Submit a comment
Abstract
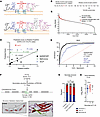
How SARS-CoV-2 causes a wide range of clinical manifestations and disease severity remains poorly understood. SARS-CoV-2 encodes 2 proteases (3CLPro and PLPro), vital for viral production, but also promiscuous with respect to host protein targets. Pharmacological inhibition of 3CLPro markedly reduced hospitalization and death in Phase 2/3 clinical studies. Here, we develop a bioinformatic algorithm, leveraging experimental data from SARS-CoV, to predict host cleavage targets of 3CLPro. We capture targets of 3CLPro described previously for SARS-CoV-2, as well as thousands of putative targets. We validate numerous targets cleaved during infection, including the giant sarcomeric protein obscurin and the innate immune protein OAS1. A long form of OAS1, p46, has been associated in numerous GWAS studies with lesser COVID disease severity. We show that 3CLPro cleaves p46 OAS1 immediately upstream of a known prenylation domain, relocalizing OAS1 from subcellular membranes to the cytosol, rendering it akin to the nonprotective, cytosolic p42 isoform. Similar OAS1 relocalization occurs upon infection by SARS-CoV-2. Our data provide a high-throughput resource to identify putative host cleavage targets of 3CLPro and reveal a mechanism by which SARS-CoV-2 antagonizes host innate immunity in individuals with the protective p46 isoform of OAS1.
Authors
Nora Yucel, Silvia Marchiano, Evan Tchelepi, Germana Paterlini, Ivan A. Kuznetsov, Kristina Li, Quentin McAfee, Nehaar Nimmagadda, Andy Ren, Sam Shi, Alyssa Grogan, Aikaterini Kontrogianni-Konstantopoulos, Charles Murry, Zoltan Arany
Guidelines
The Editorial Board will only consider comments that are deemed relevant and of interest to readers. The Journal will not post data that have not been subjected to peer review; or a comment that is essentially a reiteration of another comment.
- Comments appear on the Journal’s website and are linked from the original article’s web page.
- Authors are notified by email if their comments are posted.
- The Journal reserves the right to edit comments for length and clarity.
- No appeals will be considered.
- Comments are not indexed in PubMed.
Specific requirements
- Maximum length, 400 words
- Entered as plain text or HTML
- Author’s name and email address, to be posted with the comment
- Declaration of all potential conflicts of interest (even if these are not ultimately posted); see the Journal’s conflict-of-interest policy
- Comments may not include figures