recipients of BCR:ABL1-transduced marrow were sacrificed in CP, and marrow transplanted into secondary recipients. Secondary recipients in TKI-induced remission were treated with Alum to induce emergency granulopoiesis, or saline steady state control, and sacrificed 2 weeks later. Marrow was transduced with vectors to overexpress short or long Oxr1 isoforms, or empty control vector. (A–C) Experimental schema. Oxr1 isoform expression was verified (A), and overexpression of one isoform did not influence expression of the other isoform for protein (B) and mRNA expression (C). Western blots of GFP+Lin– cells were probed with Oxr1 or GAPDH antibodies (experiment was repeated 3 times and a representative blot shown). mRNA was quantified with isoform specific primers. Significant differences were found by 1-way ANOVA with Tukey correction. *P < 0.05, **P < 0.01, or ****P < 0.0001. Data are shown as mean ± SD, n = 5. (D) Expressing short Oxr1 increases, and long Oxr1 decreases, total and mitochondrial ROS and DNA damage. The effect of short Oxr1 is reversed by N-acetylcysteine. *P < 0.05, **P < 0.01, ***P < 0.001, or ****P < 0.0001. Data are shown as mean ± SD, n = 3 and a representative histogram is shown.
Overexpressing long Oxr1 in GFP+LSKs from Alum-injected CML mice in TKI-induced remission decreased total and mitochondrial ROS and DNA damage compared with control vector (P = 0.005, P = 0.0009, P = 0.04 [n = 3], respectively) (Figure 6D). Conversely, overexpressing short Oxr1 increased total and mitochondrial ROS and DNA damage compared with control vector, consistent with the studies above (P = 0.0048, P = 0.0019, P = 0.0002 [n = 3], respectively). To determine if the effect of short Oxr1 on DNA damage was via enhanced ROS, we treated some cells with N-acetylcysteine (NAC; a free radical scavenger). NAC treatment decreased total and mitochondrial ROS, and DNA damage in CML marrow from Alum injected mice overexpressing short Oxr1 (P = 0.0002, P = 0.0004, P = 0.0001 [n = 3], respectively, with versus without NAC). Levels of DNA damage in short Oxr1-overexpressing, NAC-treated LSKs were not different than in cells from Alum-injected CML mice without short Oxr1 overexpression.
Oxr1-knockdown and emergency granulopoiesis enhance BCR:ABL1-KD mutations in CML. To determine if increased intracellular ROS during emergency granulopoiesis favored BCR:ABL1-KD mutations, we analyzed GFP+ marrow cells of individual TKI-treated CML mice by next-generation sequencing of the KD. Such KD mutations occur in human CML and murine models and are known to be responsible for TKI resistance. We performed sequence analysis using murine marrow cells and primer sets that only amplified the KD in the context of the BCR:ABL1 fusion.
We studied CML mice during emergency granulopoiesis or steady state, and we compared recipients with or without Oxr1 knockdown in the marrow. Marrow was obtained after the second injection when 50% of Alum-injected CML mice had relapsed in CP, and 100% of CML mice with Oxr1 knockdown had relapsed in CP or progressed to BC (Figure 3A). We found the number of BCR:ABL1-KD mutations per mouse increased during emergency granulopoiesis episodes (P < 0.0001 with or P = 0.008 without Oxr1 knockdown [n = 5], by 1-way ANOVA with Tukey correction), but Oxr1 knockdown exaggerated this effect (P = 0.007, n = 5 with versus without Oxr1 knockdown) (Figure 7A). We found different KD mutation profiles during emergency granulopoiesis versus steady state, and with versus without Oxr1 knockdown. Alum-injected CML mice with Oxr1 knockdown had the most mutations, with each arising in 1 mouse except for E505K, which arose in 5 of 6 mice with Oxr1 knockdown (Figure 7B). This mutation was described in patients with CML with TKI resistance (27). Mutations were verified by Sanger Sequencing (Figure 7C).
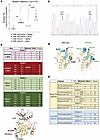 Figure 7
Figure 7Oxr1 knockdown and emergency granulopoiesis increase BCR:ABL1-KD mutations in CML. Recipients of BCR:ABL1-transduced marrow were sacrificed in CP, and marrow transduced with Oxr1-specific shRNA or scrambled control vectors was transplanted into secondary recipients. Secondary recipients were TKI treated and injected every 4 weeks with Alum to induce emergency granulopoiesis or saline as a steady state control. Marrow was analyzed 2 weeks later. (A) Oxr1 knockdown and emergency granulopoiesis cooperate to increase BCR:ABL1-KD mutations. KD-cDNA libraries from individual mice were sequenced and compared with human ABL1 sequence. Significant differences in mutations shown by 1-way ANOVA with Tukey correction. **P < 0.01 or ****P < 0.0001. Data are shown as mean ± SD, n = 4. (B) CML mice with Oxr1 knockdown in TKI-remission develop an E505K KD mutation during emergency granulopoiesis. Mutations in individual mice under various conditions are indicated. Frame-shift mutations are individual base pair losses. Ribbon diagram indicating identified mutations (labels color coded by cohort as in A). (C) Sanger sequencing confirmed specific BCR:ABL1-KD mutations. Representative sequencing shown, n = 4. (D) Mutations specific to Oxr1-knockdown plus emergency granulopoiesis block the autoinhibitory BCR:ABL1 myristoyl domain. Ribbon diagram showing most frequent mutations. (E) Patients with CML with long TFR have fewer BCR:ABL1-KD mutations than patients with short TFR. Mutations in individual patients are indicated, n = 7.
By AlphaFold analysis, we determined the identified mutations did not influence BCR:ABL1-KD secondary structure (Figure 7D). However, E505K is predicted to block myristoyl-group binding to a hydrophobic pocket in the KD: an autoinhibitory mechanism (27). For this analysis, the per-atom confidence score was > 90, indicating high confidence, and the predicted template modeling score was > 0.90, indicating high similarity of the predicted fold and true structure.
We similarly analyzed KD mutations in samples from patients with CML at the time of TKI discontinuation, using the same primer sets for KD amplification. These patients were in TKI induced remission with a ≥ 4 log reduction in BCR:ABL1 transcripts, and no KD mutations associated with TKI resistance were identified (Figure 7E). We separately analyzed patients with short versus long TFR and found an increased number of total KD mutations in those with rapid relapse. This was not unexpected, given that these patients also had a higher short/long Oxr1 isoform ratio compared with those with sustained TFR.
Oxr1 knockdown enhances mutagenesis in CML. Since BC progression involves accumulation of point mutations and chromosomal deletions or rearrangements, we analyzed the effect of Oxr1 knockdown on mutagenesis by whole exome sequencing of marrow mononuclear cells (28). We quantified mutations and clones (defined as variant allelic frequency > 12%) arising during repeated emergency granulopoiesis episodes in non-CML mice or CML mice in TKI-induced remission, with or without Oxr1 knockdown. We analyzed GFP+ (BCR:ABL1+) and GFP– (BCR:ABL1–) cells separately to assess mutagenesis in the CML clone and the effect of this clone on non-CML cells in the marrow. Marrow cells from 30-week-old non-CML mice were controls for GFP– cells in CML marrow (age match for secondary recipients), and 50-week-old non-CML mice were controls for GFP+ cells in CML mice (age match for transduced cells from initial donors).
Without Oxr1 knockdown, half of the CML mice in TKI remission relapsed in CP during 3 Alum injections, but all of the CML mice with Ox1 knockdown developed transient CP relapse with rapid BC progression (Figure 3A). In GFP+ cells from CML mice in CP relapse during emergency granulopoiesis, Oxr1 knockdown did not alter numbers of mutations (P = 0.357, n = 3 by 1-way ANOVA with Tukey correction) or clones (P = 0.107, n = 3) (Figure 8A). These mice also had no difference in clone numbers compared with non-CML mice under these conditions (P = 0.103, n = 3 for comparison of 3 groups by 1-way ANOVA) and a minor difference in mutations (P = 0.045, n = 3). CML mice with Oxr1 knockdown and BC progression had increased mutations and clones in GFP+ cells compared with CML mice with Oxr1 knockdown and CP relapse, or CML mice without Oxr1 knockdown in CP relapse (P = 0.0006 BC versus CP with or without Oxr1 knockdown, n = 3 by 1-way ANOVA for comparison of groups). Consistent with these data, mutation rates were; 7.6 ± 0.21 mutations/megabase for 50-week-old control mice; 8.4 ± 0.20 for CML mice without Oxr1-knockdown in CP relapse; 9.0 ± 0.81 for CML mice with Oxr1 knockdown in CP relapse; and 257.6 ± 99.5 for CML mice with Oxr1 knockdown in BC.
 Figure 8
Figure 8Mutagenesis in CML is increased by Oxr1 knockdown and emergency granulopoiesis. Recipients of BCR:ABL1-transduced marrow were sacrificed in CP, and marrow transduced with Oxr1-specific shRNAs or scrambled control vectors was transplanted into secondary recipients. Secondary recipients were TKI-treated and injected every 4 weeks with Alum to induce emergency granulopoiesis episodes. Whole exome sequencing performed on GFP+ versus GFP– marrow mononuclear cells, or age-matched cells from non-CML mice were identified. (A) Increased mutations and clones in GFP+ cells during BC versus other conditions and in GFP– cells from CML mice during CP relapse versus cells from non-CML mice. Significant differences in mutations were found by 1-way ANOVA with Tukey correction. *P < 0.05, **P < 0.01, or ***P < 0.001. Data are shown as mean ± SD and n = 3. (B) Distinct mutation profiles are found during CP relapse with versus without Oxr1 knockdown or with CP relapse versus BC progression. Gene ontology identified differences in pathways regulated by identified mutations. For comparisons, an uncorrected P < 0.05 and FDR-adjusted P < 0.05 were considered significant, and differences are indicated by Log10. Three mice per cohort were analyzed. (C) Mutation profiles in CML mice with Oxr1 knockdown and BC during emergency granulopoiesis are consistent with ROS-induced DNA damage. Mutation profiles were subjected to COSMIC analysis to implicate mutagenesis mechanisms. Three mice per cohort were analyzed. (D) Venn diagram demonstrating BC-unique mutations. Few common mutations were identified in CP relapse with versus without Oxr1 knockdown.
During emergency granulopoiesis, we found more mutations in GFP– marrow cells from CML mice compared with similarly aged cells from non-CML mice, and this was not influenced by Oxr1 knockdown (P = 0.002 versus with Oxr1 knockdown, or P = 0.001 without, n = 3). We also found GFP– cells from CML marrow with Oxr1 knockdown, but not without, had increased clones during emergency granulopoiesis compared with non-CML marrow (P = 0.042 non-CML versus CML with Oxr1 knockdown, or P = 0.93 non-CML versus CML without Oxr1 knockdown, n = 3).
Mutation profiles were analyzed by gene ontology to suggest an effect on related pathways (Figure 8B). In CML mice in CP relapse during TKI treatment, mutation profiles with Oxr1 knockdown were enriched for genes involved in cell cycle progression and inflammatory cytokine activity versus without Oxr1 knockdown. With BC progression, mutation profiles were additionally enriched for genes involved in receptor tyrosine kinase and Rho-GTPase signaling, RNA metabolism and catabolism, and differentiation block.
To identify possible mutagenesis mechanisms under the various experimental conditions, mutation profiles were subjected to Catalog of Somatic Mutations in Cancer (COSMIC) analysis (29). This compares mutation profiles to the COSMIC database of 6 million mutations found in 1.4 million tumor samples and curated using 25,000 publications (29). Mutations were globally enhanced in CML-BC compared with other conditions, but 2 profiles suggested unique mutagenesis mechanisms in BC: SBS22 and SBS25 (Figure 8C). The former is associated with ROS-induced DNA adducts, and the latter with exposure to alkylating or intercalating agents. Profiles with the greatest overlap were from TKI-treated CML mice with Oxr1 knockdown in CP relapse and those in BC, consistent with a continuum between the two (Figure 8D).
-
Discussion
Our studies used a murine model that mimics human CML for rates of TKI-induced remission and relapse, BC progression after development of TKI resistance, and relapse after TKI discontinuation (see; 8, 9). In this model, mice are transplanted with BCR:ABL1 p210 transduced LSKs. We study treatment outcomes in secondary bone marrow recipients from donors with established CML to address heterogeneity in rates of CML development in primary recipients. To study heterogeneity of molecular events during CP, we study multiple independent transductions and multiple primary donors. TKI-discontinuation studies are performed in tertiary recipients to track outcomes and molecular events from the same donor under multiple conditions. Relapse rates in these tertiary recipients are comparable with relapse after TKI discontinuation in secondary recipients. Outstanding investigators developed BCR:ABL1 transgenic mice, but constitutive, Sca1 promoter–driven, or CD34 promoter–driven BCR:ABL1 expression induced ALL, AML, solid tumors, or a megakaryocyte MPN rather than CML. Tet-inducible expression driven by the SCL promoter induced a CML CP like picture that was rapidly fatal but not TKI responsive nor BCR:ABL dependent in secondary recipients (30). Our approach solved these problems (8, 9).
In prior studies, we used this model to target intermediates implicated in CML apoptosis resistance (9). Compared with TKI alone, we found adding a Survivin inhibitor (Ym155) prolonged remission during TKI treatment and after discontinuation, but adding an Xiap1 inhibitor (Embelin) accelerated BC. In the current work, we investigated molecular mechanisms for these differences. By transcriptome analysis, we found increased VegfR2 and calreticulin in GFP+Lin– marrow cells from CML mice treated with TKI plus Embelin versus TKI alone, but we found decreased expression of these genes with TKI plus Ym155. Autocrine Vegf signaling occurs in BC, and calreticulin inhibits differentiation and the unfolded protein response in myeloproliferative neoplasms (31–33). Conversely, Oxr1 was increased by adding Ym155 to TKI but deceased by adding Embelin, associating it with sustained remission and protection from BC.
Oxr1 mediates detoxification of ROS generated by mitochondria or the phagocyte-NADPH oxidase (21–23). We found increased Oxr1 expression in CML HSPCs compared with HSPCs from control mice, perhaps reflecting the greater neutrophil commitment in the former. We also found an increase in total Oxr1 during emergency granulopoiesis in CML mice in TKI remission and in non-CML mice, suggesting neutrophilia induces Oxr1 expression to handle ROS generated by this process. Both long and short isoforms of human and murine OXR1 include an arginine that is oxidized by ROS, but OXR1 is a weak free radical scavenger (22). Long, but not short, human and murine OXR1 are hypothesized to have domains that activate antioxidant genes, and dominant negative activity has been proposed for short OXR1 (25). Interaction of short OXR1 with partner proteins might prevent their interaction with long OXR1, leading to a dominant negative effect. Our studies were consistent with this hypothesis.
At steady state, we found a similar ratio of short to long Oxr1 isoforms in non-CML control mice compared with CML mice, with or without TKI remission. During emergency granulopoiesis, Oxr1 increased with a constant short to long isoform ratio in control mice, but only short Oxr1 increased in CML mice during this process, suggesting dysregulation of both isoforms. Only short Oxr1 increased during CP relapse, suggesting dysregulation during inflammatory stress may contribute to progression. We also found a relative increase in short Oxr1 in CML LSKs versus non-CML LSKs, but no difference in more committed progenitors. If short Oxr1 is a dominant negative, this suggests increased susceptibility of CML LSKs to ROS-induced stress.
Consistent with a role in marrow protection, total Oxr1 knockdown enhanced intracellular ROS and DNA damage during emergency granulopoiesis in HSPCs from TKI-treated CML mice; enhancing TKI resistance and BC. Increased mitochondrial ROS was a minor component of the increase, suggesting ROS from activated neutrophils cells diffused into HSPCs. Specific knockdown of long Oxr1 had even greater effects, suggesting it is the major isoform protecting from oxidant stress. With total Oxr1 knockdown, residual expression of short and long Oxr1 were equivalent, consistent with a dominant negative effect of short Oxr1. To study this directly, we overexpressed short versus long Oxr1 in CML marrow. During emergency granulopoiesis, overexpression of short Oxr1 increased ROS and DNA damage, but long Oxr1 decreased ROS and DNA damage. Enhanced DNA damage due to short Oxr1 overexpression was reversed by the free radical scavenger NAC, implicating ROS in short Oxr1 effects.
We found enhanced and sustained levels of cytokines involved in emergency granulopoiesis in CML mice with Oxr1 knockdown, suggesting Oxr1 is negative regulator of this response. This is consistent with known ROS activation of the NLRP3 inflammasome and NF-κB (34, 35). Sustained ROS production by the phagocyte NADPH oxidase indicates unresolved infection, suggesting this mechanism is of interest for additional study.
Since ROS-induced point mutations in the BCR:ABL1-KD are a common mechanism for TKI resistance, we studied the effect of Oxr1 on such mutations. We found enhanced KD-mutations with either Oxr1 knockdown or emergency granulopoiesis with more than additive effects; both are demonstrated to increase ROS and DNA damage. Profiles of KD mutation varied by Oxr1-knockdown status and emergency granulopoiesis stress. Most mice with Oxr1 knockdown developed E505K-mutation during emergency granulopoiesis episodes. This mutation blocks autoinhibitory interaction of the BCR:ABL1 myristoyl domain with the hydrophobic ATP binding pocket and is associated with TKI resistance in patients (27). Our results therefore associate impaired handling of phagocyte-generated ROS during the innate immune response, due to aberrant Oxr1-isoform expression, with a known TKI-resistance mechanism.
We found emergency granulopoiesis episodes also enhanced clonal genomic mutations in GFP+ (BCR:ABL1-expressing) marrow cells from CML mice, with or without Oxr1 knockdown. Mice with Oxr1 knockdown rapidly progressed to BC during this stress, while 50% of those without Oxr1 knockdown relapsed in CP after 2 episodes. We found a significant increase in mutations and clones in CML mice with Oxr1 knockdown and BC progression compared with those with Oxr1 knockdown and CP relapse, as anticipated. There was also a significant increase in mutagenesis in GFP+ cells from CML mice in CP relapse versus non-CML mice. Interestingly, CML mice, but not non-CML mice, developed mutations in GFP– non-CML marrow cells during emergency granulopoiesis. This is consistent with observations that leukemia-associated mutations in human CML may involve non-CML clones and reflect sustained inflammation in CML marrow during unresolved inflammatory episodes. During BC in CML mice with Oxr1 knockdown, mutation profiles exhibited a pattern consistent with mutagenesis due to DNA adducts, providing independent confirmation of a role for aberrant ROS handling in BC pathogenesis.
Our results suggest differential regulation of short versus long Oxr1 effects on CML. The promoters for human and murine long OXR1 include IRF8 and STAT1 binding consensus sequences, but the short OXR1 promoter does not (24, 36). Decreased IRF8 and STAT1 are found in CML and may implicate these transcription factors in decreased long OXR1 expression, with no effect on short OXR1 expression, altering the isoform ratio (12, 14). Once this occurs, TKI resistance with proliferation and apoptosis resistance may favor ongoing mutagenesis and BC.
In human and murine CML, increased short OXR1 was associated with short remission after therapy discontinuation. We found an increase in various KD mutations at therapy discontinuation in patients with subsequent short TFR. Although we did not find mutations associated with TKI resistance, this finding suggests use of a different TKI for second remission induction may be a valid strategy. OXR1 isoforms are a molecular marker of interest for further study of TKI resistance, BC, and TFR duration.
Activation of innate immune response pathways and decreased IRF8 are also observed in aging, MDS, and some AML subtypes. Clarifying OXR1 regulation in CML may have implications for understanding the role of ROS in such myeloid malignancies with higher mutagenesis rates.
-
Methods
Sex as a biological variable. Equal numbers of male and female mice were used in murine studies. Male and female patients with CML were equally recruited.
Murine bone marrow transduction/transplant. 293T cells were cotransfected with p210-Bcr-abl-MiGR1 (from Ravi Bhatia, University of Alabama, Birmingham) and pCL-Eco plasmids and supernatants collected at 48 hours (8, 9). This line was obtained from American Type Culture Collection (ATCC, Manassas VA) and verified annually by STR.
C57BL/6 mice (originally obtained from Jackson Labs, Bar Harbor, ME) were injected i.p. with 150 mg/kg of 5-fluorouracil (Sigma Aldrich, St. Louis, MO, F6627) (8, 9). Marrow harvested 4 days later was incubated with retroviral supernatant (~107 pfu/mL) in DME, 10% FCS, 1% pen-strep, 10 ng/mL Il3, 100 ng/mL Scf, 10 ng/mL Il6 (R & D Systems Inc., Minneapolis MN) plus polybrene (6 μg/mL) (8, 9, 37). Transgene expression was confirmed by BCR:ABL1 PCR and GFP flow cytometry. Four independent transductions were performed with pooled bone marrow from 10 donors each. Transduced bone marrow was transplanted into recipients at a 1:1 ratio of donors to recipients. No transduced marrow or from primary recipients was frozen for use in subsequent studies.
Lethally irradiated, syngeneic recipients were injected with 1 × 106 GFP+ cells and sacrificed in CP (>15,000 GFP+ neutrophils, <5% blasts in peripheral blood [PB]). Marrow from some mice was transduced with a set of retroviral vectors to express Oxr1-specific shRNAs (2 to knockdown all isoforms; AGGTCTTTAAGTGGACAGGAGATAATATG or CTTTATCGAACAATGACAGGTTTA GACAC; and 1 to knockdown the long isoform only; TTCTAAGAAGGAAGATTTCTTTATCCA AG) or scrambled controls (Origene Technologies, Rockville, MD, TR506064), or to express cDNAs for Oxr1 short or long isoforms or vector control (Origene, MC212326, MC221322). Transduced marrow was transplanted into sublethally irradiated secondary recipients (2 × 106 cells). Each cohort included recipients from 4 different primary donors (representing 3–4 independent transductions) and PB counts were not different between groups at experiment start.
Some secondary recipient cohorts were i.p. injected with Imatinib (IM) 100 mg/kg/day; LC Laboratories, Woburn MA, I-5508) versus vehicle control a starting 4 weeks after transplant. Four weeks later, cohorts were i.p.-injected every 4 weeks with ovalbumin/aluminium chloride (Imject Alum, ThermoFisher Scientific, Waltham, MA, 77161) to induce emergency granulopoiesis versus saline as a steady state control (10–13). Other secondary recipients were i.p. injected with Imatinib IM plus Ym155 (5 mg/kg/day; Selleckchem, Houston TX, S1130), IM plus Embelin (10 mg/kg/day; Selleckchem, S7025), or saline control (9). Each cohort included recipients from 4 different primary donors (and 3–4 independent transductions), and initial peripheral blood counts were not significantly different between groups.
At 24 weeks, 2 × 106 bone marrow cells from IM-treated secondary recipients in molecular remission (≥3 log bone marrow BCR:ABL transcript reduction) were transplanted into sublethally irradiated tertiary recipients. Three tertiary recipients were transplanted with bone marrow from each of 4 different secondary recipients for each treatment cohort (each with a different primary donor). Tertiary recipients were observed without treatment.
Automated PB counts were performed on tail vein blood. Blast counts were determined on May-Grunwald-Giemsa stained peripheral smears (300 cells/slide) (8, 9). Ten mice were studied per cohort for 80% power in a 1-sided test with continuous measurement (α = 0.05) to detect differences between groups occurring at a 40% rate. All mice were analyzed without preselection and blinding was not required. Variance was significantly different between but not within groups.
ELISA. Bone marrow was collected from mice 12 hours or 4 days following injection with ovalbumin/aluminium chloride to induce emergency granulopoiesis, or saline as a steady state control. Femurs and tibias were flushed with 3 mL of phosphate buffered saline, and the BM suspensions were centrifuged at 300g at 4°C to obtain supernatants. G-CSF, IL-1β, IL-6, or TNF-α were quantified in the supernatants according to the manufacturers’ instructions with: G-CSF DuoSet (R&D Systems, DY414), IL-1β Ready-SET-Go! (ThermoFisher, 88-7013-22), TNF-α ELISA MAX Deluxe (BioLegend, 430904), or IL-6 OptEIA (BD Biosciences, 555240). A450 was measured using a BioTek microplate reader (BioTek Instruments, Winooski, VT), and cytokine concentrations calculated based on standard curves.
qPCR. RNA was isolated with RNeasy plus micro kit (Qiagen, Germantown MD, 74034). Primers were designed with Applied Biosystems software (Waltham MA) and qPCR performed with PowerUp SYBR Green Master Mix (Applied Biosystems, A25742). Results were normalized to 18S and actin. Four independent experiments were performed in triplicate.
Flow cytometry and cell sorting. Cells were analyzed on a Becton-Dickinson LSR-Fortessa flow cytometer (BD Biosciences, Franklin Lakes, NJ) and fluorescence intensities analyzed with Flowjo software (Flowjo LLC, Ashland, Oregon). Average RLU and SD of independent triplicate samples was determined. LSK cells were identified with anti-mouse V450 lineage cocktail (BD Biosciences, 561301) and PE-Cy7-Sca1, PE-ckit, or APC-ckit (eBioscience, San Diego, CA, 25-5981-82, 17-1171-82, 12-1171-82, respectively). DNA damage was assessed by anti-phospho-H2AX (S139) APC staining (eBioscience, 12-9865-42). Total intracellular or mitochondrial ROS were quantified with ROS Deep Red or Mitochondrial Superoxide Detection Kit, respectively (Abcam Inc., Cambridge, MA, ab186029 or ab219943).
Murine bone marrow cells were sorted by staining with an anti-mouse V450 lineage cocktail (BD Biosciences, 561301), APC-conjugated anti-Sca1 antibody (eBioscience, 17-5981-82), and PE-conjugated anti-Kit antibody (eBioscience, 12-1171-82). Lineage-negative (Lin–) cells were defined using a Becton-Dickenson FACSAria IIIu cell sorter. LSK (Lin-Sca1+ckit+) and Lin-Sca1-ckit+ cells were isolate from this population.
RNA, whole exome, and BCR:ABL1 KD sequencing. Stranded mRNA-Seq of GFP+Lin– marrow was conducted in the Northwestern University NUSeq Core (5/cohort, nonpooled samples). RNA quality was determined using an Agilent Bioanalyzer 2100 (Agilent Research Laboratories, Santa Clara, CA). Libraries were prepared and validated with TruSeq Stranded mRNA kit (Illumina Inc, San Diego, CA, 20020594). Single-end 75 bp reads were generated with an lllumina NextSeq 500 Sequencer; read quality was evaluated using FastQC v0.11.7 (38); adapters were trimmed; and reads of poor quality or aligning to rRNA were filtered. Cleaned reads were aligned to the Mus musculus genome using STAR and read counts calculated by htseq-count in conjunction with mm10 gene annotation file (http://genome.ucsc.edu). Differential expression was determined using DESeq2 (39). For gene expression comparisons between groups, uncorrected P < 0.05 and FDR-adjusted P value of 0.05 were determined to identify statistical significance.
To identify BCR:ABL1 mutations, KDs were amplified from cDNA by nested PCR (set 1 BCR-F: ACAGCATTCCGCTGACCATCAATAAG and ABL1-R1: ATGGTCCAGAGGATCGCTCTCT; set 2 ABL1-KD-F: TGGTTCATCATCATTCAACGGTGG and Abl-KD-R: TCCACTTCGTCTGAGA TACTGGATT). Libraries were constructed with Nextera XT DNA Library Prep Kit (Illumina, FC-131-1024), purified with SPRIselect beads (Beckman Coulter, Brea, CA, A66514), and next-generation sequenced on an Illumina HiSeq 4000 with single-ended 50 bp mode. Read quality was evaluated using FastQC v0.11.7 (38). Reads were trimmed to remove adapters using cutadapt v1.14 and aligned to the amplicon sequence using Burrows Wheeler Aligner v0.7.12 (38, 40); aligned files were cleaned, sorted, and marked for duplicates with Picard Tools v1.85; variants were detected using BCFtools mpileup with minimum base quality and maximum depth parameters set to 20 and 8,000; and mutations confirmed with Sanger sequencing.
Predicted structures of the BCR:ABL1 KD (amino acids 250–535) and 7 identified mutations were obtained by AlphaFold3.0 (41). Mutant and nonmutant sequences were uploaded separately to the AlphaFold server (AlphaFoldserver.com). Predicted secondary structure and tertiary fold exhibited a high degree of structural similarity, with excellent alignment upon superimposition. Predicted structures demonstrated a high degree of 3D homology with the crystal structure (PDB 2FO0) with nearly identical spatial arrangements of key functional groups.
For whole exome sequencing, genomic DNA was extracted from murine marrow cells using the DNeasy Blood & Tissue Kit (Qiagen, 69504). Libraries were constructed with 100 ng of QC-passed gDNA using the Illumina DNA Prep Kit with exome 2.0 Plus enrichment kit (Illumina, 20077595), hybridized to exome oligo probes (Twist Bioscience, San Francisco, CA, 102035) with unique barcodes, validated with Agilent Bioanalyzer High Sensitivity DNA kit (Agilent Technologies, 5067-4626) and Qubit DNA HS assay kit (ThermoFisher, Q32854), and sequenced on an Illumina Novaseq X Plus to generate paired-end 150 bp reads. Reads in FASTQ format were evaluated using FastQC (version 0.11.7) (38); trimmed with cutadapt (version 1.14); and aligned to the mouse genome (mm10) with Burrows Wheeler Aligner (version 0.7.12) (38). Processed BAM files were cleaned, sorted, and marked for duplicates with Picard Tools (version 1.85), and they were filtered using the TruSeq_Exome_TargetedRegions_v1.2.bed file. Variants were detected using bcftools mpileup (42) with minimum base quality and maximum depth parameters set to 20 and 1,000; VCF files were filtered for quality and depth of 20 and 10; annotated with SnpEff (version 4.3); and analyzed for mutational patterns with the MutationalPatterns package in R (43). Mutation rates were calculated as the number of high-confidence variants divided by callable bases, defined as exome target regions covered at ≥ 10× depth with base quality ≥ 20. Callable coverage was estimated at ~40–43 Mb per sample using depth-of-coverage metrics; 42 Mb was used as a standard denominator to report mutation rates in mutations per megabase (mut/Mb).
Statistics. Significance was determined 1-way ANOVA with Tukey correction using SigmaPlot (Grafti LLC). Data are reported as average ± SD with P < 0.05 considered significant. Survival/relapse differences were analyzed by Mann-Whitney rank sum test. Blood counts were analyzed by Kruskal-Wallis 1-way ANOVA on ranks.
Study approval. Approved by IRB or IACUC of Northwestern University and Jesse Brown VA. Informed consent was obtained from all patients.
Data availability. Datasets were submitted to the GEO with an accession number of GSE281856 and NCBI tracking system no. 24898598 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE281956). Values for all data points in graphs are reported in the Supporting Data Values file (see supplemental material; available online with this article; https://doi.org/10.1172/JCI190833DS1).
-
Author contributions
WH performed the majority of experiments, analyzed data, and assisted in experimental design and in writing the manuscript. BL performed experiments involving animal models and flow cytometry. LH assisted with development of animal models and experiments involving animals. CHL performed analysis of structural effects of KD mutations. PP performed statistical analysis of large datasets in conjunction with WH. EB contributed to design of statistical approaches and supervised analysis. EAE planned the project and developed the experimental approach, analyzed data, and wrote the manuscript.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- I0-CX001864, R01-CA174205, R01-DK121354 (to EAE).
- Robert H. Lurie Comprehensive Cancer Center.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2025, Huang et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(1):e190833.https://doi.org/10.1172/jci.insight.190833
-
References
- Rowley JD. The Philadelphia chromosome translocation. A paradigm for understanding leukemia. Cancer. 1990;65(10):2178–2184.
- Hochhaus A, et al. Long-term outcomes of imatinib treatment for chronic myeloid leukemia. N Engl J Med. 2017;376(10):917–927.
- Padula WV, et al. Cost-effectiveness of tyrosine kinase inhibitor treatment strategies for chronic myeloid leukemia in chronic phase after generic entry of imatinib in the United States. J Natl Cancer Inst. 2016;108(7):djw003.
- Patel AB, et al. Mechanisms of resistance to ABL kinase inhibition in chronic myeloid leukemia and the development of next generation ABL kinase inhibitors. Hematol Oncol Clin North Am. 2017;31(4):589–612.
- Ross DM, et al. Safety and efficacy of imatinib cessation for CML patients with stable undetectable minimal residual disease: results from the TWISTER study. Blood. 2013;122(4):515–522.
- Rea D, et al. Discontinuation of dasatinib or nilotinib in chronic myeloid leukemia: interim analysis of the STOP 2G-TKI study. Blood. 2017;129(7):846–854.
- Pallera A et al. NCCN Guidelines insights: chronic myeloid leukemia, version 1.2017. J Natl Compr Canc Netw. 2017;14(12):1505–1512.
- Huang W, et al. The role of Fas-associated phosphatase 1 in leukemia stem cell persistence during tyrosine kinase inhibitor treatment of chronic myeloid leukemia. Leukemia. 2016;30(7):1502–1509.
- Huang W, et al. Investigating the role of the innate immune response in relapse or blast crisis in chronic myeloid leukemia. Leukemia. 2020;34(9):2364–2374.
- Cain DW, et al. Inflammation triggers emergency granulopoiesis through a density-dependent feedback mechanism PLoS One. 2011;6(5):e19957.
- Ueda Y, et al. IL-1R type I-dependent hemopoietic stem cell proliferation is necessary for inflammatory granulopoiesis and reactive neutrophilia. J Immunol. 2009;182(10):6477–6484.
- Hu L, et al. The interferon consensus sequence binding protein (Icsbp/Irf8) is required for termination of emergency granulopoiesis. J Biol Chem. 2016;291(8):4107–4120.
- Hu L, et al. Increased Fanconi C expression contributes to the emergency granulopoiesis response. J Clin Invest. 2013;123(9):3952–3966.
- Hao SX, Ren R. Expression of interferon consensus sequence binding protein (ICSBP) is downregulated in Bcr-Abl-induced murine chronic myelogenous leukemia-like disease, and forced coexpression of ICSBP inhibits Bcr-Abl-induced myeloproliferative disorder. Mol Cell Biol. 2000;20(4):1149–1161.
- Schmidt M, et al. Expression of nuclear transcription factor interferon consensus sequence binding protein in chronic myeloid leukemia correlates with pretreatment risk features and cytogenetic response to interferon-alpha. Blood. 2001;97(11):3648–3650.
- Huang W, et al. Fas-associated phosphatase 1 mediates Fas resistance in myeloid progenitor cells expressing the Bcr-abl oncogene. Leuk Lymphoma. 2013;54(3):619–630.
- Huang W, et al. Fas-associated phosphatase 1 influences βcatenin activity in myeloid progenitor cells expressing the Bcr-abl oncogene. J Biol Chem. 2013;288(18):12766–12776.
- Huang W, et al. The interferon consensus sequence-binding protein (ICSBP/IRF8) represses PTPN13 gene transcription in differentiating myeloid cells. J Biol Chem. 2008;283(12):7921–7935.
- Huang W, etInterferon consensus sequence binding protein (ICSBP) decreases beta-catenin activity in myeloid cells by repressing GAS2 transcription. Mol Cell Biol. 2010;30(19):4575–4594.
- Huang W, et al. Decreased calpain activity in chronic myeloid leukemia impairs apoptosis by increasing survivin in myeloid progenitors and xiap1 in differentiating granulocytes. Oncotarget. 2017;8(4):50629–50641.
- Matsui A, et al. Oxidation resistance 1 functions in the maintenance of cellular survival and genome stability in response to oxidative stress-independent DNA damage. Genes Environ. 2020;42(1):29.
- Yang M, et al. Human OXR1 maintains mitochondrial DNA integrity and counteracts hydrogen peroxide-induced oxidative stress by regulating antioxidant pathways involving p21. Free Radic Biol Med. 2014;77:41–48.
- Yang M, et al. Transcriptome analysis of human OXR1 depleted cells reveals its role in regulating the p53 signaling pathway. Sci Rep. 2015;5:17409.
- Murphy KC, Volkert MR. Structural/functional analysis of the human OXR1 protein: identification of exon 8 as the anti-oxidant encoding function. BMC Mol Biol. 2012;13:26.
- Volkert MR, Crowley DJ. Preventing neurodegeneration by controlling oxidative stress: the role of OXR1. Front Neurosci. 2020;14:611904.
- Paudel S, et al. Regulation of emergency granulopoiesis during infection. Front Immunol. 2022;13:961601.
- Lee B, Shah N. Identification and characterization of activating ABL1 1b kinase mutations: impact on sensitivity to ATP-competitive and allosteric ABL1 inhibitors. Leukemia. 2017;31(5):1096–1107.
- Ochi Y, et al. Clonal evolution and clinical implications of genetic abnormalities in blastic transformation of chronic myeloid leukaemia. Nat Commun. 2021;12(1):2833.
- Bamford S, et al. The COSMIC (catalogue of somatic mutations in cancer) database and website. Br J Cancer. 2004;91(2):355–358.
- Sontakke P, et al. Modeling of chronic myeloid leukemia: an overview of in vivo murine and human xenograft models. Stem Cells Int. 2016;2016:1625015.
- Dias S, et al. Autocrine stimulation of VEGFR-2 activates human leukemic cell growth and migration. J Clin Invest. 2000;106(4):511–521.
- Yang Y, et al. The ER-localized Ca2+-binding protein calreticulin couples ER stress to autophagy by associating with microtubule-associated protein 1A/1B light chain 3. J Biol Chem. 2019;294(3):772–782.
- Shago M, et al. Modulation of the retinoic acid and retinoid X receptor signaling pathways in P19 embryonal carcinoma cells by calreticulin. Exp Cell Res. 1997;230(1):50–60.
- Abais JM, et al. Redox regulation of NLRP3 inflammasomes: ROS as trigger or effector? Antioxid Redox Signal. 2015;22(13):1111–1129.
- Morgan M, Liu Z. Crosstalk of reactive oxygen species and NF-κB signaling. Cell Res. 2011;21(1):103–115.
- MotifMap webtools. University of California, Irvine. https://motifmap.ics.uci.edu/.
- Chen Y, et al. Loss of the Alox5 gene impairs leukemia stem cells and prevents chronic myeloid leukemia. Nat Genet. 2009;41(7):783–792.
- Liu H, et al. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754–1760.
- Love MI, et al. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550.
- Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10–12.View this article via: CrossRef Google Scholar
- Abramson J, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature. 2024;630(8016):493–500.
- Li H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics. 2011;27(21):2987–2993.
- Manders F, et al. MutationalPatterns: the one stop shop for the analysis of mutational processes. BMC Genomics. 2022;23(1):134.
-
Version history
- Version 1 (November 25, 2025): In-Press Preview
- Version 2 (January 9, 2026): Electronic publication
- Version 3 (March 19, 2026): Electronic publication











