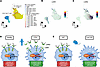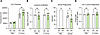Research ArticleDermatologyImmunologyOncology
Open Access |  10.1172/jci.insight.183935
10.1172/jci.insight.183935
LAIR1 prevents excess inflammatory tissue damage in Staphylococcus aureus skin infection and cutaneous T cell lymphoma
Hannah K. Dorando,1 Evan C. Mutic,1 Kelly L. Tomaszewski,2 Yulia Korshunova,1 Ling Tian,1 Mellisa K. Stefanov,1 Chaz C. Quinn,1 Deborah J. Veis,3 Juliane Bubeck Wardenburg,2 Amy C. Musiek,2 Neha Mehta-Shah,2 and Jacqueline E. Payton1
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Dorando, H. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Mutic, E. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Tomaszewski, K. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Korshunova, Y. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Tian, L. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Stefanov, M. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Quinn, C. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by
Veis, D.
in:
PubMed
|
Google Scholar
|

1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Wardenburg, J. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Musiek, A. in: PubMed | Google Scholar
1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by
Mehta-Shah, N.
in:
PubMed
|
Google Scholar
|

1Department of Pathology and Immunology,
2Department of Pediatrics, and
3Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
Find articles by Payton, J. in: PubMed | Google Scholar
Published November 13, 2025 - More info
JCI Insight. 2026;11(1):e183935. https://doi.org/10.1172/jci.insight.183935.
© 2025 Dorando et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: June 14, 2024; Accepted: November 11, 2025
-
Abstract
Patients with cutaneous T cell lymphoma (CTCL) experience high morbidity and mortality due to S. aureus skin infections and sepsis, but the underlying mechanisms remain unclear. We have previously identified high levels of LAIR2, a decoy protein for the inhibitory receptor LAIR1, in advanced CTCL. Mice lack a LAIR2 homolog, so we used Lair1-KO mice to model LAIR2 overexpression. In a model of S. aureus skin infection, Lair1-KO mice had significantly larger abscesses and areas of dermonecrosis compared with WT despite similar bacterial burdens. Lair1 KO exhibited a pattern of increased inflammatory responses in infection and sterile immune stimulation, with increased production of proinflammatory cytokines and myeloid chemokines, neutrophil ROS, and collagen/extracellular matrix (ECM) pathway proteins, including collagens and complement factors. These findings support the notion that loss of LAIR1 signaling causes an excessive inflammatory response that exacerbates tissue damage and does not improve infection control. Underscoring the clinical relevance of our findings, CTCL skin lesions exhibited similarly increased expression in cytokine and collagen/ECM remodeling pathways, suggesting that high levels of LAIR2 promote excessive inflammatory tissue damage and compromise host defense against S. aureus infection. LAIR signaling represents a promising target for therapeutic development in CTCL and other inflammatory diseases.
-
Introduction
Cutaneous T cell Lymphoma (CTCL) is a cancer of skin-homing T lymphocytes. Patients with late-stage disease have poor survival rates, and the only curative treatment is allogeneic stem cell transplant. Advanced CTCL is marked by inflammatory tissue damage in CTCL-involved skin and immune defects that lead to more severe Staphylococcus aureus (S. aureus) skin and soft tissue infections associated with high morbidity and mortality (1–3). Notably, the underlying mechanisms remain undefined (4–7).
We previously identified aberrant epigenetic regulation and high mRNA and protein expression of leukocyte-associated immunoglobulin-like receptor 2 (LAIR2) in advanced and treatment-resistant CTCL (8). LAIR2 and its paralog, LAIR1, are Ig-like receptors that bind collagens and other proteins with collagen-like domains, including complement pathway proteins such as C1q, which are important in host defense and tissue remodeling (9–12). We and others demonstrated that LAIR1 is highly expressed on macrophages and other innate immune cells; in contrast, LAIR2 is produced primarily by T and NK cells but is secreted into the extracellular space and plasma (8, 13–15). LAIR1 is a transmembrane inhibitory receptor with 2 intracellular ITIM motifs. LAIR2 acts as a decoy receptor, reducing LAIR1 inhibitory signaling and increasing activation signaling in immune cells (10). Upon binding collagen domain-containing ligands, LAIR1 signaling via SHP1 and SHP2 phosphatases negatively regulates many cellular immune activation processes, including cytokine production, calcium signaling, and differentiation (16–20).
Available evidence supports the notion that LAIR proteins likely function in inflammatory responses in infection and inflammatory diseases. Indeed, we and others demonstrated that LAIR1 and LAIR2 expression varies in cell type– and factor-specific patterns in response to pathogen- and host-derived inflammatory factors (13–15). Decreased levels of LAIR1 are associated with rheumatoid arthritis, systemic lupus erythematosus, inflammatory bowel disease, and psoriasis (21, 22). LAIR1 expression in innate immune cells (monocytes, NK cells, neutrophils) may play a modulatory role in immune response to RSV infection (21, 23, 24), severe COVID-19 (25), and pediatric malarial infection (26, 27). These reports support a role for LAIR proteins in inflammatory diseases and immune response to viral and malarial infection, but a role in bacterial infection has not been tested, to our knowledge.
We previously identified LAIR2 upregulation in patients with CTCL, a population with increased susceptibility to invasive S. aureus infection (1–3). Because mice do not have a LAIR2 gene, Lair1-KO mice effectively model loss of LAIR1 inhibitory signaling caused by high levels of LAIR2 (10, 28). We used a well-characterized mouse model of S. aureus skin infection established by the Bubeck Wardenburg lab in which mice are s.c. inoculated with a high dose of a clinically relevant isolate of methicillin-resistant S. aureus (MRSA), USA300/LAC (29), causing areas of abscess and dermonecrosis (death of overlying epithelium) that spontaneously resolve within 2 weeks in WT mice (30–32).
The results of in vivo and in vitro studies presented here demonstrate that loss of LAIR1 causes striking differences in the response to S. aureus. In skin infections, Lair1-KO mice developed much larger areas of abscess and dermonecrosis that did not resolve within the experimental timeframe, with no significant difference in bacterial burden. Lair1-KO S. aureus skin lesions also had higher levels of proinflammatory cytokines; these same cytokines were elevated in Lair1-KO bone marrow–derived macrophages (BMDMs) infected with S. aureus in vitro. Lair1-KO neutrophils produced higher levels of reactive oxygen species (ROS) yet demonstrated equivalent phagocytosis of S. aureus compared with WT. In addition, S. aureus skin lesions from Lair1 KO exhibited increased expression of genes in extracellular matrix remodeling, collagen biosynthesis and degradation, and complement immune defense pathways. The same collagen/extracellular matrix (ECM) pathways were upregulated in TLR2-activated BMDMs from Lair1 KO. Comparison of these findings to samples from patients with CTCL with advanced disease revealed that many of the same proinflammatory cytokines and collagen/ECM factors were also significantly elevated in CTCL skin biopsies. Taken together, our findings provide insights into the role of LAIR1 inhibitory signaling as an important modulator of immune response during S. aureus infection that prevents excess tissue damage. These studies also provide further support for LAIR2’s role in promoting the immune dysfunction and immunopathology observed in CTCL.
-
Results
LAIR2 is elevated in CTCL tumor cells. We previously reported that high levels of LAIR2 mRNA and protein are associated with advanced and therapy-resistant cutaneous T cell lymphoma (CTCL) (8). LAIR2 is a decoy receptor that disrupts inhibitory signals from its paralog, LAIR1, which normally suppresses inflammatory signaling in immune cells (10). Inflammatory damage to skin and underlying soft tissue, as well as deficits in cellular immunity, are hallmarks of CTCL and are associated with increased incidence of S. aureus skin and soft tissue infections (1–3, 8). We therefore hypothesized that reduced LAIR1 inhibitory signaling promotes inflammation in CTCL and alters immune response to S. aureus infection.
To begin to test this hypothesis, we first examined single-cell RNA-Seq (scRNA-Seq) from 16 serial peripheral blood samples from patients with CTCL. CD4+ cells were enriched from peripheral blood mononuclear cells (PBMCs) to enhance capture of CTCL cells, which are CD4+ in the vast majority of patients (33). We used 10x Genomics Chromium Single Cell 5′ Gene Expression and V(D)J Immune Profiling (v1.0) to generate expression and TCR (T cell receptor) repertoire profiles. In total, 92,496 cells passed quality control filters; annotation of cell type was performed using the Human Primary Cell Atlas (HPCA) data set as reference (34). Of 92,496 cells from all samples, the majority were CD4+ T cells (76,924); monocytes, which are also CD4+, comprised most of the remaining cells (Figure 1A and Supplemental Figure 1A; supplemental material available online with this article; https://doi.org/10.1172/jci.insight.183935DS1). Automated cell annotations were confirmed using canonical marker genes: pan T cell CD3 (CD3D), mature T cell (TRBC2), helper (CD4) versus cytotoxic T cell (CD8A, CD8B), CTCL cell (TOX, KIR3DL2, PLS3), and monocyte (CD14, LYZ, S100A9) (35, 36) (Supplemental Figure 1, B–D). Cell type assignments show a clear separation of CD4+ non–T cells, most of which are monocytes, and T cells (Figure 1A). In accordance with our previous work (8, 13), we found that LAIR1 was expressed primarily in myeloid cells (Figure 1B), while LAIR2 is expressed highly in CTCL T cells (Figure 1C). Thus, we hypothesized that LAIR2 secreted from CTCL cells blocks LAIR1 inhibitory signaling in myeloid cells, causing increased inflammation and associated tissue damage, which negatively affect host defense and create a permissive environment for invasive S. aureus infection (Figure 1D). To test the role of LAIR proteins in S. aureus skin infection, we used Lair1-KO mice to achieve loss of LAIR1 signaling because mice do not have a LAIR2 homolog (Figure 1E) (10, 28).
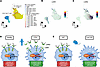 Figure 1
Figure 1LAIR2 is overexpressed in CTCL and reduces LAIR1 signaling, analogous to Lair1 KO. (A–C) CD4+ cells were isolated from CTCL patient peripheral blood and analyzed by single cell RNA-Seq. UMAP projections show 92,496 mononuclear cells from 16 peripheral blood samples from 6 patients with CTCL across multiple time points. (A) Cell type assignment by highest spearman rho of purified immune populations in the Human Primary Cell Atlas. (B and C) UMAP plots show expression of LAIR1 (B) and LAIR2 (C). (D) Illustration depicts hypothesis that LAIR2 overexpression reduces LAIR1 signaling in CTCL. (E) Illustration depicts mouse model for loss of LAIR1 signaling via KO of Lair1.
LAIR1 is protective in S. aureus skin infection. We employed a well-established mouse model of S. aureus skin infection (30–32) in which WT and Lair1-KO mice were inoculated s.c. with 1 × 107 CFU S. aureus USA300/LAC. In WT mice, this results in a self-limited skin infection that resolves spontaneously over the course of 2 weeks. The infectious lesions consist of an abscess, which has a collagen-rich capsule generated within 1 day postinfection (dpi) that serves to contain the bacteria, leukocytes, and debris. Dermonecrosis, defined as death of epithelium overlying the abscess, develops at 2 dpi in WT mice. Strikingly, Lair1-KO mice develop abscesses that are 2-fold larger on average (Figure 2A) and areas of dermonecrosis that are 2-fold larger and manifest more rapidly (by 1 dpi; Figure 2B) compared with WT mice. Differences in abscess and dermonecrosis sizes were seen beginning at 1 dpi and persisted throughout the experimental timeline, with the greatest differences at 1–8 dpi. This phenotypic difference early in infection suggests a predominant role for myeloid cells. Additionally, while WT mice completely resolve the infection by 14 dpi, an average of over 10% of lesion area remained in Lair1 KO at 14 dpi. Representative gross images of S. aureus abscess and dermonecrosis at 4 dpi (Figure 2C) and H&E staining of tissue excised from mice with S. aureus skin infections at 2 dpi (Figure 2D) demonstrate the larger areas of abscess and dermonecrosis in Lair1 KO compared with WT. Surprisingly, S. aureus CFU recovered from skin lesions at 2 dpi or from spleen at 2 or 5 dpi was not statistically different in Lair1 KO compared with WT (Supplemental Figure 2, A and B). Together, these findings suggest that absence of LAIR1 in S. aureus skin infection causes greater tissue damage yet does not meaningfully affect bacterial clearance.
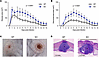 Figure 2
Figure 2LAIR1 is protective in S. aureus skin infection. WT and Lair1-KO mice were infected s.c. with 1 × 107 CFU S. aureus USA300 LAC. (A and B) Lesions were monitored over 14 days and quantified as abscess (A) and dermonecrosis (B) area (noninvasive measurements, area calculated: A = ([pi/2] × l × w). Statistical analysis by repeated measures 2-way ANOVA with Bonferroni post hoc tests. Results are representative of 5 independent experiments (n = 40 WT, n = 40 Lair1 KO). (C) Representative gross images of S. aureus skin lesions in WT (left) and Lair1-KO (right) mice on 4dpi. Abscess boundaries are indicated with the dashed lines. Dermonecrosis is the central erythematous lesion. (D) Representative H&E stains of lesions at 2 dpi. Scale bars: 1 mm. Asterisks indicate abscesses, and solid lines indicate areas of dermonecrosis, which are larger in Lair1 KO.
α-Hemolysin (Hla) is a pore-forming S. aureus virulence factor that causes lysis of host cells. Isogenic hla-negative S. aureus strains caused significantly smaller skin lesions with little to no dermonecrosis compared with WT strains (32, 37). To determine whether the larger abscess and dermonecrosis areas in Lair1-KO SSTI were dependent on Hla-mediated tissue damage, we used an isogenic hla-negative strain of USA300/LAC (Δhla) (37). We inoculated each WT and Lair1-KO mouse with USA300/LAC on 1 flank and Δhla USA300/LAC on the opposite flank and compared abscess area (Supplemental Figure 2, C and D) and dermonecrosis area (Supplemental Figure 2, E and F) over 2 weeks. Overall, infection with the Δhla strain reduced the abscess and dermonecrosis sizes in both WT and Lair1 KO compared with the WT strain. However, Δhla abscesses in Lair1 KO were larger than WT mice with WT or Δhla infections (Supplemental Figure 2C). Moreover, areas of dermonecrosis in Δhla-infected Lair1 KO were similar to WT infections in WT mice and were larger than Δhla infections in WT mice (Supplemental Figure 2E).
Statistical analysis of these results confirmed that there are significant differences due to S. aureus strain and genotype. A 3-way ANOVA was conducted to compare the main effects of genotype and S. aureus strain as well as their interaction effects on either size of abscess (Supplemental Figure 2D) or dermonecrosis (Supplemental Figure 2F). Effect of genotype (WT vs. Lair1 KO) was statistically significant for both abscess and dermonecrosis at P < 0.0001. Effect of S. aureus strain (USA300 vs. Δhla) was statistically significant for both abscess (P = 0.0008) and dermonecrosis (P < 0.0001). The 2-way interaction effect for genotype and S. aureus strain was not significant for abscess but was significant for dermonecrosis at P = 0.0249, indicating that the effect of genotype on dermonecrosis depended on S. aureus strain. To determine significant differences, 2-way ANOVA with Bonferroni adjustment for α significance level were used as post hoc tests to determine significant differences within the dermonecrosis 2-way interaction effect: dermonecrosis is significantly greater in KO than WT when using USA300 strain (P = 0.0004) and showed a nonsignificant trend toward a larger size in KO versus WT when using Δhla strain (P = 0.0692). This result was expected due to the overall low amount of dermonecrosis in Δhla infection. Taken together, these data demonstrate that the increased abscess and dermonecrosis sizes in S. aureus–infected Lair1-KO mice were not solely due to the activity of Hla. Therefore, we next sought to define host-derived differences in response to S. aureus infection.
Lair1-KO mice produce higher levels of proinflammatory cytokines in skin lesions. Because we observed larger lesions and more tissue damage in S. aureus SSTI in Lair1 KO with no difference in bacterial burden, we hypothesized that Lair1-KO mice produce increased proinflammatory cytokines. We therefore measured production of 32 cytokines and growth factors from skin punch biopsies from WT and Lair1-KO SSTI at 18 hours after infection by multiplex cytokine array. Lair1 KO skin had an overall pattern of increased proinflammatory cytokine expression (Figure 3A), with higher protein levels of 5 cytokines compared with WT (Figure 3, B–F). Neutrophil-activating chemokines CXCL1 and CXCL2 are both significantly increased, with CXCL1 elevated by more than 100% and CXCL2 elevated by about 8% on average (Figure 3, B and C). Monocyte-activating chemokines CCL2 (MCP1) and CCL3 (MIP1α) were both elevated by greater than 60% on average (Figure 3, D and E). IL-1β, which is classically associated with response to bacterial infection, was increased by over 90% on average (Figure 3F).
 Figure 3
Figure 3Lair1-KO mice produce higher levels of proinflammatory cytokines. (A–F) Skin lesions were collected at 18 hours after infection, homogenized, and cytokine production measured by multiplex cytokine array. (A) Heatmap shows average WT and Lair1 KO z-scored cytokine concentrations. (B–F) Bar graphs show all cytokines with significantly different expression between WT and Lair1 KO. (G–L) BMDMs were generated in vitro and treated with PBS or S. aureus at a multiplicity of infection (MOI) of 10:1 for 18 hours. Cell supernatant was analyzed by multiplex cytokine array. (G) Heatmap shows average WT and Lair1-KO cytokine concentrations in PBS and S. aureus groups. (H–L) Bar graphs show all cytokines with significantly different expression between WT and Lair1 KO (n = 2 independent infections, 2–6 mice per group). (B–F and G–L) Two-way ANOVA with Fisher’s LSD for multiple comparisons. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
Because dermal macrophages express high levels of LAIR1 and are initial sensors of skin infection (14, 38), we hypothesized that macrophages were responsible for the pattern of increased cytokine production in skin infection at 18 hours after infection. To test this hypothesis, we cultured BMDMs from WT and Lair1-KO mice and measured cytokine production after PBS treatment or S. aureus infection using the same cytokine array method. We again observed a significantly increased overall pattern of cytokine expression in Lair1 KO (Figure 3G), with 5 significantly elevated chemokines, 3 of which were also increased in skin infection: CXCL1, CXCL2, and CCL2 (Figure 3, H–L). CXCL1 and CXCL2 were both increased by over 90% on average (Figure 3, H and I). Strikingly, CCL2 levels in Lair1 KO were over 10-fold higher than WT in both PBS- and S. aureus–treated conditions (Figure 3K). CXCL10 was also significantly higher at baseline in Lair1 KO (Figure 3J). Basal expression of several other chemokines and cytokines, including CCL3, CCL4, CXCL2, and VEGF, were nonsignificantly elevated at baseline in Lair1 KO, suggesting that loss of LAIR1 releases inhibition of macrophage proinflammatory pathways at rest and may predispose cells to an excessive inflammatory response. Significant increases in proinflammatory cytokines that can recruit and activate multiple immune cell types, including monocytes and macrophages, were detected: CXCL10 was 3-fold elevated (Figure 3J) and CCL4 was elevated by over 130% (Figure 3L) (39, 40). In summary, these data confirm our hypothesis that loss of LAIR1 inhibitory signaling in macrophages results in greater production of proinflammatory cytokines and support the notion that dermal macrophages are at least partially responsible for increased proinflammatory cytokine levels in Lair1-KO skin abscess.
Collagen and ECM remodeling pathways are elevated in Lair1-KO skin abscess and macrophages. Our results presented thus far demonstrate that Lair1 KO macrophages produce high levels of proinflammatory mediators at baseline and in SSTI, leading us to hypothesize that the larger skin lesions and tissue damage in SSTI were due to excess inflammation. To begin to test this, we compared the transcriptomes of WT and Lair1-KO S. aureus SSTI skin biopsies and WT and Lair1-KO BMDMs treated with PBS or PAM3CSK4. PAM3CSK4 is a synthetic agonist of TLR2, which is stimulated in infection by bacterial lipoproteins, such as S. aureus lipotechoic acid, to induce NF-κB pathways (41–43). We defined upregulated genes in RNA-Seq from Lair1-KO BMDMs and Lair1-KO S. aureus SSTI compared with WT and used GSEA to identify the top 20 pathways in these genes. Surprisingly, the top 20 Reactome pathways for both BMDMs and skin abscesses included 4 collagen-related extracellular matrix remodeling pathways: “collagen degradation”, “assembly of collagen fibrils and other multimeric structures” (denoted “collagen fibril assembly”), “collagen formation”, and “collagen biosynthesis and modifying enzymes” (denoted “collagen biosynthesis”) (Figure 4, A and B). Notably, there is substantial overlap in the genes in these pathways. Therefore, we combined them into a single set of 98 genes and plotted a heatmap of z-scored expression for BMDMs (Figure 4C) and S. aureus SSTI (Figure 4D) with genes grouped according to function: collagen, collagen synthesis, and ECM remodeling. In BMDMs and SSTI, Lair1 KO exhibit a clear pattern of increased collagen-associated gene expression in response to PAM3CSK4 and S. aureus. Several collagen genes are also elevated in Lair1-KO BMDMs at baseline (KO + PBS control treatment; Figure 4C), providing further evidence that loss of LAIR1 alters the macrophage phenotype prior to inflammatory insult.
 Figure 4
Figure 4Collagen pathway gene expression is elevated in Lair1 KO S. aureus–infected skin and macrophages. (A and C) BMDM from WT and Lair1-KO mice were treated with PBS or PAM3CSK4 for 6 hours and were then subjected to RNA-Seq. (B and D) Skin punch biopsies from WT and Lair1-KO mouse S. aureus skin infections were taken at 2 dpi and were then subjected to RNA-Seq. GSEA was used to define the top 20 mouse Reactome pathways with enrichment scores and nominal P value in genes upregulated in Lair1 KO in BMDM treated with PAM3CSK4 (A) and 2 dpi SSTI (B); collagen-related pathways common to both analyses are bolded in teal. Genes from the 4 pathways, many of which overlapped, were classified into 3 general categories: collagen genes, collagen synthesis genes, and extracellular matrix (ECM) remodeling genes. Heatmaps show z-scored expression for these genes for BMDM (C) and S. aureus SSTI (D). (E and F) BMDM from WT and Lair1-KO mice were treated with PBS or PAM3CSK4 for 24 hours and were then fixed and stained for Serpin H1. Representative images from 3 independent experiments are shown (E). Scale bars: 50 μm. Images were analyzed in CellProfiler and mean Serpin H1 cytoplasmic fluorescence intensity per cell was calculated (F). Mean ± SEM are indicated by black bars. Statistical analysis was done by 2-way ANOVA with multiple comparisons.
To further support these findings, we verified increased protein levels of collagen pathway genes that had significantly increased mRNA expression in Lair1-KO BMDM and skin treated with PAM3CSK4 and S. aureus. As shown in Figure 4, E and F, we used immunofluorescence to evaluate protein levels of Serpinh1, a molecular chaperone that is essential for the proper folding and assembly of collagen. We used CellProfiler to measure average cell fluorescence intensity of Serpin H1 protein in Lair1-KO and WT BMDM treated with PAM3CSK4. These studies revealed that Serpin H1 protein expression is 1.35-fold significantly increased (P < 2.22 × 10–16) in PAM3CSK4-treated Lair1 KO compared with WT (Figure 4F). In addition, we measured Collagen I by Western blot in BMDM treated with PAM3CSK4, which demonstrated 2.5-fold higher levels in Lair1-KO BMDM compared with WT (Supplemental Figure 3, A and B). Consistent with mRNA expression, protein levels of Collagen I are elevated at baseline (PBS control) and with PAM3CSK4 treatment in Lair1 KO compared with WT. We additionally evaluated thickness of the collagen capsule in frozen sections from WT and Lair1-KO skin abscesses stained with Gomori trichrome, which stains collagen fibers (Supplemental Figure 3, C–E). While not statistically significant, there is a trend toward thicker collagen capsule walls in Lair1 KO; moreover, the overall abscess areas were significantly larger in Lair1 KO (Figure 1), meaning total abscess wall circumference was correspondingly larger in Lair1 KO. The larger size would necessitate greater production of collagen and associated ECM remodeling enzymes to achieve, which is consistent with the RNA-Seq, Western blot, and immunofluorescence results. Collagens are also defense molecules, and the increased expression of these pathways may be another facet of the enhanced inflammatory phenotype caused by loss of LAIR inhibitory signaling.
Because LAIR1 also binds collagen domain-containing complement and lectin proteins (10, 15), which are involved in bacterial defense, we next investigated whether these LAIR1 ligands were differentially expressed in Lair1-KO PAM3CSK4-treated BMDM and skin abscess. We first determined that C1qa, C1qb, and C1qc were the only members of this group that had detectable expression (averaged normalized counts > 100 for at least 1 group). In BMDMs, C1qa, C1qb, and C1qc exhibited significantly higher expression in Lair1-KO cells versus WT when stimulated with PAM3CSK4 (Supplemental Figure 3F). In addition, levels of C1qa and C1qc in murine skin abscess were significantly higher in Lair1 KO compared with WT (Supplemental Figure 3G). Together, these findings demonstrate that Lair1-KO cells produce higher levels of collagens and collagen defense molecules when infected with S. aureus or stimulated by TLR2 ligands, providing what we believe to be novel insights into the mechanism by which loss of LAIR1 causes a hyperinflammatory phenotype.
Lair1 KO produce higher levels of ROS, but phagocytosis of S. aureus is equivalent. We hypothesized that Lair1-KO neutrophils may also be hyperactivated in S. aureus infection because CXCL1 and CXCL2 were elevated in S. aureus infected skin. To test this hypothesis, we isolated neutrophils from WT and Lair1-KO bone marrow, treated them with S. aureus, and measured ROS production. Compared with WT, Lair1-KO neutrophils treated with S. aureus produced significantly more hydrogen peroxide (Figure 5A), a major component of ROS critical for killing bacteria. Hydrogen peroxide can also cause extensive tissue damage (44, 45). We next assessed intracellular lysosome acidification because it is an important step in bacterial killing, and SHP1 plays a role in regulating this process (46, 47). We incubated neutrophils with S. aureus and Lys-NIR probe and measured fluorescence by flow cytometry. We did not detect any difference in lysosome acidification between WT and Lair1-KO neutrophils (Figure 5B). To determine whether there were differences in phagocytosis between WT and Lair1-KO neutrophils, we incubated neutrophils with either S. aureus bioparticles (Figure 5C) or live mCherry-expressing S. aureus (Figure 5D); we then measured S. aureus fluorescence in washed cells by flow cytometry. Neither measure was significantly different in WT and Lair1-KO neutrophils.
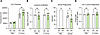 Figure 5
Figure 5Lair1-KO neutrophils produce increased reactive oxygen species but phagocytosis is equivalent to WT. WT and Lair1-KO neutrophils were isolated from mouse bone marrow. (A) Amplex red fluorescence measured by plate reader indicates hydrogen peroxide (H2O2) production in neutrophils treated with PBS or S. aureus at MOI 50:1 for 30 minutes. (B) Ratio of WT/KO lysosome probe MFI measured by flow cytometry indicates lysosome acidification in neutrophils treated with PBS or S. aureus at MOI 100:1 for 30 minutes. (C) Neutrophils incubated for 30 minutes with PBS or S. aureus bioparticles (pHrodo) that fluoresce when acidified in the lysosome were subjected to flow cytometry. (D) Neutrophils incubated for 30 minutes with PBS or mCherry-expressing S. aureus at MOI 50:1 were subjected to flow cytometry. mCherry MFI is shown as a ratio of KO to WT. Statistical analysis for all panels was done by 2-way ANOVA with Fisher’s LSD. **P < 0.01.
Monocyte-recruiting and -activating cytokines, particularly CCL2, were also elevated in Lair1 KO (Figure 3); therefore, we also assessed lysosome acidification and phagocytosis in monocytes. Classical monocytes isolated from WT and Lair1-KO bone marrow were assessed using the same methods as for neutrophils (Figure 5). Like neutrophils, monocytes exhibited similar levels of phagocytosis and lysosome acidification in Lair1 KO and WT (Supplemental Figure 4, A and B). Together, these data demonstrate that Lair1-KO neutrophils produce higher levels of ROS in SSTI, but neutrophil and monocyte phagocytosis and lysosome acidification are similar to WT. These data suggest that higher levels of ROS produce excess tissue damage in Lair1-KO skin infection yet do not enhance bacterial clearance.
Cytokines elevated in Lair1 KO are also higher in advanced CTCL. To determine whether the cytokines that were elevated in Lair1 KO were also increased in CTCL, we evaluated 3 publicly available RNA-Seq datasets from CTCL lesional skin biopsies (7, 48, 49). We examined cytokine expression for all the significantly increased cytokines in Lair1-KO BMDM and SSTI. Skin biopsies were collected from healthy donors and a range of CTCL stages, including: healthy donor, early stage CTCL, and late stage CTCL (Figure 6A) (7); healthy donor, CTCL plaques, or CTCL tumors/patches (Figure 6B) (48); or paired healthy and lesional skin from the same donors (Figure 6C) (49). Notably, none of the biopsies contained active infection. CXCL1, CXCL10, and CCL2 were significantly higher in at least 2 of 3 datasets; CCL3 and CCL4 were significantly elevated in 3 of 3 datasets evaluated. CXCL2 and IL-1b were not elevated in any of the 3 datasets we evaluated, though other proinflammatory cytokines associated with CTCL pathogenesis, including CCL5, CXCL9, IL-15, TNF, and IFN-g, were also elevated (Supplemental Figure 5, A–C). Overall, CTCL-involved skin exhibited a pattern of significantly increased expression of cytokines involved in myeloid cell recruitment and activation.
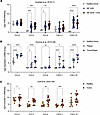 Figure 6
Figure 6Cytokines elevated in Lair1 KO are also higher in CTCL. (A–C) Publicly available CTCL skin biopsy RNA-Seq from ref. 7 (A), ref. 48 (B), and ref. 49 (C) were analyzed using DEseq2. Normalized expression for CCL2, CCL3, CCL4, CXCL1, and CXCL10 is shown. Statistical analysis for all panels was done by DEseq adjusted P value. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
We next evaluated these 3 datasets for expression of the genes in collagen pathways that were elevated in Lair1-KO S. aureus–infected skin and macrophages. Pathway analyses of upregulated genes identified significantly enriched gene sets in immune activation pathways such as CD28 costimulation, IL-3/IL-5/GM-CSF signaling, and IL-10 signaling, though no collagen/ECM remodeling pathways (Supplemental Figure 6, A–C). However, we found that individual genes in these pathways were significantly increased: 5 ECM remodeling enzyme genes were elevated in 3 of 3 CTCL datasets (CTSS, MMP1, MMP12, CTSB, and MMP9), and 3 collagen genes were elevated in 2 of 3 CTCL datasets (COL4A4, COL6A6, COLGALT2) (Supplemental Figure 6, D–F). These findings are not surprising, given that the biopsies did not involve active bacterial infection or abscess but, rather, ongoing inflammation associated with the cutaneous lymphoma microenvironment (7, 48, 49). In summary, like Lair1-KO S. aureus skin abscess and infected BMDMs, CTCL skin lesions exhibit an overall pattern of increased expression of inflammatory factors, including myeloid cell–recruiting and –activating cytokines and some collagen/ECM remodeling pathway genes.
-
Discussion
Our studies presented here reveal roles for LAIR1 inhibitory signaling in CTCL and S. aureus bacterial infection that, to our knowledge, have not been previously shown. In CTCL, a cutaneous cancer that exhibits striking inflammatory tissue damage, including ulceration, S. aureus skin and soft tissue infections are common and serious contributors to morbidity and mortality (1–3, 8). We previously identified high levels of LAIR2, a decoy receptor for the immune inhibitor LAIR1, in advanced and therapy-resistant CTCL (8). In studies presented here, we expand upon those findings and demonstrate that loss of LAIR1 signaling causes high levels of proinflammatory cytokines, bacterial defense molecules, and inflammatory tissue damage in S. aureus infection. Similar elevations were detected in cytokines and complement pathways in CTCL samples. Together, these results demonstrate that LAIR1 is an important regulator of the host immune response and that loss of LAIR1, as in excessive LAIR2 production in CTCL, causes greater inflammation and tissue damage, compromising the host response to S. aureus infection.
CTCL progression is marked by skin inflammation and barrier breakdown, which culminate in frequent S. aureus skin and soft tissue infections. LAIR1 had been previously shown to dampen activation in diverse immune cell subsets (9, 12, 18, 50). Based on overexpression of LAIR1 decoy receptor LAIR2 in patients with CTCL with progressive disease (8), we hypothesized that loss of LAIR1 inhibitory signaling may cause increased inflammation in CTCL skin lesions. Inflamed skin has long been recognized as a risk factor for S. aureus infection, particularly in CTCL (51, 52). We observed a striking phenotype in Lair1-KO mice; they suffered larger skin lesions with more tissue damage, as well as higher levels of proinflammatory cytokines and immune defense molecules in S. aureus infection. It is well established that effective host response to infection requires a fine balance between too little inflammation (inadequate response to control microbial proliferation) and too much inflammation (causing excess damage to host tissues) (53–55). This excess inflammation does not enhance pathogen clearance but, rather, impairs host immune responses and creates opportunities for microbes such as S. aureus to invade damaged tissues, particularly in CTCL (1, 2, 56, 57). Inhibitory immune pathways, such as LAIR signaling, are an essential component of immune balance and enable nuanced control of immune activation (58). Our results are consistent with this notion and demonstrate that LAIR signaling is an important molecular mechanism underlying the inflammatory tissue damage and increased incidence of S. aureus infection in CTCL.
LAIR2 is produced primarily by NK and T cells, including CTCL tumor cells, and is secreted, thereby blocking LAIR1 signaling in other cells (8, 10, 59). Innate immune cells, including monocytes and macrophages, express high levels of LAIR1 (13) and, thus, would be affected by LAIR2 blockade. Innate immune cells are the first responders in acute bacterial infection. Therefore, we predicted that reduced inhibitory signaling in these cells due to LAIR1 loss would result in an early and excessive inflammatory response to acute bacterial infection. Consistent with our prediction, our studies here show that Lair1 KO exhibit increased proinflammatory cytokine levels and larger areas of abscess and dermonecrosis by 1–2 days after S. aureus infection. While other studies showed an effect of LAIR1 on viral and malarial infection (24, 25, 27), our studies are the first to our knowledge to demonstrate that LAIR1 plays a role in modulating inflammatory response to bacterial infection.
Proinflammatory cytokine production in S. aureus–infected Lair1-KO BMDMs mirrored that of infected skin, suggesting that skin macrophages may be responsible, at least at early infection time points. This notion is further supported by the fact that most are myeloid chemokines, including CXCL1 and CXCL2, which serve to recruit neutrophils from the peripheral blood to the site of infection, resulting in abscess formation (60). Indeed, Lair1 KO had increased abscess size, likely due to increased neutrophil recruitment and influx. These results are consistent with a study in Lair1-KO lung that showed increased neutrophil recruitment upon oropharyngeal administration of CXCL1 and RSV infection compared with WT, although RSV infection did not cause increased cytokine production in Lair1 KO in this study (61). Our findings thus demonstrate what we believe to be a novel role for LAIR1 in regulating cytokine production and neutrophil recruitment in bacterial infection.
Given these results, we next sought to determine if loss of LAIR1 alters relevant functions of neutrophils or monocytes. ROS production was higher in Lair1-KO neutrophils compared with WT, though phagocytic ability in neutrophils and monocytes was equivalent to WT. ROS production causes bacterial as well as host cell death and, if unchecked, leads to substantial tissue injury (62, 63). The larger areas of dermonecrosis in Lair1 KO may be due, at least in part, to increased ROS production.
Additionally, there are no other studies to our knowledge that demonstrate increased expression of complement, collagens, and ECM remodeling genes in Lair1-KO skin infection and BMDMs. Complement proteins are important in host defense against microbial pathogens (64). Collagens and enzymes involved in production and degradation of the ECM are essential for abscess formation (60). Complement component C1q and collagen proteins are ligands of LAIR1. These findings raise the intriguing possibility of a role for LAIR1 in the setting of infection, in which collagen breakdown and complement consumption decrease the pool of LAIR1 ligands, which reduces LAIR1 inhibitory signaling, in turn increasing production of C1q and collagen via a feed-forward mechanism.
Our studies revealed several similarities in CTCL biopsy tissue and Lair1-KO S. aureus infection. Both exhibit increases in macrophage- and monocyte-activating cytokines, including CXCL10, which has previously been linked to early CTCL pathogenesis (65–67). Neutrophil infiltration in CTCL is rare (68, 69), and accordingly, we did not find similar increases in neutrophil activating cytokines in CTCL. Notably, the CTCL skin biopsies evaluated in this study did not harbor active S. aureus infection. Overall, our results support that inhibition of LAIR1 signaling via LAIR2 overexpression causes macrophage-initiated inflammatory tissue damage and S. aureus susceptibility in CTCL.
Particularly intriguing are our findings that some proinflammatory cytokines and collagen pathway genes have higher basal expression levels in Lair1-KO macrophages. These results suggest that loss of LAIR1 may cause macrophages to be preactivated — i.e., exhibit some characteristics of activation prior to immune stimulus — which could be indicative of altered regulation of immune response at the epigenetic level. These observations require further study to elucidate the underlying mechanisms.
In summary, our studies reveal LAIR1 as a critical modulator of pathogenic inflammation that is important for immune response to S. aureus infection and protection from CTCL immunopathology. We additionally demonstrate that LAIR1 loss increases the expression of cytokine signaling pathways, which has broader implications for other inflammatory diseases and malignancies in which LAIR1 signaling has been implicated (21, 70–73). Indeed, recent studies in solid tumor models demonstrated reduction in tumor size and increased survival by targeting LAIR signaling using chimeric antigen receptor (CAR) T cells (74, 75). Our findings suggest that LAIR signaling represents a promising target for therapeutic development for CTCL. Future studies are needed to investigate these compelling ideas.
-
Methods
Sex as a biological variable. For human studies, both male and female patients are included in approximately equal numbers. For animal studies, both male and female mice were used in equal proportions for in vitro experiments, including cytokine analysis from BMDM lysate, RNA-Seq of BMDMs, and experiments involving neutrophil and monocyte isolation. Male and female mice exhibit relevant differences in the stratum corneum — the surface epithelium of the skin. Visualization of epithelial lesions in these models is best performed in male mice as the stratum corneum is several cell layers thicker than in female mice. We confirmed that S. aureus skin infection in female mice exhibits a similar disease course as in male mice, including differences in Lair1 KO vs WT. Tissue-based analysis of the clinical correlates of disease are less informative in female mice, due to the very thin stratum corneum. Therefore, to minimize the number of animals needed to obtain consistency in the S. aureus skin infection models, only male mice were used, which is in agreement with previous reports on this model (76).
Mice. WT C57BL/6J and Lair1–/– (B6.Cg-Lair1tm1.1Jco/J, JAX Strain #:032788) mice (28) were purchased from The Jackson Laboratory and housed in a pathogen-free environment in the WUSM animal facility. Breeders were fed high-fat chow, and all other mice were fed standard rodent chow (5058; Purina). Inhaled isoflurane was used for all anesthesia.
Bacteria. S. aureus MRSA strain USA300/LAC (gift from Juliane Bubeck Wardenburg lab, WUSM) was used for all infections. Colonies from tryptic soy agar plates were grown overnight at 37°C in tryptic soy broth before being subcultured at a 1:10 dilution and grown until 0.5–0.7 OD600. Culture was diluted with tryptic soy broth to OD600 0.48, and bacteria were pelleted, washed in PBS, and diluted to the desired inoculum using the estimated dilution factor 2 × 108 bacteria/mL. Cultures were serially diluted and then plated onto tryptic soy agar to confirm desired inoculum.
Skin infections. On day 0, 2- to 3-month-old male mice were anesthetized; then, flanks were shaved and treated with hair removal cream for 2–3 minutes. After 24 hours, flanks were injected s.c. with expected dose of 1 × 107 CFU S. aureus (actual dose ranged from 1 × 107 to 2 × 107 CFU). Abscess and dermonecrosis length and width were measured over 14 days using digital calipers; then, area of abscess and dermonecrosis was calculated using the formula A = ([pi/2] × l × w). Dedicated mice were sacrificed at 24 or 48 hours after infection for 8 mm skin punch biopsy. For histology, skin punch biopsies were fixed in 10% neutral-buffered formalin for 24 hours at 4°C for paraffin-embedding and H&E staining (Washington University Division of Comparative Medicine Research Animal Diagnostic Laboratory) or fixed for 2 hours in 10% neutral-buffered formalin at room temperature before embedding in OCT Compound (Fisher, 4585) prior to trichrome and H&E staining (WUSM Pulmonary and Critical Care Morphology Core). For CFU analysis or cytokine production analysis, skin punch biopsies were homogenized in PBS and either serially diluted and plated onto tryptic soy agar (31) or stored at –80°C until analysis, respectively. For RNA-Seq, skin punch biopsies were homogenized in TRI reagent (Sigma, 93289) and stored at –80°C until RNA isolation. Results from an entire 8 mm skin punch biopsy are reported for each analysis.
Preparation of primary cells from mouse bone marrow. Bone marrow from age- and sex-matched WT and Lair1–/– mice was used for all primary cells. BMDMs were produced by treating bone marrow cells with the equivalent of 10 ng/mL M-CSF from CMG14-12 cells (gift from Deborah Veis lab, Washington University in St. Louis) for 6 days. Primary monocytes and neutrophils were isolated from mouse bone marrow using negative isolation kits (Miltenyi Biotec, 130-100-629 and 130-097-658). Cells were cultured in RPMI 1640 media (monocytes, neutrophils) or α-MEM media (BMDMs) supplemented with 10% FBS, 100 U/mL penicillin (Corning), 100 mg/mL streptomycin (Corning), and 50 μM 2-mercaptoethanol at 37°C in 5% CO2.
Immunofluorescence staining and image analysis. BMDMs prepared as stated above were plated onto glass coverslips at a density of 3 × 105 cells/mL and incubated for 24 hours with 500 ng/mL PAM3CSK4 (Tocris Bioscience, 4633) or PBS. After 24 hours, cells were fixed in 4% paraformaldehyde for 5 minutes at room temperature and were then stored in PBS until staining. All of the following steps were performed with three 5-minute PBS washes in between: cells were permeabilized with 0.1% TritonX-100 for 10 minutes at room temperature, blocked at 4° C for 24 hours in 2% BSA, incubated with Serpin H1 polyclonal antibody (Invitrogen, PA5-27832) at 1:100 dilution overnight, and finally incubated with Donkey α-Rabbit IgG-Alexa 488 (Invitrogen, A32790) for 1 hour at room temperature in the dark at 1:2,000 dilution. Coverslips were mounted onto slides using Everbrite Mounting Medium with DAPI (Biotium, 23002) and imaged on a Zeiss LSM 880 II FAST confocal microscope. Images were analyzed in CellProfiler (77) in order to quantify expression intensity, with default module settings used except as specified. Nuclei were identified with the IdentifyPrimaryObjects module (using the default Minimum Cross-Entropy thresholding method), cell outlines were identified using the IdentifySecondaryObjects module (using the adaptive threshold strategy, robust background thresholding method, threshold correction factor 1.1, lower threshold bound of 0.07), and cytoplasm was isolated using the IdentifyTertiaryObjects module. Mean fluorescence intensity per cell was exported and analyzed in R.
Western blotting. BMDM prepared and treated as for immunofluorescence were lysed in RIPA buffer, heated in Laemmli loading buffer (without 2-mercaptoethanol) at 99°C for 5 minutes, and 40 μg protein per lane loaded on 4%–20% gradient Mini-PROTEAN TGX Precast Gels (BioRad 4561093), transferred to nitrocellulose, blocked with 5% nonfat dried milk in TBS-T, washed, and incubated with Invitrogen Collagen I Polyclonal Antibody (Fisher Scientific, PA126204) 1:500 dilution or Invitrogen GAPDH Polyclonal Antibody (Fisher Scientific, PA1988) 1:4000 dilution before being washed and incubated with anti-rabbit secondary antibody (Invitrogen Goat anti-Rabbit IgG (H+L) Secondary Antibody, HRP, Fisher Scientific, PI32460), prior to incubation with SuperSignal West Pico PLUS Chemiluminescent Substrate (ThermoFisher Scientific, 34577). They were then imaged and bands quantified on an iBright CL1500 Imaging System.
In vitro functional assays. Amplex Red Hydrogen Peroxide Assay Kit (Invitrogen, A22188) was used according to manufacturer’s instructions to measure ROS production. To measure phagocytosis and lysosome acidification, neutrophils and monocytes were treated with either pHrodo Green S. aureus BioParticles (Invitrogen, P35382) or mCherry-expressing S. aureus (gift from the Veis lab, Washington University) (29) at multiplicity of infection 50:1 with or without lysosome probe NIR (BioLegend, 421916) at a concentration of 1:500. After incubation at 37° C in 5% CO2 for 30 minutes, cells were immediately transferred to ice and pHrodo, mCherry, or lysosome probe fluorescence analyzed by flow cytometry.
Cytokine analysis. Multiplex cytokine array (32-plex discovery panel, Eve Technologies) was conducted for skin homogenate and BMDM supernatant.
Patient sample collection. Deidentified peripheral blood was obtained from patients seen at the Washington University School of Medicine Cutaneous Lymphoma Clinic under IRB-approved protocols with patients providing informed consent.
Peripheral blood mononuclear cell isolation and scRNA-Seq. PBMCs were subjected to enrichment and/or depletion using antibody cocktails as detailed below to enable purification of the desired cells. Monocytes and neutrophils were depleted by incubation of peripheral blood samples with RosetteSep Human Monocyte (CD36) Depletion Cocktail (StemCell Technologies, 15628) for 15 minutes at room temperature, layered onto a Histopaque-1077 gradient, and centrifuged at 400g for 30 minutes with no brake. The interphase was collected and washed with 10 mL of sort buffer (PBS, 1% FBS, 2 mM EDTA), followed by red blood cell lysis (155 mM NH4Cl, 10 mM KHCO3, 0.1 mM EDTA) for 10 minutes. For each sample, 10,000–20,000 viable cells were submitted for processing using the 10x Genomics Chromium Controller and the Chromium Single Cell 5′ Library & Gel Bead Kit v2 (PN-1000006), Chromium Single Cell A Chip Kit (PN-1000152), Chromium Single Cell V(D)J Enrichment Kit, Human, Tcell (96rxns) (PN-1000005), and Chromium Single Index Kit T (PN-1000213) following the manufacturer’s protocols. Normalized libraries were sequenced on a NovaSeq6000 S4 Flow Cell using the XP workflow and a 151 × 10 × 10 × 151 sequencing recipe according to manufacturer protocol. A median sequencing depth of 50,000 reads/cell was targeted for each sample.
scRNA-Seq data processing and analysis. Alignment and gene counting were performed using the Cell Ranger pipeline (10x Genomics, v3.0). Genes found in fewer than 15 cells in a given sample were removed. For each patient, gene counts and cells were pooled into a single Seurat (v3.1.4) object (78), and cells containing fewer than 200 or more than 3,000–3,750 expressed genes, more than 8%–10% mitochondrial reads, fewer than 300 or more than 10,000–20,000 UMIs, or classification as a doublet by the R package scDblFinder with parameters dims = 30, clust.method = “fast_greedy” were removed. Normalization and regression of technical variation due to mitochondrial read percentage and read depth was performed with the SCTransform function with variables.features.n = 4,000. Integration to account for experimental variability due to differences in ficoll or buffy coat preparation and batch effects was performed using the Seurat wrapper around the fastMNN function from the batchelor R package (v1.4.0) with n.features = 3,000. Gene expression was normalized and the top 1,500 variable using the “VST” method were calculated. Data were integrated using the harmony (v1.0.0) R package (79) using both patient and batch information to correct for batch effect with up to 10 iterations. The UMAP and neighbors were calculated with Seurat, using 20 dimensions of the harmony calculations. Cell annotation was performed using the singler (v1.4.1) R package (80) using the highest Spearman rho of purified immune populations in the HPCA (34). Cell type designations with less than 50 cells in the entire cohort were reduced to “other.” Automated annotations were checked manually using canonical marker genes. All single-cell visualizations were performed with Seurat and ggplot2 R package (v3.5.1) (81).
RNA isolation and RNA-Seq. RNA was isolated from homogenized mouse skin punch biopsies using a Direct-zol RNA MiniPrep Plus kit (Zymo research, R2072). BMDMs were plated at 2 × 105 cells/mL and treated with PBS or 500 ng/mL PAM3CSK4 (Tocris Bioscience, 4633) for 6 hours. RNA was isolated using an RNeasy Plus Mini kit (Qiagen, 74134). RNA was sequenced by the Genome Technology Access Center at the McDonnell Genome Institute at Washington University School of Medicine on an Illumina NovaSeq-6000 to generate 150 bp paired end reads. RNA-Seq reads were then aligned to the Ensembl release 101 primary assembly with STAR (v 2.7.9a1). Gene counts were derived from the number of uniquely aligned unambiguous reads by Subread:featureCount (v 2.0.32). Isoform expression of known Ensembl transcripts were quantified with Salmon (v 1.5.23).
RNA-Seq analysis. R (v 4.1.3) was used for all analyses. DEseq2 (v 1.4.1) (82) was used to normalize gene counts and evaluate significance of individual genes for our own RNA-Seq and publicly obtained datasets. Genes with counts above expression cutoffs (averaged normalized counts > 100 for at least 1 group), significance < 0.5, and absolute log2 fold change > 0.1 were analyzed by GSEA (v 4.3.2) (83, 84) desktop software using the MSigDB mouse or human reactome (v.2023.2) gene set for pathway analysis. Heatmaps were generated using pheatmap (v 1.0.12) R package.
Statistics. GraphPad Prism 9 (Version 9.4.1) was used for all statistical tests other than RNA-Seq. All t tests were 2-tailed and all ANOVA were 2-way. A P value less than 0.05 was considered significant.
Study approval. Deidentified peripheral blood draws were obtained from patients seen at the Washington University School of Medicine Cutaneous Lymphoma Clinic under IRB-approved protocols with patients providing informed consent. All animal experimental protocols were approved by IACUC of Washington University in St. Louis. All animal experiments were performed in accordance with the Guide for the Care and Use of Laboratory Animals (National Academies Press, 2011)s.
Data availability. Sequencing data were deposited in GEO under accession nos. GSE269177 and GSE269178. External datasets used include GEO datasets GSE59307, GSE113113, and GSE221148. Additional datasets are available on Zenodo (DOI: 10.5281/zenodo.org/records/17527215). Values for all data points in graphs other than sequencing data are reported in Supporting Data Values file.
-
Author contributions
Conception and design were contributed by HKD, DJV, JBW, and JEP. Development of methodology was contributed by HKD, KLT, ECM, and JEP. Acquisition of data was contributed by HKD, KLT, YK, ECM, LT, ACM, NMS, CCQ, MKS, and JEP. Analysis, interpretation of data, and generation of figures were contributed by HKD, KLT, and JEP. Funding acquisition, writing, review, and/or revision of the manuscript were contributed by HKD, ECM, NMS, and JEP. Study supervision was contributed by JEP. All authors reviewed and approved the manuscript.
-
Funding support
This work is in part the result of NIH funding and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- American Heart Association Predoctoral Fellowship grant #P23-01044 to HKD.
- Washington University BioSURF funding to ECM.
- Siteman Cancer Center Foundation to NMS and JEP.
- Barnes-Jewish Hospital Foundation Steinbeck Designated Fund to NMS and JEP.
- McDonnell Genome Institute and Illumina research support to NMS and JEP.
- Gateway for Cancer Research Awards to NMS and JEP.
- The Siteman Cancer Center is supported in part by an NCI Cancer Center Support Grant #P30 CA091842.
- ICTS is funded by the National Institutes of Health’s NCATS Clinical and Translational Science Award (CTSA) program grant #UL1 TR00234.
-
Acknowledgments
We thank the Alvin J. Siteman Cancer Center at Washington University School of Medicine and Barnes-Jewish Hospital in St. Louis, Missouri, USA, and the Institute of Clinical and Translational Sciences (ICTS) at Washington University in St. Louis, for the use of the Genome Technology Access Center, which provided library preparation, sequencing, and data analysis services. We acknowledge the Genome Technology Access Center at the McDonnell Genome Institute at Washington University School of Medicine for help with genomic analysis. The Center is partially supported by NCI Cancer Center Support Grant #P30 CA91842 to the Siteman Cancer Center from the National Center for Research Resources (NCRR), a component of the NIH, and NIH Roadmap for Medical Research. We also acknowledge the Research Infrastructure Services (RIS) group at Washington University in St. Louis for providing computational resources and services needed to generate the research results delivered within this paper (https://ris.wustl.edu). We thank Ezri Perrin for assistance with editing and genotyping mice.
Address correspondence to: Jacqueline E. Payton, Washington University Department of Pathology and Immunology, 660 S. Euclid Ave., Box 8118, St. Louis, Missouri 63110, USA. Phone: 314.362.5935; Email: jpayton@wustl.edu.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2025, Dorando et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(1):e183935.https://doi.org/10.1172/jci.insight.183935.
-
References
- Dummer R, et al. Cutaneous T cell lymphoma. Nat Rev Dis Primers. 2021;7(1):61.
- Glinos G, et al. Characteristics and outcomes for hospitalized patients with cutaneous T-cell lymphoma. JAMA Dermatol. 2023;159(2):192–197.
- Allen PB, et al. Risk of bacteremia in patients with cutaneous T-cell lymphoma (CTCL). Leuk Lymphoma. 2020;61(11):2652–2658.
- Guenova E, et al. TH2 cytokines from malignant cells suppress TH1 responses and enforce a global TH2 bias in leukemic cutaneous T-cell lymphoma. Clin Cancer Res. 2013;19(14):3755–3763.
- Vieyra-Garcia P, et al. Benign T cells drive clinical skin inflammation in cutaneous T cell lymphoma. JCI Insight. 2019;4(1):e124233.
- Gonzalez BR, et al. Tumor microenvironment in mycosis fungoides and Sézary syndrome. Curr Opin Oncol. 2016;28(1):88–96.
- Querfeld C, et al. Primary T cells from cutaneous T-cell lymphoma skin explants display an exhausted immune checkpoint profile. Cancer Immunol Res. 2018;6(8):900–909.
- Andrews JM, et al. Novel cell adhesion/migration pathways are predictive markers of HDAC inhibitor resistance in cutaneous T cell lymphoma. EBioMedicine. 2019;46:170–183.
- Meyaard L, et al. LAIR-1, a novel inhibitory receptor expressed on human mononuclear leukocytes. Immunity. 1997;7(2):283–290.
- Lebbink RJ, et al. The soluble leukocyte-associated Ig-like receptor (LAIR)-2 antagonizes the collagen/LAIR-1 inhibitory immune interaction. J Immunol. 2008;180(3):1662–1669.
- Jacquet M, et al. C1q and mannose-binding lectin interact with CR1 in the same region on CCP24-25 modules. Front Immunol. 2018;9:453.
- Meyaard L. The inhibitory collagen receptor LAIR-1 (CD305). J Leukoc Biol. 2008;83(4):799–803.
- Dorando HK, et al. LPS and type I and II interferons have opposing effects on epigenetic regulation of LAIR1 expression in mouse and human macrophages. J Leukoc Biol. 2023;115(3):547–564.
- Carvalheiro T, et al. Leukocyte associated immunoglobulin like receptor 1 regulation and function on monocytes and dendritic cells during inflammation. Front Immunol. 2020;11:1793.
- Keerthivasan S, et al. Homeostatic functions of monocytes and interstitial lung macrophages are regulated via collagen domain-binding receptor LAIR1. Immunity. 2021;54(7):1511–1526.
- Verbrugge A, et al. Differential expression of leukocyte-associated Ig-like receptor-1 during neutrophil differentiation and activation. J Leukoc Biol. 2006;79(4):828–836.
- Maasho K, et al. The inhibitory leukocyte-associated Ig-like receptor-1 (LAIR-1) is expressed at high levels by human naive T cells and inhibits TCR mediated activation. Mol Immunol. 2005;42(12):1521–1530.
- Van der Vuurst de Vries AR, et al. Leukocyte-associated immunoglobulin-like receptor-1 (LAIR-1) is differentially expressed during human B cell differentiation and inhibits B cell receptor-mediated signaling. Eur J Immunol. 1999;29(10):3160–3167.
- Jin J, et al. LAIR-1 activation inhibits inflammatory macrophage phenotype in vitro. Cell Immunol. 2018;331:78–84.
- Poggi A, et al. p40/LAIR-1 regulates the differentiation of peripheral blood precursors to dendritic cells induced by granulocyte-monocyte colony-stimulating factor. Eur J Immunol. 1998;28(7):2086–2091.
- Hammad R, et al. Immunoregulatory complement receptor-1 and leukocyte-associated Ig-like receptor-1 expression on leukocytes in Psoriasis vulgaris. Innate Immun. 2020;26(8):683–692.
- Van Laethem F, et al. LAIR1, an ITIM-containing receptor involved in immune disorders and in hematological neoplasms. Int J Mol Sci. 2022;23(24):16136.
- Besteman SB, et al. Signal inhibitory receptor on leukocytes (SIRL)-1 and leukocyte- associated immunoglobulin-like receptor (LAIR)-1 regulate neutrophil function in infants. Clin Immunol. 2020;211:108324.
- Geerdink RJ, et al. LAIR-1 limits neutrophil extracellular trap formation in viral bronchiolitis. J Allergy Clin Immunol. 2018;141(2):811–814.
- Van der Wijst MGP, et al. Type I interferon autoantibodies are associated with systemic immune alterations in patients with COVID-19. Sci Transl Med. 2021;13(612):eabh2624.
- Achieng AO, et al. Integrated OMICS platforms identify LAIR1 genetic variants as novel predictors of cross-sectional and longitudinal susceptibility to severe malaria and all-cause mortality in Kenyan children. EBioMedicine. 2019;45:290–302.
- Achieng AO, et al. Molecular basis of reduced LAIR1 expression in childhood severe malarial anaemia: implications for leukocyte inhibitory signalling. EBioMedicine. 2019;45:278–289.
- Tang X, et al. Leukocyte-associated Ig-like receptor-1-deficient mice have an altered immune cell phenotype. J Immunol. 2012;188(2):548–558.
- Roper PM, et al. Contemporary clinical isolates of Staphylococcus aureus from pediatric osteomyelitis patients display unique characteristics in a mouse model of hematogenous osteomyelitis. Infect Immun. 2021;89(10):e0018021.
- Inoshima N, et al. Genetic requirement for ADAM10 in severe Staphylococcus aureus skin infection. J Invest Dermatol. 2012;132(5):1513–1516.
- Becker REN, et al. Tissue-specific patterning of host innate immune responses by Staphylococcus aureus α-toxin. J Innate Immun. 2014;6(5):619–631.
- Kennedy AD, et al. Targeting of alpha-hemolysin by active or passive immunization decreases severity of USA300 skin infection in a mouse model. J Infect Dis. 2010;202(7):1050–1058.
- Dorando H, et al. Clonal evolution of cutaneous T cell lymphoma revealed at single cell resolution. Blood. 2022;140(suppl 1):754–755.
- Mabbott NA, et al. An expression atlas of human primary cells: inference of gene function from coexpression networks. BMC Genomics. 2013;14:632.
- Borcherding N, et al. Single-cell profiling of cutaneous T-cell lymphoma reveals underlying heterogeneity associated with disease progression. Clin Cancer Res. 2019;25(10):2996–3005.
- Borcherding N, et al. Single-cell analysis of Sézary syndrome reveals novel markers and shifting gene profiles associated with treatment. Blood Adv. 2023;7(3):321–335.
- Sampedro GR, et al. Targeting Staphylococcus aureus α-toxin as a novel approach to reduce severity of recurrent skin and soft-tissue infections. J Infect Dis. 2014;210(7):1012–1018.
- Abtin A, et al. Perivascular macrophages mediate neutrophil recruitment during bacterial skin infection. Nat Immunol. 2014;15(1):45–53.
- Liu M, et al. CXCL10/IP-10 in infectious diseases pathogenesis and potential therapeutic implications. Cytokine Growth Factor Rev. 2011;22(3):121–130.
- Maurer M, von Stebut E. Macrophage inflammatory protein-1. Int J Biochem Cell Biol. 2004;36(10):1882–1886.
- Satoh T, Akira S. Toll-like receptor signaling and its inducible proteins. Microbiol Spectr. 2016;4(6)
- Bubeck Wardenburg J, et al. Host defenses against Staphylococcus aureus infection require recognition of bacterial lipoproteins. Proc Natl Acad Sci U S A. 2006;103(37):13831–13836.
- Hashimoto M, et al. Not lipoteichoic acid but lipoproteins appear to be the dominant immunobiologically active compounds in Staphylococcus aureus. J Immunol. 2006;177(5):3162–3169.
- Dovi JV, et al. Accelerated wound closure in neutrophil-depleted mice. J Leukoc Biol. 2003;73(4):448–455.
- Wang J. Neutrophils in tissue injury and repair. Cell Tissue Res. 2018;371(3):531–539.
- Khouili SC, et al. SHP-1 regulates antigen cross-presentation and is exploited by leishmania to evade immunity. Cell Rep. 2020;33(9):108468.
- Gómez CP, et al. The protein tyrosine phosphatase SHP-1 regulates phagolysosome biogenesis. J Immunol. 2012;189(5):2203–2210.
- Humme D, et al. Aurora kinase A is upregulated in cutaneous T-cell lymphoma and represents a potential therapeutic target. J Invest Dermatol. 2015;135(9):2292–2300.
- Licht P, Mailänder V. Multiomic data integration reveals microbial drivers of aetiopathogenesis in mycosis fungoides [preprint]. https://doi.org/10.1101/2024.01.15.575621 Posted on bioRxiv January 15, 2024.
- Steevels TAM, Meyaard L. Immune inhibitory receptors: essential regulators of phagocyte function. Eur J Immunol. 2011;41(3):575–587.
- Axelrod PI, et al. Infections complicating mycosis fungoides and Sézary syndrome. JAMA. 1992;267(10):1354–1358.
- Posner LE, et al. Septicemic complications of the cutaneous T-cell lymphomas. Am J Med. 1981;71(2):210–216.
- Germain RN. Maintaining system homeostasis: the third law of Newtonian immunology. Nat Immunol. 2012;13(10):902–906.
- Goldszmid RS, Trinchieri G. The price of immunity. Nat Immunol. 2012;13(10):932–938.
- Murray PJ, Smale ST. Restraint of inflammatory signaling by interdependent strata of negative regulatory pathways. Nat Immunol. 2012;13(10):916–924.
- Lee B, et al. Staphylococcus aureus toxin suppresses antigen-specific T cell responses. J Clin Invest. 2020;130(3):1122–1127.
- Liu Q, et al. Immune and inflammatory reponses to Staphylococcus aureus skin infections. Curr Dermatol Rep. 2018;7(4):338–349.
- Rumpret M, et al. Functional categories of immune inhibitory receptors. Nat Rev Immunol. 2020;20(12):771–780.
- Ramos MIP, et al. Cancer immunotherapy by NC410, a LAIR-2 Fc protein blocking human LAIR-collagen interaction. Elife. 2021;10:e62927.
- Kobayashi SD, et al. Pathogenesis of Staphylococcus aureus abscesses. Am J Pathol. 2015;185(6):1518–1527.
- Kumawat K, et al. LAIR-1 limits neutrophilic airway inflammation. Front Immunol. 2019;10:842.
- Mittal M, et al. Reactive oxygen species in inflammation and tissue injury. Antioxid Redox Signal. 2014;20(7):1126–1167.
- Auten RL, Davis JM. Oxygen toxicity and reactive oxygen species: the devil is in the details. Pediatr Res. 2009;66(2):121–127.
- De Jong NWM, et al. Immune evasion by Staphylococcus aureus. Microbiol Spectr. 2019;7(2):10.1128/microbiolspec.gpp3-0061-2019.
- Sarris AH, et al. Interferon-inducible protein 10 as a possible factor in the pathogenesis of cutaneous T-cell lymphomas. Clin Cancer Res. 1997;3(2):169–177.View this article via: PubMed Google Scholar
- Tensen CP, et al. Epidermal interferon-gamma inducible protein-10 (IP-10) and monokine induced by gamma-interferon (Mig) but not IL-8 mRNA expression is associated with epidermotropism in cutaneous T cell lymphomas. J Invest Dermatol. 1998;111(2):222–226.
- Lu D, et al. The T-cell chemokine receptor CXCR3 is expressed highly in low-grade mycosis fungoides. Am J Clin Pathol. 2001;115(3):413–421.
- Franck N, et al. Mycosis fungoides-type cutaneous T-cell lymphoma and neutrophilic dermatosis. Arch Dermatol. 2005;141(3):353–356.
- Cirée A, et al. Expression and activity of IL-17 in cutaneous T-cell lymphomas (mycosis fungoides and Sezary syndrome). Int J Cancer. 2004;112(1):113–120.
- Son M, et al. Evidence for C1q-mediated crosslinking of CD33/LAIR-1 inhibitory immunoreceptors and biological control of CD33/LAIR-1 expression. Sci Rep. 2017;7(1):270.
- Nordkamp MJMO, et al. Enhanced secretion of leukocyte-associated immunoglobulin-like receptor 2 (LAIR-2) and soluble LAIR-1 in rheumatoid arthritis: LAIR-2 is a more efficient antagonist of the LAIR-1-collagen inhibitory interaction than is soluble LAIR-1. Arthritis Rheum. 2011;63(12):3749–3757.
- Lovewell RR, et al. LAIR-1 agonism as a therapy for acute myeloid leukemia. J Clin Invest. 2023;133(22):e169519.
- Kang X, et al. The ITIM-containing receptor LAIR1 is essential for acute myeloid leukaemia development. Nat Cell Biol. 2015;17(5):665–677.
- Perrin EP, et al. Targeting LAIR1-mediated immunosuppression adds a new weapon to our immunotherapy arsenal. J Clin Invest. 2025;135(16):e194924.
- Tao H, et al. Blocking immune checkpoint LAIR1 with antibody blockade or 3-in-1 CAR T cells enhances antitumor response. J Clin Invest. 2025;135(16):e184042.
- Bubeck Wardenburg J, et al. Panton-Valentine leukocidin is not a virulence determinant in murine models of community-associated methicillin-resistant Staphylococcus aureus disease. J Infect Dis. 2008;198(8):1166–1170.
- Stirling DR, et al. CellProfiler 4: improvements in speed, utility and usability. BMC Bioinformatics. 2021;22(1):433.
- Stuart T, et al. Comprehensive integration of single-cell data. Cell. 2019;177(7):1888–1902.
- Korsunsky I, et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods. 2019;16(12):1289–1296.
- Aran D, et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol. 2019;20(2):163–172.
- Wickham H, ed. ggplot2. Cham: Springer International Publishing; 2016.
- Love MI, et al. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 2014;15(12):550.
- Subramanian A, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102(43):15545–15550.
- Mootha VK, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34(3):267–273.
-
Version history
- Version 1 (November 13, 2025): In-Press Preview
- Version 2 (January 9, 2026): Electronic publication