Research ArticleImmunologyMetabolism
Open Access |  10.1172/jci.insight.193837
10.1172/jci.insight.193837
CD9 regulates macrophage-mediated remodeling of adipose tissue in obesity
Julia Chini,1,2,3 Nicole DeMarco,2 Dana V. Mitchell,4 Sam J. McCright,1,2,3 Kaitlyn M. Shen,5 Divyansi Pandey,1,2,3 Rachel L. Clement,1,2,3 Jessica Miller,2 Rajan Jain,5 Deanne M. Taylor,1,4 Mitchell A. Lazar,6,7 and David A. Hill1,2,3,7
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by Chini, J. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by DeMarco, N. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by Mitchell, D. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by McCright, S. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by Shen, K. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by Pandey, D. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by Clement, R. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by Miller, J. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by Jain, R. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by
Taylor, D.
in:
PubMed
|
Google Scholar
|

1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by Lazar, M. in: PubMed | Google Scholar
1Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
2Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
3Institute for Immunology and Immune Health, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Department of Biomedical and Health Informatics, Children’s Hospital of Philadelphia, Philadelphia, Pennsylvania, USA.
5Departments of Medicine and Cell and Developmental Biology, Penn Cardiovascular Institute, Penn Epigenetics Institute,
6Department of Medicine, Division of Endocrinology, and
7Institute for Diabetes, Obesity and Metabolism, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
Find articles by
Hill, D.
in:
PubMed
|
Google Scholar
|

Published February 10, 2026 - More info
JCI Insight. 2026;11(6):e193837. https://doi.org/10.1172/jci.insight.193837.
© 2026 Chini et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: March 25, 2025; Accepted: January 28, 2026
-
Abstract
Dysfunctional white adipose tissue contributes to the development of obesity-related morbidities, including insulin resistance, dyslipidemia, and other metabolic disorders. Adipose tissue macrophages (ATMs) accumulate in obesity and play both beneficial and harmful roles in the maintenance of adipose tissue homeostasis and function. Despite their importance, the molecules and mechanisms that regulate these diverse functions are not well understood. Lipid-associated macrophages (LAMs), the dominant subset of obesity-associated ATMs, accumulate in crown-like structures and are characterized by a metabolically activated and proinflammatory phenotype. We previously identified CD9 as a surface marker of LAMs. However, the contribution of CD9 to the activation and function of LAMs during obesity is unknown. Using a myeloid-specific CD9-KO model, we show that CD9 supports ATM-adipocyte adhesion and crown-like structure formation. Furthermore, CD9 promotes the expression of profibrotic and extracellular matrix remodeling genes. Loss of myeloid CD9 reduces adipose tissue fibrosis, increases visceral adipose tissue accumulation, and improves global metabolic outcomes during diet-induced obesity. These results identify CD9 as a causal regulator of pathogenic LAM functions, highlighting CD9 as a potential therapeutic target for treating obesity-associated metabolic disease.
-
Introduction
White adipose tissue undergoes dramatic remodeling in response to excess caloric intake, resulting in tissue expansion, inflammation, and fibrosis. This pathological remodeling promotes insulin resistance and ectopic lipid deposition in peripheral organs, predisposing individuals to the development of metabolic sequelae such as metabolic dysfunction–associated fatty liver disease, atherosclerosis, and type 2 diabetes (1). Although newer therapeutics, including glucagon-like peptide-1 (GLP-1) receptor agonists, can induce meaningful weight loss, emerging evidence indicates that the inflammatory and fibrotic features of adipose tissue remodeling often persist (2–6). Defining the mechanisms that drive adipose tissue remodeling is therefore essential for developing more effective strategies to prevent and treat metabolic disease.
In response to excess caloric intake, adipose tissue initially undergoes adaptive expansion through adipocyte hypertrophy and hyperplasia, thereby increasing its storage capacity for lipids. However, with prolonged overnutrition, adipocyte hypertrophy can outpace angiogenesis, leading to tissue hypoxia and adipocyte death (7). Adipocyte dysfunction and apoptosis promote the infiltration and proliferation of immune cells, particularly adipose tissue macrophages (ATMs). ATMs accumulate in crown-like structures surrounding dead and dying adipocytes, where they support tissue homeostasis through the efferocytosis of apoptotic adipocytes and the uptake of free fatty acids released during adipocyte lipolysis (8–12). At the same time, ATMs can contribute to adipose tissue dysfunction by producing pro-inflammatory and profibrotic mediators (10, 13–19). Despite their central role in adipose tissue remodeling, the molecular pathways that direct beneficial versus pathogenic ATM functions remain poorly understood.
During obesity, a subset of ATMs acquires a metabolically activated phenotype characterized by increased lipid accumulation and processing, fatty acid–dependent cellular metabolism, and proinflammatory cytokine production (9, 10, 13, 14, 20). This population, termed lipid-associated macrophages (LAMs), arises largely from recruited monocytes and becomes the dominant ATM subset in adipose tissue during obesity (19). LAMs localize to crown-like structures surrounding dead and dying adipocytes, where they mediate efferocytosis and lipid handling in part through the lipid receptor TREM2 (11, 21, 22).
We previously identified CD9 as a surface marker of LAMs in both mice and humans (11, 23). CD9 is a tetraspanin molecule that recruits and stabilizes proteins such as integrins, cytokine receptors, and metalloproteinases to tetraspanin-enriched microdomains (24–28). Through these interactions, CD9 contributes to cellular adhesion, migration, signal transduction, and extracellular vesicle formation (24, 26, 29–32). Since our initial report, CD9 has been widely used to identify LAMs across multiple other tissues, including the liver and aorta (33–38). Scar-associated macrophages (SAMs), which accumulate during liver and lung fibrosis, also share features with LAMs, including the expression of CD9 (36, 37, 39, 40). Despite the strong association between macrophage CD9 expression and metabolic dysfunction, the influence of CD9 on LAM functions during obesity remains unknown.
To investigate the functional role of CD9 in LAMs during obesity, we developed and characterized a myeloid-specific CD9-KO mouse model. Using this model, we identified a role for CD9 in mediating macrophage adhesion to adipocytes, which is critical for crown-like structure formation during obesity. Moreover, we found that CD9 promotes the expression of fibrosis-associated genes in macrophages. Consequently, loss of CD9 in myeloid cells led to attenuated adipose tissue remodeling and fibrosis, resulting in improved systemic metabolic outcomes during diet-induced obesity. Together, these findings establish CD9 as a pathogenic regulator of LAM function, suggesting that CD9 may be a potential therapeutic target for obesity-associated metabolic disease.
-
Results
CD9+ ATMs display a transcriptional profile enriched for extracellular matrix remodeling and adhesion pathways. We previously compared the transcriptional profiles of CD9+ ATMs and Ly6c+ monocytes and found that CD9+ ATMs were enriched for proinflammatory and lysosomal-dependent lipid metabolism gene programs (23). During obesity, CD9+ ATMs are thought to arise from monocytes via a CD9– ATM transitional state (11, 41). To further investigate the role of CD9 in ATMs, we compared the transcriptomes of sort-purified CD9+ and CD9– ATMs isolated from the visceral epididymal white adipose tissue (eWAT) of WT mice fed a high-fat diet (HFD) for 12 weeks (23). We found that 97 genes were upregulated and 84 genes were downregulated in CD9+ ATMs compared with CD9– ATMs (Figure 1A). CD9– ATMs demonstrated increased expression of genes encoding pattern recognition receptors (Cd209b, Cd209f, and Cd209g) and genes associated with antiinflammatory and perivascular macrophages (Lyve1, Retnla, Cd163, and Ednrb) (42). In contrast, CD9+ ATMs were enriched for genes previously identified in LAMs and SAMs, including Spp1, Gpnmb, Mmp12, Fabp5, Lpl, Cd36, and Lgals3 (9, 11, 13, 14, 36, 43) (Figure 1, A and B).
 Figure 1
Figure 1CD9+ adipose tissue macrophages have increased expression of extracellular matrix remodeling and adhesion genes. Bulk RNA-Seq analysis of CD9+ and CD9– adipose tissue macrophages (ATMs) sorted from epididymal white adipose tissue (eWAT) of WT male mice fed an HFD for 12 weeks (n = 3–5). (A) Volcano plot of gene expression in CD9+ versus CD9– ATMs showing the log2 fold-change (FC; x axis) and adjusted P value (–log10 FDR; y axis) of genes. Significantly differentially expressed genes (log2 FC > 0.58 and FDR < 0.05) are shown in purple. (B) Differential expression of genes previously associated with CD9+ macrophages (log2 FC of CD9+ ATMs/CD9– ATMs). Genes are listed in order from highest to lowest log2 FC. (C) Gene set enrichment analysis showing all Reactome pathways significantly upregulated in CD9+ ATMs compared with CD9– ATMs. (D) Heatmap of differentially expressed genes (FDR > 0.1) from Extracellular Matrix Organization Pathway (Reactome: R-MMU-1474244). P2RY1, p2Y purinoreceptor 1.
Gene set enrichment analysis revealed that CD9+ ATMs were enriched for pathways involved in extracellular matrix (ECM) remodeling, including programs related to both ECM formation and degradation (Figure 1, C and D). Furthermore, CD9+ ATMs were enriched for genes associated with cell adhesion, including Ncam1 (CD56) and integrin-related interactions (Figure 1C). These data indicate that CD9+ ATMs have a transcriptional profile characterized by enhanced expression of ECM-remodeling and adhesion-associated genes.
CD9 stabilizes ATM-adipocyte interactions in vivo. During obesity, CD9+ LAMs accumulate in adipose tissue in crown-like structures surrounding dead or dying adipocytes (44–46). We therefore investigated whether CD9 regulates ATM accumulation and localization in adipose tissue during obesity. To assess the functional role of CD9 in macrophages, we first generated a Cd9 conditional KO mouse line (Cd9fl/fl) by inserting loxP sites in the endogenous Cd9 locus (Supplemental Figure 1A; supplemental material available online with this article; https://doi.org/10.1172/jci.insight.193837DS1). Cd9fl/fl mice were then crossed to mice expressing Cre recombinase under the endogenous Lyz2 promoter (LysMCre) to generate a myeloid-specific CD9-KO (CD9-MKO) mouse model (47). Successful insertion of loxP sites was confirmed by DNA PCR genotyping (Supplemental Figure 1B). A reduction in gene expression of Cd9 and cell-surface expression of CD9 was confirmed by qPCR and flow cytometry of unpolarized BM-derived macrophages (BMDMs) (Supplemental Figure 1, C and D). Because CD9-MKO BMDMs exhibited only partial CD9 deletion, we also crossed Cd9fl/fl mice with Vav-iCre mice to generate a hematopoietic-wide CD9 KO (48). BMDMs from Cd9fl/fl Vav-iCre+/– mice had complete ablation of CD9 expression, thereby confirming the efficiency and fidelity of the Cd9fl/fl conditional allele (Supplemental Figure 1, E and F).
Next, we placed CD9-MKO and control (Cd9fl/fl) mice on an HFD for 12 weeks and examined the visceral eWAT depot. Since female mice are known to be resistant to adipose tissue inflammation during diet-induced obesity (49–51), initial studies were performed in male mice. Immunofluorescence staining of eWAT sections revealed decreased expression of the pan-macrophage markers F4/80 and CD68 in CD9-MKO HFD mice compared with control HFD mice. In addition, the number of F4/80+ and CD68+ cells were also reduced in CD9-MKO HFD mice compared with control HFD mice (Figure 2A and Supplemental Figure 2A). Further, CD9-MKO HFD eWAT also displayed reduced expression of the LAM markers TREM2 and OPN compared with control HFD eWAT (Supplemental Figure 2, B and C). Overall, these data indicate that myeloid-specific deletion of CD9 decreases LAM accumulation in eWAT during diet-induced obesity.
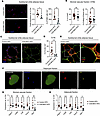 Figure 2
Figure 2CD9 promotes adipose tissue macrophage accumulation, crown-like structure formation, and adipocyte interactions. Analysis of epididymal white adipose tissue (eWAT) from control (Cd9fl/fl) and CD9-MKO (Cd9fl/fl LysMCre+/–) male mice fed an HFD for 12 weeks. (A) Representative images and quantification of immunofluorescence staining of F4/80 and DAPI in eWAT tissue sections. Data shown as relative MFI of F4/80 (n = 6–8) and number of F4/80+ DAPI+ cells per mm2 (n = 8). Scale bar: 200 μm. (B) Quantification by flow cytometry of adipose tissue macrophages (ATMs) from control and CD9-MKO HFD mice. Data are shown as frequency of total CD45+ immune cells (n = 15–16) and cells per gram of tissue (n = 13–16). (C) Representative high-resolution images of eWAT tissue sections from control and CD9-MKO HFD mice stained with DAPI (blue), PLIN1 (green), F4/80 (red), and CD9 (yellow) (n = 8). Scale bar: 50 μm. (D) F4/80+ crown-like structures (CLSs) per mm2 quantified from images from samples shown in A (n = 6–8). (E) 3D image reconstruction of PLIN1 (green), F4/80 (red), and CD9 (yellow) from control HFD mice (n = 8). Scale bar: 50 μm (left) or 20 μm (right). (F) Representative images of adipocyte fraction isolated from control HFD mice eWAT and stained for Hoechst (blue), BODIPY (green), F4/80 (red), and CD9 (yellow) (n = 3). Scale bar: 50 μm. (G and H) Relative expression of Adgre1 (F4/80), Cd68, Cd9, Trem2, and Spp1 (OPN) in stromal vascular fraction (n = 6–9; G) and adipocyte fraction (n = 6–9; H) isolated from eWAT. Data are from 2 (A, B, D, and E) or 3 (C, and F–H) independent pooled experiments. Data presented as mean ± SEM (A–C, G, and H). qPCR shown as ΔCt relative to a normalization factor relative to control HFD samples. Unpaired 2-tailed Student’s t test (A–C, G, and H). ns, not significant; *P < 0.05, **P < 0.01, ***P < 0.001.
Paradoxically, flow cytometric analyses of the stromal vascular fraction revealed no difference in the frequency or number of total ATMs between CD9-MKO and control HFD mice, despite a significant reduction in CD9 expression in ATMs and other myeloid cells in CD9-MKO HFD mice (Figure 2B, Supplemental Figure 3, and Supplemental Figure 4, A–E). Likewise, the total number of CD45+ immune cells and the distribution of other immune cell subsets in the stromal vascular fraction were unchanged (Supplemental Figure 4, F and G).
To explain the discrepancy between our immunofluorescence and flow cytometry findings, we hypothesized that a subset of ATMs might be lost during stromal vascular fraction preparation. Prior work has shown that some ATMs can remain tightly adherent to adipocytes and therefore partition into the adipocyte fraction, which is typically discarded during flow cytometry processing (52). To assess whether CD9+ ATMs associate with adipocytes, we performed high-resolution imaging of eWAT from control HFD and CD9-MKO HFD mice. Consistent with our prior studies (11, 23), CD9+ ATMs localized to crown-like structures surrounding adipocytes, including those lacking Perilipin-1 (PLIN1) expression, a feature of dying adipocytes (Figure 2C and Supplemental Figure 5) (53). In addition, CD9-MKO HFD mice exhibited fewer F4/80+ crown-like structures compared with control HFD mice (Figure 2, C and D). Three-dimensional reconstruction further confirmed that CD9+ ATMs reside in close proximity to adipocytes and are thus appropriately positioned for direct adhesion (Figure 2E).
Since CD9+ ATMs were closely associated with adipocytes, we hypothesized that CD9+ ATMs may be found within the adipocyte fraction. Indeed, we observed that CD9+ F4/80+ ATMs could be found attached to adipocytes within the floating adipocyte fraction (Figure 2F). To test whether ATM-intrinsic CD9 influences this retention, we quantified macrophage markers in both the stromal vascular fraction and adipocyte fraction from eWAT of control HFD and CD9-MKO HFD mice. Consistent with flow cytometry data, there were minimal differences in the expression of pan-macrophage markers (Adgre1 [F4/80] and Cd68) or LAM markers (Cd9, Trem2, and Spp1 [OPN]) in the stromal vascular fraction (Figure 2G). However, all 5 markers were reduced in the eWAT adipocyte fraction from CD9-MKO HFD mice compared with control HFD mice (Figure 2H). These findings demonstrate that macrophage-intrinsic CD9 promotes ATM-adipocyte interactions and is essential for the efficient formation of crown-like structures during obesity.
CD9 regulates integrin expression and macrophage-adipocyte adhesion. To investigate the functional role of CD9 in macrophage-adipocyte adhesion, we utilized an in vitro model in which BMDMs were metabolically activated with palmitate. Palmitate-treated BMDMs recapitulate key transcriptional and functional features of obesity-associated ATMs (9, 20). However, it is not known whether CD9 expression is induced by this metabolic stimulus. Metabolic activation with palmitate strongly induced the expression of Cd9 transcript and surface CD9 protein expression in BMDMs (Figure 3, A and B, and Supplemental Figure 6, A–C). CD9 deletion did not affect BMDM survival, differentiation, or proliferation of palmitate-treated BMDMs (Supplemental Figure 6, D–F). Further, palmitate also induced Cd9 expression in sort-purified blood monocytes from WT mice fed a normal chow diet (NCD; Figure 3C). Together, these results suggest that CD9 upregulation is a general feature of monocyte and macrophage metabolic activation.
 Figure 3
Figure 3CD9 promotes macrophage integrin expression and adhesion. (A and B) BMDMs from control (Cd9fl/fl) and CD9-MKO (Cd9fl/fl LysMCre+/–) mice were treated with BSA or palmitate for 24 hours. (A) Expression of Cd9 was assessed by qPCR (n = 6). Shown as expression relative to BSA controls (Ctrl). (B) Surface expression of CD9 shown as relative MFI compared with BSA controls. BMDMs were gated as live CD45+ CD11b+ F4/80+ CD64+ cells (n = 4). (C) Blood monocytes isolated from WT mice fed a normal chow diet were treated with BSA or palmitate for 24 hours. Expression of Cd9 was assessed by qPCR (n = 7–8). Monocytes were gated using the gating strategy in Supplemental Figure 3. (D) Expression of integrins Itgax and Itgav by qPCR in BMDMs from control and CD9-MKO mice treated with BSA or palmitate (n = 6). (E) Flow cytometry of CD11c (Itgax) and CD51 (Itgav) shown as MFI relative to control BMDMs treated with palmitate (n = 5). (F) Adhesion assay of control or CD9-MKO BMDMs on differentiated 3T3-L1 adipocytes (n = 6). (G) Relative expression of Itgax and Itgav in epididymal white adipose tissue (eWAT) adipocyte fraction isolated from control and CD9-MKO mice fed an HFD (n = 6–9). (H and I) Representative images and quantification of immunofluorescence staining of CD11c (H) and CD51 (I) in eWAT. Data shown as relative MFI (n = 8–9). Scale bar: 200 μm. Data shown as combined data from 2–3 independent experiments (C, G–I) or representative data from 3 independent experiments (A, B, and D–F). qPCR is shown as ΔCt relative to Hprt normalized to BSA controls. Data presented as mean ± SEM (A–I); 2-way ANOVA with Fisher’s LSD test (A, B, and D) or unpaired 2-tailed Student’s t test (C, and E–I). ns, not significant; **P < 0.01, ***P < 0.001, ****P < 0.0001.
Given the established role of CD9 in regulating cell-cell adhesion, we hypothesized that CD9 may promote macrophage-adipocyte adhesion by recruiting and stabilizing integrins at the cell surface (26, 30, 31). Consistent with this idea, metabolic activation of BMDMs with palmitate increased expression of the integrins Itgax (CD11c) and Itgav (CD51) in a CD9-dependent manner (Figure 3D). Further, palmitate-treated BMDMs from CD9-MKO mice had reduced surface levels of CD11c and CD51 compared with palmitate-treated control BMDMs (Figure 3E). As it has previously been described that ATMs form tight adhesions to adipocytes via integrin interactions, we next assessed whether CD9 affects this interaction (52, 54). We found that CD9-MKO BMDMs had reduced adhesion to 3T3-L1 adipocytes compared with control BMDMs, identifying a key role for CD9 in macrophage-adipocyte interactions (Figure 3F).
Next, to determine whether CD9 deletion also alters integrin expression in vivo, we assessed Itgax and Itgav levels in eWAT of control and CD9-MKO HFD mice. Consistent with our in vitro data, deletion of CD9 reduced Itgax and Itgav expression in the eWAT adipocyte fraction (Figure 3G). On the other hand, Itgax but not Itgav was decreased in the eWAT stromal vascular fraction of CD9-MKO HFD mice compared with control HFD mice (Supplemental Figure 6G). Immunofluorescence staining similarly showed decreased CD11c and CD51 protein expression in CD9-MKO HFD eWAT compared with controls (Figure 3, H and I). Overall, these results suggest that CD9 regulates integrin expression in metabolically activated macrophages, both in vitro and in vivo, providing a potential mechanism by which CD9 influences macrophage-adipocyte interactions.
Myeloid-intrinsic CD9 mediates adipose tissue remodeling during obesity. Given that ATMs are known to have both beneficial and pathological effects on adipose tissue adaptation in obesity, we next assessed how myeloid-specific CD9 deletion influences adipose tissue remodeling during obesity. To investigate this, we examined eWAT from CD9-MKO and control mice fed either an NCD or an HFD for 12 weeks. CD9-MKO mice exhibited slightly increased eWAT depots following HFD, but not NCD, compared with controls (Figure 4A). Histological analysis further demonstrated larger adipocyte size in CD9-MKO HFD eWAT compared with controls (Figure 4, B and C, and Supplemental Figure 7, A and B).
 Figure 4
Figure 4Myeloid-intrinsic CD9 alters adipose tissue remodeling during obesity. Epididymal white adipose tissue (eWAT) was collected from male control (Cd9fl/fl) and CD9-MKO (Cd9fl/fl LysMCre+/–) mice fed either an NCD or HFD for 12 weeks. (A) eWAT mass (n = 10–15). (B) Representative H&E staining of eWAT in control or CD9-MKO HFD mice (n = 11–12). Scale bar: 400 μm. (C) Frequency distribution and mean adipocyte diameter calculated from H&E images in B (n = 11–12). (D and E) Bulk RNA-Seq was performed on eWAT depots from control and CD9-MKO mice (n = 4). (D) Volcano plot showing the log2 fold-change (FC; x axis) and adjusted P value (–log10 FDR; y axis) of genes between CD9-MKO HFD and control HFD groups. Significantly differentially expressed genes (log2 FC > 0.58 and FDR < 0.05) are shown in purple. (E) Pathway enrichment analysis showing the most significant (by normalized enrichment score) nonredundant pathways downregulated in eWAT. Pooled data from 3 independent experiments (A–C) or data from 1 experiment (D and E). Data presented as mean ± SEM (A and C); 1-way ANOVA with Fisher’s LSD test (A) or unpaired 2-tailed Student’s t test between means (C). ns, not significant; *P < 0.05, ****P < 0.0001. Avg, average; MMP, matrix metalloproteinases; NOXs, NADPH oxidases.
To further assess how loss of CD9 influences adipose tissue remodeling, we performed bulk RNA-Seq of eWAT from CD9-MKO HFD mice and control HFD mice. We identified 1,873 genes upregulated and 2,204 genes downregulated in the eWAT of CD9-MKO HFD mice compared with control HFD mice (Figure 4D). Gene set enrichment analysis revealed that the top downregulated pathways in CD9-MKO HFD eWAT were associated with inflammatory cell activation and function, ECM remodeling, and cell adhesion (Figure 4E). Consistent with our previous data, expression of Itgax and Itgav was reduced in whole eWAT from CD9-MKO HFD mice compared with control HFD mice (Supplemental Figure 7, C and D). In contrast, the top upregulated pathways included those related to cellular metabolism and mitochondrial function (Supplemental Figure 7E). Collectively, these findings demonstrate that myeloid-intrinsic CD9 broadly regulates inflammatory, adhesive, and profibrotic remodeling programs in adipose tissue during diet-induced obesity.
Myeloid-intrinsic CD9 promotes adipose tissue fibrosis during diet-induced obesity. Given that CD9+ ATMs were enriched for ECM remodeling genes and that these pathways were downregulated in CD9-MKO HFD eWAT compared with controls, we next investigated whether loss of myeloid-intrinsic CD9 affected eWAT fibrosis during obesity. Histological analysis demonstrated reduced collagen deposition in CD9-MKO HFD eWAT compared with controls (Figure 5A). Next, we investigated whether this reduction in fibrosis was associated with changes in the transcriptional profile of eWAT adipocyte fraction. Fibrosis-associated genes (Lgals3, Pdgfb, Mmp12, Gpnmb, and Adam8) were decreased in the adipocyte fraction of CD9-MKO HFD mice compared with control HFD mice (Figure 5B). Notably, with the exception of Lgals3, these genes have either very low or no expression in adipocytes, suggesting that the reduction in expression of these genes primarily reflects the loss of CD9+ ATMs adhering to adipocytes within the adipocyte fraction (55).
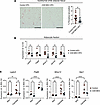 Figure 5
Figure 5Myeloid-intrinsic CD9 promotes adipose tissue fibrosis and macrophage profibrotic gene expression during obesity. Epididymal white adipose tissue (eWAT) was collected from male control (Cd9fl/fl) and CD9-MKO (Cd9fl/fl LysMCre+/–) mice fed an HFD for 12 weeks. (A) Representative images and quantification of Picrosirius red staining of eWAT from control HFD and CD9-MKO HFD mice (n = 13–15). Scale bar: 400 μm. (B) qPCR for fibrosis-associated genes in the adipocyte fraction isolated from control and CD9-MKO eWAT (n = 7–9). Data shown as ΔCt relative to a normalization factor relative to control HFD samples. (C) Expression of fibrosis-associated genes Lgals3, Pdgfb, Mmp12, and Spp1 in BMDMs from control and CD9-MKO mice treated with BSA or palmitate for 24 hours (n = 5–6). Data shown as ΔCt relative to Hprt normalized to BSA controls. Pooled data (A and B) or representative data (C) from 3 independent experiments. Data presented as mean ± SEM; unpaired 2-tailed Student’s t test (A and B) or 2-way ANOVA with Fisher’s LSD test (C). ns, not significant; *P < 0.05, **P < 0.01, ****P < 0.0001.
CD9 regulates the expression of fibrosis-associated genes in macrophages in vitro. LAMs are characterized by the expression of lipid metabolic, proinflammatory, and profibrotic programs (11, 23, 40). To determine whether CD9 contributes to these hallmark features, we assessed the effects of CD9 deficiency on metabolically activated macrophages. We found that CD9 did not alter the expression of lipid uptake and metabolism genes, including Plin2, Cd36, Lpl, and Lipa, in palmitate-treated BMDMs (Supplemental Figure 8A). Similarly, induction of the proinflammatory cytokines Tnfa and Il6 was unaffected by CD9 deletion (Supplemental Figure 8B). Finally, loss of CD9 did not influence intracellular lipid accumulation, as assessed by boron-dipyrromethene (BODIPY) staining (Supplemental Figure 8C).
Since loss of CD9 did not affect lipid metabolism or proinflammatory genes, we next examined whether CD9 influences expression of other LAM-associated genes. Palmitate treatment induced the expression of LAM genes Lgals3, Pdgfb, Mmp12, and Spp1, but not Trem2, Gpnmb, or Adam8, in BMDMs (Figure 5C and Supplemental Figure 8D). Furthermore, the induction of Lgals3, Pdgfb, Mmp12, and Spp1 was dependent on CD9 expression, whereas Trem2, Gpnmb, and Adam8 expression was unaffected by CD9 deficiency. Consistent with our RNA-Seq findings, Spp1, Lgals3, Mmp12, and Pdgfb are well-established mediators of ECM remodeling and are expressed by macrophages in fibrotic disease (11, 36, 40, 41, 56–58). Together, these results suggest that myeloid-intrinsic CD9 promotes the development of adipose tissue fibrosis during obesity by modulating the expression of fibrosis-associated genes.
To further understand how CD9 regulates fibrosis-associated gene expression, we examined the contribution of integrin signaling to the expression of CD9-dependent genes. Treatment of BMDMs with blocking antibodies against CD11c or CD51 resulted in reduced expression of Lgals3, Pdgfb, and Mmp12, but not Spp1, compared with treatment with isotype controls (Supplemental Figure 9). These findings suggest that integrin-mediated binding or signaling may represent one mechanism through which CD9 modulates the expression of fibrosis-associated genes in macrophages.
Myeloid-intrinsic CD9 worsens obesity-associated metabolic dysfunction. Given that adipose tissue remodeling contributes to global metabolic dysfunction, we next examined how loss of myeloid-intrinsic CD9 affects systemic metabolic outcomes during diet-induced obesity. Despite having slightly larger eWAT depots, CD9-MKO mice gained less overall weight on an HFD than control mice (Figure 6A). This reduced weight gain was accompanied by a decrease in subcutaneous adipose tissue and liver mass, with no differences in muscle or brown adipose tissue weight (Figure 6, B and C, and Supplemental Figure 10, A–C). Notably, CD9-MKO HFD mice had increased perirenal adipose tissue, another form of visceral adipose tissue, relative to control HFD mice (Supplemental Figure 10D).
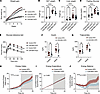 Figure 6
Figure 6Myeloid-intrinsic CD9 contributes to obesity-associated metabolic dysfunction. (A) Weekly body weights of male control (Cd9fl/fl) and CD9-MKO (Cd9fl/fl LysMCre+/–) mice fed either an NCD or HFD for 12 weeks (n = 10–29). (B and C) Tissue weights in control or CD9-MKO mice fed either an NCD or HFD for 12 weeks. (B) Subcutaneous (inguinal) adipose tissue weight (n = 6–15). (C) Liver (left lobe) weight (n = 9–18). (D) Glucose levels after a 16-hour fast in control or CD9-MKO mice fed an HFD for 12 weeks (n = 11–14). (E) Intraperitoneal glucose tolerance test (GTT) was performed in control or CD9-MKO mice after 12 weeks of NCD or HFD (n = 11–14). (F) Plasma insulin levels measured by ELISA after fasted (16 hours) or refed (4 hours) conditions (n = 6–8). (G) Plasma triglyceride levels after fasted (16 hours) or refed (4 hours) conditions (n = 6–7). (H–J) Cumulative energy intake (n = 6–8; H), energy expenditure (n = 7–9; I), and energy balance (n = 6–8; J) in control and CD9-MKO mice over a 24-hour period. Gray sections indicate dark photoperiods. Pooled data from 2–6 independent experiments (A–J). Data presented as mean ± SEM; 2-way ANOVA with Fisher’s LSD test (A, and E–J) performed separately for NCD and HFD groups (A and E) or 1-way ANOVA with Fisher’s LSD test (B–D). ns, not significant; *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Additionally, CD9-MKO HFD mice exhibited reduced fasting glucose levels and improved glucose tolerance compared with control HFD mice (Figure 6, D and E). CD9-MKO HFD mice also had a reduction in fed plasma insulin and triglyceride levels compared with control HFD mice (Figure 6, F and G). On the other hand, CD9-MKO HFD mice had no differences in plasma non-esterified fatty acids or glycerol compared with control HFD mice in both the fed and fasted conditions (Supplemental Figure 10, E and F). Notably, the differences in weight gain, tissue mass, fasting glucose, and glucose intolerance were not observed under NCD conditions (Figure 6, A–E).
To investigate the mechanism underlying reduced weight gain in CD9-MKO HFD mice, we performed indirect calorimetry studies. CD9-MKO mice exhibited decreased energy intake, with no corresponding differences in energy expenditure (Figure 6, H and I, and Supplemental Figure 10, G and H). This reduction in caloric intake resulted in a lower net energy balance, likely contributing to the diminished weight gain observed in CD9-MKO mice on HFD (Figure 6J and Supplemental Figure 10I). Notably, differences in energy intake persisted after adjusting for lean mass, indicating that the reduced levels of energy intake observed in CD9-MKO mice were independent of differences in lean mass (Supplemental Figure 10J).
Finally, we assessed whether the weight and glucose homeostasis phenotypes observed in male mice were also present in female mice. In contrast to males, female CD9-MKO HFD mice showed no differences in weight gain, adipose tissue mass, or liver mass compared with control HFD mice (Supplemental Figure 11, A–D). Further, female CD9-MKO mice had no differences in gonadal white adipose tissue adipocyte size, crown-like structure formation, or fibrosis (Supplemental Figure 11, E–H). Consistent with these findings, fasting glucose levels and glucose intolerance were unchanged in female CD9-MKO HFD mice (Supplemental Figure 11, I and J). These sex-specific differences may reflect the lower frequency of CD9+ ATMs in female gonadal white adipose tissue, as female mice did not exhibit the prominent ATM accumulation observed in male mice (Supplemental Figure 11, K–M), in agreement with previous reports (49, 51).
-
Discussion
ATMs play both beneficial and detrimental roles during obesity-associated adipose tissue remodeling. A deeper understanding of the molecules and mechanisms that drive the pathogenic activation of ATMs is critical for developing new therapeutic strategies for obesity and its metabolic complications. In this study, we demonstrate that CD9 is required for macrophage-adipocyte adhesion and the formation of crown-like structures. Additionally, loss of CD9 attenuated the profibrotic activation of ATMs and protected against pathogenic adipose tissue remodeling during diet-induced obesity. Consistent with CD9 serving a primarily pathogenic role in macrophages, myeloid-specific CD9 deletion improved systemic metabolic outcomes in obese mice. These findings establish CD9 as a critical regulator of pathogenic ATM functions, highlighting it as a potential target for mitigating obesity-associated metabolic dysfunction.
Since the initial identification of CD9 as a marker of LAMs in adipose tissue, CD9 upregulation has been observed in LAM-like macrophages across multiple murine and human tissues. This macrophage subset is characterized by the expression of lipid uptake and metabolism genes, intracellular lipid accumulation, proinflammatory cytokine production, and presence in crown-like structures (11, 23). Emerging evidence suggests that the upregulation of lipid metabolism may be an early feature of LAM activation that occurs prior to crown-like structure formation and the induction of CD9 expression (10, 41). Despite CD9’s widespread use as a defining LAM marker, it has remained unclear whether CD9 actively contributes to LAM lipid metabolic programs or is simply upregulated as a downstream consequence of their metabolic activation state.
CD9– ATMs are thought to represent a transitional state between monocytes and CD9+ ATMs (11, 23, 41, 59). Therefore, we compared the transcriptomes of ATMs during obesity based solely on CD9 expression. Although CD9+ ATMs expressed higher levels of certain genes involved in lipid metabolism and uptake, we found that the loss of CD9 did not alter the expression of key lipid metabolism genes or lipid accumulation. Notably, palmitate robustly induced the expression of CD9 in BMDMs and sort-purified blood monocytes. Together, these findings suggest that CD9 does not drive LAM development but is instead upregulated as a consequence of macrophage metabolic activation. Further studies are needed to elucidate the molecules and mechanisms governing LAM development and metabolic activation, as well as the specific role of CD9 in these processes.
Crown-like structures are sites of close interactions between adipocytes and LAMs and are thought to facilitate the exchange of lipids, cytokines, and other signaling molecules (12, 32, 46, 60–63). Previous studies have demonstrated that ATMs adhere tightly to adipocytes within crown-like structures, partly through integrin-mediated interactions (52, 54). We identified CD9 as a key positive regulator of these interactions, as myeloid-specific deletion of CD9 disrupted ATM-adipocyte associations both in vitro and in vivo, leading to the loss of ATMs within the adipocyte fraction and a marked reduction in crown-like structures. Our data suggest that CD9 may contribute to this adhesive phenotype by positively regulating the expression of integrins, such as CD11c (Itgax) and CD51 (Itgav), which are critical for cell-cell and cell-ECM adhesion (31, 64, 65). Additionally, CD9 may directly facilitate cell-cell adhesion through direct interactions with partner molecules on adjacent cells (30, 66).
Importantly, our findings have methodological implications. A subset of CD9+ LAMs remains associated with adipocytes and partitions into the floating adipocyte fraction during standard adipose tissue processing, potentially leading to underestimation of this highly adherent ATM subset when only the stromal vascular fraction is analyzed. Conversely, as previously noted, adipocyte fractions may contain contaminating adherent ATMs (52), underscoring the need for careful interpretation of studies utilizing either fraction.
There is evidence that ATMs play important protective functions during adipose tissue remodeling through the phagocytosis and clearance of dead and dying adipocytes (10, 11). However, they also contribute to adipose tissue dysfunction through the secretion of proinflammatory and profibrotic mediators (10, 15–19). Although the specific role of CD9+ LAMs in adipose tissue fibrosis remains incompletely understood, CD9+ macrophages in the liver (SAMs) have been implicated in fibrosis promotion during steatohepatitis (36, 39, 40, 67). We found that CD9+ ATMs express genes involved in ECM remodeling, including those involved in both ECM synthesis and degradation. Metabolic activation with palmitate further induced the expression of fibrosis-associated genes such as Spp1, Lgals3, Pdgfb, and Mmp12 in a CD9-dependent manner. On the other hand, we did not observe an upregulation of other LAM/SAM markers, including Trem2 and Gpnmb. Prior work has demonstrated that type 3 cytokines can induce CD9+ macrophages that express high levels of Spp1, Gpnmb, Trem2, and Fabp5 (40). This suggests that additional signals, such as type 3 cytokines, may be required to drive the development of CD9+ LAMs in vivo. Moreover, our data suggests that integrin-mediated binding and signaling may contribute to the expression of a subset of fibrosis-associated genes, highlighting a potential mechanistic link between CD9’s role in modulating integrin function and its regulation of profibrotic transcriptional programs. Future work is needed to address the mechanisms by which CD9 modulates fibrosis-associated gene expression and effector molecules during metabolic activation.
Adipose tissue remodeling is an adaptive response to excess caloric intake. However, prolonged exposure to surplus nutrients can lead to dysregulated ECM deposition and metabolic dysfunction (1, 68, 69). Adipose tissue hypoxia and inflammatory infiltration contribute to pathogenic ECM accumulation and remodeling (16, 17, 68, 70). Macrophages can exert both profibrogenic and fibrolytic functions in tissue fibrosis (15–19, 71, 72). Our study suggests that CD9+ LAMs are primarily profibrogenic, despite expressing genes involved in both ECM formation and degradation. CD9+ LAMs may contribute to fibrosis through multiple mechanisms, including promoting the proliferation and activation of fibrogenic mesenchymal stromal cells via secretion of proinflammatory cytokines such as TNF-α and IL-6, as well as CD9-dependent factors such as PDGFβ and Spp1 (OPN) (19, 56, 73). Additionally, CD9+ LAMs may directly contribute to ECM deposition, as observed in other fibrotic diseases, suggested by the expression of ECM components in our RNA-Seq analysis of sorted CD9+ ATMs (74, 75). Indeed, interstitial tissue fibrosis colocalizes with crown-like structures during obesity, supporting the idea that these sites are enriched for profibrotic ATMs (16). Thus, the reduced eWAT fibrosis observed in CD9-MKO mice may reflect both diminished profibrotic activation and impaired localization of LAMs to crown-like structures.
Myeloid-specific deletion of CD9 improved multiple obesity-associated outcomes, including insulin resistance, hyperlipidemia, and ectopic lipid deposition. Notably, these effects were sex-specific, as male mice exhibited worse metabolic outcomes, whereas female mice were largely unaffected. Female mice are known to be more resistant to obesity-associated sequelae, such as insulin resistance and adipose tissue inflammation (49, 50). This suggests that CD9+ LAM accumulation may play a less prominent role in adipose tissue remodeling in female mice, at least at the time point we assessed.
Our findings demonstrate that myeloid-intrinsic CD9 contributes to eWAT dysfunction, a central driver of obesity-associated metabolic impairments (1, 68, 69). Given the conserved presence of CD9+ macrophages across tissues, it is plausible that CD9 similarly regulates macrophage activation in other organs, such as the liver. Future studies are needed to define the role of CD9 in macrophage biology across different tissue environments during obesity. Importantly, LysMCre yielded only partial deletion of CD9 in macrophages, raising the possibility that more complete CD9 ablation could produce even more pronounced effects on adipose remodeling and metabolic outcomes. In addition, because LysMCre-mediated deletion reduced CD9 expression not only in macrophages but also in monocytes and neutrophils, CD9 may still influence additional myeloid cell populations during obesity. Further investigation is therefore warranted to determine the broader contribution of CD9 to myeloid cell function in metabolic disease.
Therapeutic targeting of ATMs has shown limited success, likely because ATMs play both beneficial and pathological roles during obesity-associated adipose tissue remodeling. In this study, we found that CD9 selectively mediates the pathogenic features of LAMs, driving the formation of crown-like structures and promoting profibrotic activation while preserving beneficial lipid metabolic functions. Moreover, myeloid-specific CD9 deletion improved adipose tissue health and mitigated obesity-associated metabolic complications. Together, these findings identify CD9 as a promising therapeutic target with the potential to modulate pathogenic ATM activity while preserving essential homeostatic functions.
-
Methods
Sex as a biological variable. It is known that female mice are protected from adipose tissue inflammation and the development of obesity-associated sequelae. Therefore, the majority of experiments were performed using male mice. Metabolic and inflammatory phenotyping of female mice is shown in Supplemental Figure 11.
Mice. Mice were maintained under specific pathogen–free conditions in an animal facility at the Children’s Hospital of Philadelphia (CHOP), which has been accredited by the American Association for the Accreditation of Laboratory Animal Care. All studies were performed according to protocols the IACUC approved (24-001324). Experiments were performed using sex- and age-matched littermate controls. Cd9fl/fl mice were generated by our lab as described below. C57BL/6J (JAX stock 000664), LysMCre (B6.129P2-Lyz2tm1(cre)Ifo/J; JAX stock 004781), and Vav-iCre (B6.Cg-Commd10Tg(Vav1-icre)A2Kio/J; JAX stock 008610) mice were purchased from The Jackson Laboratory and subsequently bred in-house. All experimental mice were used between 10 and 18 weeks of age.
Generation of Cd9fl/fl mouse line. The Cd9fl/fl mouse line was generated on a C57BL/6J background by the University of Pennsylvania CRISPR-Cas9 Mouse Targeting Core. Sanger sequencing confirmed the insertion of LoxP sites. Mice were backcrossed to C57BL/6J at least 6 times before further breeding. Cd9fl/fl mice were then crossed to LysMCre mice (B6.129P2-Lyz2tm1(cre)Ifo). Cd9fl/fl mice were genotyped using the following primers: F: CGGTAGCCCTTGTGATTTA; R: CAAACTCGGCAGGCAAGAC. The Cd9fl/fl mouse line is available from David Hill’s lab upon request.
Diet studies. Mice were provided with ad libitum access to water and the indicated diet. An NCD (LabDiet, 5015) was provided to all mice until 6 weeks of age. For HFD studies, mice were switched to an irradiated diet (60 kcal% fat, Research Diets, D12492) from 6 to 18 weeks of age.
Tissue collection. Mice were euthanized by carbon dioxide inhalation followed by cervical dislocation. Transcardiac perfusion was performed with 10 mL of PBS, and tissues were excised and weighed. Tissues used for RNA-Seq were immediately flash-frozen in liquid nitrogen and stored at –80°C. Adipose tissue used for other purposes was collected in complete media (DMEM with 10% FBS, 10 mM HEPES, 1 mM sodium pyruvate, and penicillin/streptomycin) at 37°C.
Adipose tissue digestion. Adipose tissue was minced and digested for 40 minutes at 37°C with end-over-end rotation in 1 mg/mL collagenase type IV (Gibco, 17104019) and 0.1 mg/mL DNase I grade II (Roche, 10104159001) in complete media. Digested tissue was passed through a 100 μM cell strainer (CELLTREAT, 229485). After filtration, samples were centrifuged for 5 minutes at 450g. The adipocyte fraction was removed using a 1 mL pipette, and the remainder of the sample was used for isolation of the stromal vascular fraction.
The adipocyte fraction was washed with PBS 3 times. Following each wash, adipocyte fraction samples were left upright for 10 minutes to allow adipocytes to float. The floating fraction was collected with a 1 mL pipette and transferred to a new tube. The adipocyte fraction was then frozen in TRIzol (Invitrogen, 15596018) for qPCR or further processed for immunofluorescence imaging.
To isolate the stromal vascular fraction, samples were centrifuged at 450g for 5 minutes and washed with PBS. Red blood cells were lysed using ACK lysing buffer (Quality Biological, 118-156-721EA). Samples were frozen in TRIzol for qPCR or used for flow cytometry experiments described below.
Flow cytometry and fluorescence-activated cell sorting. The stromal vascular fraction was isolated as detailed above. Cell number and viability were quantified using trypan blue (Gibco, 15-250-061) staining and a TC20 Automated Cell Counter (Bio-Rad). Isolated cells were blocked and stained using fluorophore-conjugated antibodies, live-dead discrimination dyes, and lipid dyes listed in Supplemental Table 1. Single-stain controls and fluorescence-minus-one controls were used as necessary. ATMs for bulk RNA-Seq were sorted using a FACS Aria II (BD Biosciences). ATMs and blood monocytes for qPCR were sorted using an Aurora CS (Cytek). ATMs were cultured in complete media for 36 hours before being frozen in TRIzol for qPCR. Monocytes were cultured in complete media with BSA or palmitate for 24 hours before being frozen in TRIzol. Flow data were collected using an Aurora flow cytometer (Cytek) and analyzed using FlowJo v10 (BD Biosciences). Immune cells were gated as previously described (23, 32).
Immunofluorescence of adipocyte fraction samples. Adipocyte fractions were isolated as detailed above. The adipocyte fraction was fixed using 4% paraformaldehyde (FD Neurotechnologies, PF101) for 15 minutes at room temperature. The adipocyte fraction was then allowed to float for 10 minutes, collected into a new tube, and washed once with PBS. Cells were then blocked with CD16/32 (1:200) for 10 minutes in PBS. After blocking, cells were stained with Hoechst 33342 (Thermo Fisher Scientific, 66249, 1:250), BODIPY 493/503 (Invitrogen 1:1,000), F4/80 (APC BD Biosciences, 1:500), and CD9 (PE BioLegend, 1:500) at room temperature for 30 minutes in the dark. Antibody information is listed in Supplemental Table 1. Samples were washed 3 additional times with PBS before imaging. Adipocyte fraction samples were imaged using a Zeiss Observer Widefield Microscope. Image analysis was conducted in Fiji.
BMDM studies. BM was collected from male mice between 10 and 15 weeks of age. BM was isolated by flushing murine femurs and tibias with cold PBS or by crushing with a mortar and pestle. Red blood cells were lysed with ACK lysis buffer, and samples were filtered through a 70 μm cell strainer (MTC Bio, C4070). BMDMs were generated by culturing BM with 10 ng/mL of M-CSF (PeproTech, 315-03) in complete media for 7 days in 10 cm non-tissue treated polystyrene plates (VWR, 89038-968). Additional media containing M-CSF was added on day 4.
Differentiated cells were harvested by scraping with ice-cold PBS. For metabolic activation studies, BMDMs were cultured with 300 μM BSA-conjugated palmitate or BSA for 24 hours. BSA-conjugated palmitate was prepared as follows: solutions of BSA (0.34 mM, Ultra Fatty Acid-Free BSA, Roche, 03117405001) and the sodium salts of palmitate (2 mM, Sigma-Aldrich, P9767-5G) were prepared in 150 mM NaCl in water. BSA and palmitate were combined 1:1 (v/v) and allowed to conjugate for 1 hour at 37°C with gentle stirring. The pH was adjusted to approximately 7.4, and aliquots were frozen in glass vials. The final concentrations were 1 mM palmitate/0.17 mM BSA, or a 6:1 molar ratio of palmitate/BSA.
For experiments with integrin blockade, BMDMs were treated with antibodies against integrins or isotype controls at the time of treatment with BSA or palmitate. BMDMs were then cultured for 24 hours prior to harvesting for qPCR. To assess the role of CD11c in BMDM gene expression, cells were treated with 1:100 anti-CD11c (Bio X Cell, BE0038, clone N418) or polyclonal Armenian hamster IgG isotype control (Bio X Cell, BE0091). To assess the role of CD51 in BMDM gene expression, cells were treated with 1:100 anti-CD51 (BioLegend, 104102, clone RMV-7) or rat IgG1κ isotype control (BioLegend, 400401).
3T3-L1 BMDM adhesion assays. First, 3T3-L1 cells (ATTC, CL-173) were grown in 24-well tissue culture-treated plates (CELLTREAT, 229124). Cells were plated at a concentration of 2 × 104 cells per well and grown until they reached 100% confluency. Cells were incubated for an additional 48 hours before adding induction media (DMEM with 10% FBS, penicillin/streptomycin, 500 μM 3-isobutyl-1-methylxanthine, 1 μM dexamethasone, and 3 μg/mL insulin). After 48 hours, induction media was replaced with adipocyte maintenance media (DMEM with 10% FBS, penicillin/streptomycin, 3 μg/mL insulin). Maintenance media was replaced every 2–3 days until adipocytes were fully differentiated (8–12 days following the addition of induction media).
BMDMs were harvested by incubation with ice-cold PBS followed by gentle scraping. BMDMs were then incubated with 5 μM CFSE (Thermo Fisher Scientific, 50-591-407) for 15 minutes at 37°C and washed with PBS. Next, 1 × 105 BMDMs were added to a 24-well plate containing differentiated 3T3-L1 adipocytes in complete media. Cells were incubated at 37°C for 30 minutes to allow BMDMs to adhere. Media was then removed and wells were washed 3 times with PBS. For each group, a subset of wells was not washed, and these samples were used as unwashed controls. Wells were then imaged using an EVOS FL auto microscope (Thermo Fisher Scientific). CFSE+ cells were counted using CellProfiler (Broad Institute). The number of CFSE+ cells was then divided by the average number of CFSE+ cells in the unwashed controls of the same genotype to calculate the percentage of adherent cells.
Histology. eWAT samples were fixed overnight in 4% buffered paraformaldehyde solution at 4°C, followed by ethanol dehydration. Dehydrated samples were sent to the University of Pennsylvania Veterinary School Comparative Pathology Core to perform H&E and Picrosirius red staining. Histological images were acquired using the EVOS FL auto microscope (Thermo Fisher Scientific). Adipocyte size was measured using the Adiposoft plugin in the Fiji application. Picrosirius red staining was quantified using Fiji. For female mice, crown-like structures were quantified manually in blinded H&E images. Histological analyses were performed on 3–5 images per biological replicate.
Immunofluorescence staining was performed using the following primary antibodies: anti-F4/80 (Cell Signaling Technology, 70076), anti-CD68 (Cell Signaling Technology, 97778), anti-CD11c (Cell Signaling Technology, 97585), anti-CD51 (Abcam, ab179475), anti-TREM2 (Abcam, ab305103), anti-OPN/SPP1 (Cell Signaling Technology, 88742), anti-CD9 (Abcam, ab307085), anti-Perilipin-1 (Cell Signaling Technology, 3470), and DAPI (Quanterix, FP1490). Staining was performed using the BOND RXm (Leica, DS9800). Immunofluorescence images were acquired on the EVOS FL auto microscope or Zeiss Observer 7. In male mice, crown-like structures were quantified manually in blinded F4/80 immunofluorescence images. Analyses were performed on 5 images per biological replicate.
For high-resolution imaging and 3D reconstruction, slides were imaged on a Leica TCS SP8 confocal microscope using a 63× oil immersion objective (NA = 1.4). Images were acquired at 1,024 × 1,024 pixels as Z-stacks (30–40 steps of 0.12 μm per field of view). Obtained images were deconvoluted using Leica Lightning Deconvolution software. Representative images represent a single plane, with brightness and contrast adjusted equivalently across samples in ImageJ (NIH). All fluorescent staining (blue, green, red, and far-red channels) were acquired using HyD detectors in the standard mode with 100% gain, and 3D image reconstruction was performed with Imaris software (v8.1.2, Oxford Instruments). Adipocyte, macrophage, and CD9 surfaces were created using the Surfaces tool with automatic settings and thresholding based on the fluorescent signals from the anti-PLIN1, anti-F4/80, or anti-CD9 antibodies, respectively.
Blood glucose measurements. For fed and fasting blood glucose, blood was collected via the tail vein. Glucose was measured using blood glucose strips and a BG1000 glucose monitor (Clarity Diagnostics) under fed or fasting (~16 hours) conditions. For intraperitoneal glucose tolerance tests, mice were fasted overnight (~16 hours) in a clean cage with ALPHA-Dri bedding (Lab Supply). Mice were weighed and given an intraperitoneal injection of glucose solution at a dose of 1 gram per kg body weight. Blood glucose levels were measured before (0 minutes) and after (15, 30, 60, 90, and 120 minutes) glucose administration.
Metabolic plasma studies. Blood was collected via tail vein under fasting (~16 hours) and refed (4 hours) conditions. Samples were collected in Microvette CB 300 lithium heparin tubes (Sarstedt) on ice. Samples were centrifuged at 13,000g for 10 minutes at 4°C. Plasma was removed and frozen at –80°C until further analysis. Serum insulin levels were measured using the Ultrasensitive Mouse Insulin ELISA kit (Crystal Chem, 90080) on an Epoch Spectrophotometer (Agilent BioTek). Plasma triglycerides were measured using the Infinity Triglyceride Reagent (Thermo Fisher Scientific, TR22421) on an Epoch Spectrophotometer (Agilent BioTek). Plasma non-esterified fatty acids were measured using a Fujifilm HR Series NEFA-HR(2) (999-34691, 995-34791, 999-34891, 999-35191). Plasma glycerol was measured using a free glycerol assay kit (Cell Biolabs, STA-398).
Indirect calorimetry. Metabolic cage experiments were performed with a Promethion CORE metabolic cage system (Sable Systems International). Mice were acclimated in the Promethion metabolic cages at 22°C for 48 hours before the recording period. Oxygen consumption (VO2) and carbon dioxide production (VCO2) were measured by indirect calorimetry every 5 minutes and used to calculate energy expenditure using the Weir equation (energy expenditure = 3.941 kcal/L × VO2 + 1.106 kcal/L × VCO2) (76). Food intake, water intake, and body mass were measured in the cages gravimetrically. Lean body mass was measured using an EchoMRI. Data were analyzed in CalR2 (https://calrapp.org/) (77). ANCOVA testing was performed in CalR2, using pretreatment lean mass as a covariate.
RT-qPCR. RNA was isolated using TRIzol (Invitrogen, 15596018) according to the manufacturer’s instructions. Reverse transcription was performed using the Verso cDNA Synthesis kits (Thermo Fisher Scientific, AB1453B) according to the manufacturer’s instructions. qPCR was performed using PowerUp SYBR Green Master Mix on a QuantStudio 7K Real-Time PCR System (Applied Biosystems, A25742). Primer sequences are listed in Supplemental Table 2. All reactions were run in duplicate or triplicate. For BMDM experiments, gene expression was calculated as 2–(Ctgene–CtHprt). For analyses of sorted ATMs, stromal vascular fraction, and adipocyte fractions, each sample’s normalization factor (NF) was calculated as the geometric mean of the Ct values for the genes Hprt, Rpl13a, Sdha, Actb, and B2m. Gene expression was calculated as 2–(Ctgene–NF).
RNA-Seq. For bulk RNA-Seq of CD9+ and CD9– ATMs, libraries were prepared from sort-purified cell populations as previously described (23). Briefly, SMARTer Stranded RNA-Seq kit (Clontech) and TruSeq Stranded mRNA Library prep kits (Illumina, 20020594) were used according to the manufacturer’s instructions. RNA-Seq libraries were sequenced on an Illumina HiSeq 2000.
For bulk RNA-Seq of adipose tissue, total RNA was isolated from snap-frozen tissue samples. RNA extraction was performed using the Maxwell RSC simply RNA tissue kit (Promega, AD1340). Briefly, tissue was homogenized in Thioglycerol Homogenization solution using a Maxwell RSC instrument (Promega). RNA was quantified using a NanoDrop 8000 spectrophotometer (Thermo Fisher Scientific) and an Agilent 4200 bioanalyzer system. RNA libraries were prepared using the standard Illumina protocol. RNA-Seq libraries were sequenced as paired-end, 75–100 bp read length on an Illumina NovaSeq 6000 with an SP 200 cycle kit by the CHOP Center for Applied Genomics.
Bulk RNA-Seq reads were aligned to the GRCmm39/mm10 genome using the nf-core RNA-Seq pipeline version 3.14.0 (78). In brief, FastQC was used to assess the quality of reads (79), and Trim Galore! was used for adapter and quality trimming (80). Alignment was performed with STAR version 2.7.10a (81) followed by quantification with RSEM version 1.3.1 (82). Duplicates were removed using Picard MarkDuplicates (83). Low-expression genes were filtered with the R package edgeR, followed by differential expression analysis (84). Genes were considered to have significant differential expression with an FDR less than 0.05. Gene set enrichment analysis was performed with Reactome, GO, and Wikipathways gene sets using the R package WebGestaltR with log2 FC as the ranking metric (85). Pathways with an adjusted P value of greater than 0.05 were considered significantly enriched.
Statistics. The method for analysis of RNA-Seq data is described above. GraphPad Prism software was used for graphing and statistical analysis. Data are presented as mean ± SEM unless otherwise stated. Comparisons between 2 groups were performed using an unpaired 2-tailed Student’s t test. An F test was used to compare the variance of the tested groups; groups with significantly different variances were compared using Welch’s t test. Comparisons between 3 or more groups were performed by 1- or 2-way ANOVA with Fisher’s LSD test for multiple comparisons. Outliers were identified by ROUT with Q = 1%. A P value of less than 0.05 was considered significant.
Study approval. Animal experiments were performed in accordance with ethical regulations and protocols approved by the IACUCs of CHOP and the Perelman School of Medicine at the University of Pennsylvania.
Data availability. Raw bulk RNA-Seq data are available in NCBI’s Gene Expression Omnibus (GEO GSE113594 and GSE311351). Raw data and values for all data points in graphs are reported in the Supporting Data Values File.
-
Author contributions
Conceptualization and design of the research study were performed by JC, SJM, MAL, and DAH. Experiments and data acquisition were conducted by JC, ND, DVM, SJM, KMS, DP, RLC, and JM. Data analysis was performed by JC, ND, and DVM. Data curation and visualization were performed by JC. Writing and preparation of the original draft were performed by JC and DAH. Project administration was performed by JC, ND, and DAH. Funding was acquired by RJ, DMT, MAL, and DAH. All authors have reviewed and approved the final manuscript.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- NIH grants K08 DK116668 and R01 HL162715 (to DAH).
- Penn Diabetes Research Center (P30DK019525) pilot grant (to DAH).
- American College of Allergy, Asthma, and Immunology Junior Faculty grant (to DAH).
- CHOP Research Institute (to DAH).
- University of Pennsylvania Theo Wilson Graduate Research Fellowship (to JC).
- NIH grant 5T32 DK007314-43 (to JC).
- American Heart Association Predoctoral Fellowship 24PRE1191220 (to JC).
- NIH grant R01 DK49780 (to MAL).
- The JPB Foundation (to MAL).
- NIH grant T32 GM007170 (to JC and SJM).
- University of Pennsylvania Michael Brown Graduate Research Fellowship (to SJM).
- NIH grant R35HL166663 (to RJ).
- Penn Rodent Metabolic Phenotyping Core (RRID: SCR_022427), supported in part by NIH grant S10-OD025098, the Cox Institute, and the Penn Institute for Diabetes, Obesity, and Metabolism.
- Penn Vet Comparative Pathology Core, supported in part by the Abramson Cancer Center Support grant (P30 CA016520) and NIH grant S10 OD023465-01A1.
-
Acknowledgments
We acknowledge the Penn Diabetes Research Center, Penn CRISPR/Cas9 Mouse Targeting Core, Penn Vet Comparative Pathology Core, CHOP Flow Cytometry Core, and CHOP Center for Applied Genomics for supporting this work. We thank the Penn Rodent Metabolic Phenotyping Core for measuring mouse plasma analytes and conducting indirect calorimetry studies. Immunofluorescence was performed by the Penn Vet Comparative Pathology Core, which is partially subsidized by the Abramson Cancer Center support grant (P30 CA016520); the Aperio Versa 200 scanner used for imaging was acquired through an NIH shared instrumentation grant (S10 OD023465-01A1); and the Leica BOND RXm instrument used for immunofluorescence was acquired through the Penn Vet IIZD Core pilot grant opportunity 2022.
Address correspondence to: David A. Hill, Division of Allergy and Immunology, Children’s Hospital of Philadelphia, Abramson Research Building, 1208B, 3615 Civic Center Blvd., Philadelphia, Pennsylvania, 19104, USA. Phone: 215.590.2549; Email: hilld3@chop.edu.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2026, Chini et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: JCI Insight. 2026;11(6):e193837.https://doi.org/10.1172/jci.insight.193837.
-
References
- Sun K, et al. Adipose tissue remodeling and obesity. J Clin Invest. 2011;121(6):2094–2101.
- Della Guardia L, Shin AC. Obesity-induced tissue alterations resist weight loss: a mechanistic review. Diabetes Obes Metab. 2024;26(8):3045–3057.
- Schmitz J, et al. Obesogenic memory can confer long-term increases in adipose tissue but not liver inflammation and insulin resistance after weight loss. Mol Metab. 2016;5(5):328–339.
- Weinstock A, et al. Single-cell RNA sequencing of visceral adipose tissue leukocytes reveals that caloric restriction following obesity promotes the accumulation of a distinct macrophage population with features of phagocytic cells. Immunometabolism. 2019;1:e190008.
- Zamarron BF, et al. Macrophage proliferation sustains adipose tissue inflammation in formerly obese mice. Diabetes. 2017;66(2):392–406.
- Cottam MA, et al. Multiomics reveals persistence of obesity-associated immune cell phenotypes in adipose tissue during weight loss and weight regain in mice. Nat Commun. 2022;13(1):2950.
- Strissel KJ, et al. Adipocyte death, adipose tissue remodeling, and obesity complications. Diabetes. 2007;56(12):2910–2918.
- Lindhorst A, et al. Adipocyte death triggers a pro-inflammatory response and induces metabolic activation of resident macrophages. Cell Death Dis. 2021;12(6):579.
- Kratz M, et al. Metabolic dysfunction drives a mechanistically distinct proinflammatory phenotype in adipose tissue macrophages. Cell Metab. 2014;20(4):614–625.
- Coats BR, et al. Metabolically activated adipose tissue macrophages perform detrimental and beneficial functions during diet-induced obesity. Cell Rep. 2017;20(13):3149–3161.
- Jaitin DA, et al. Lipid-associated macrophages control metabolic homeostasis in a Trem2-dependent manner. Cell. 2019;178(3):686–698.
- Flaherty SE, et al. A lipase-independent pathway of lipid release and immune modulation by adipocytes. Science. 2019;363(6430):989–993.
- Lumeng CN, et al. Obesity induces a phenotypic switch in adipose tissue macrophage polarization. J Clin Invest. 2007;117(1):175–184.
- Lumeng CN, et al. Phenotypic switching of adipose tissue macrophages with obesity is generated by spatiotemporal differences in macrophage subtypes. Diabetes. 2008;57(12):3239–3246.
- Askia H, et al. Obesity-elicited macrophages shape CD9hi progenitor fate to promote adipose tissue fibrosis and dysfunction [Preprint]. https://doi.org/10.1101/2023.09.26.559540 Posted on bioRxiv September 28, 2023.
- Tanaka M, et al. Macrophage-inducible C-type lectin underlies obesity-induced adipose tissue fibrosis. Nat Commun. 2014;5(1):4982.
- Takahashi M, et al. Loss of myeloid lipoprotein lipase exacerbates adipose tissue fibrosis with collagen VI deposition and hyperlipidemia in leptin-deficient obese mice. J Biol Chem. 2022;298(9):102322.
- Bourlier V, et al. TGFbeta family members are key mediators in the induction of myofibroblast phenotype of human adipose tissue progenitor cells by macrophages. PLoS One. 2012;7(2):e31274.
- Keophiphath M, et al. Macrophage-secreted factors promote a profibrotic phenotype in human preadipocytes. Mol Endocrinol. 2009;23(1):11–24.
- Xu X, et al. Obesity activates a program of lysosomal-dependent lipid metabolism in adipose tissue macrophages independently of classic activation. Cell Metab. 2013;18(6):816–830.
- Park M, et al. Triggering receptor expressed on myeloid cells 2 (TREM2) promotes adipogenesis and diet-induced obesity. Diabetes. 2015;64(1):117–127.
- Sharif O, et al. Beneficial metabolic effects of TREM2 in obesity are uncoupled from its expression on macrophages. Diabetes. 2021;70(9):2042–2057.
- Hill DA, et al. Distinct macrophage populations direct inflammatory versus physiological changes in adipose tissue. Proc Natl Acad Sci U S A. 2018;115(22):E5096–E5105.
- Huang W, et al. CD9 tetraspanin interacts with CD36 on the surface of macrophages: a possible regulatory influence on uptake of oxidized low density lipoprotein. PLoS One. 2011;6(12):e29092.
- Kaji K, et al. Functional association of CD9 with the Fc gamma receptors in macrophages. J Immunol. 2001;166(5):3256–3265.
- Machado-Pineda Y, et al. CD9 controls integrin α5β1-mediated cell adhesion by modulating its association with the metalloproteinase ADAM17. Front Immunol. 2018;9:2474.
- Mathieu M, et al. Specificities of exosome versus small ectosome secretion revealed by live intracellular tracking of CD63 and CD9. Nat Commun. 2021;12(1):4389.
- Umeda R, et al. Structural insights into tetraspanin CD9 function. Nat Commun. 2020;11(1):1606.
- Parthasarathy V, et al. Distinct roles for tetraspanins CD9, CD63 and CD81 in the formation of multinucleated giant cells. Immunology. 2009;127(2):237–248.
- Reyes R, et al. Tetraspanin CD9: a key regulator of cell adhesion in the immune system. Front Immunol. 2018;9:863.
- Peddibhotla SSD, et al. Tetraspanin CD9 links junctional adhesion molecule-A to αvβ3 integrin to mediate basic fibroblast growth factor-specific angiogenic signaling. Mol Biol Cell. 2013;24(7):933–944.
- Paneru BD, et al. Myeloid-derived miR-6236 potentiates adipocyte insulin signaling and prevents hyperglycemia during obesity. Nat Commun. 2024;15(1):5394.
- Willemsen L, de Winther MP. Macrophage subsets in atherosclerosis as defined by single-cell technologies. J Pathol. 2020;250(5):705–714.
- McArdle S, et al. Migratory and dancing macrophage subsets in atherosclerotic lesions. Circ Res. 2019;125(12):1038–1051.
- Tran S, et al. Impaired Kupffer cell self-renewal alters the liver response to lipid overload during non-alcoholic steatohepatitis. Immunity. 2020;53(3):627–640.
- Remmerie A, et al. Osteopontin expression identifies a subset of recruited macrophages distinct from Kupffer cells in the fatty liver. Immunity. 2020;53(3):641–657.
- Xiong X, et al. Landscape of intercellular crosstalk in healthy and NASH liver revealed by single-cell secretome gene analysis. Mol Cell. 2019;75(3):644–660.
- Daemen S, et al. Dynamic shifts in the composition of resident and recruited macrophages influence tissue remodeling in NASH. Cell Rep. 2021;34(2):108626.
- Ramachandran P, et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature. 2019;575(7783):512–518.
- Fabre T, et al. Identification of a broadly fibrogenic macrophage subset induced by type 3 inflammation. Sci Immunol. 2023;8(82):eadd8945.
- Stansbury CM, et al. A lipid-associated macrophage lineage rewires the spatial landscape of adipose tissue in early obesity. JCI Insight. 2023;8(19):e171701.
- Chakarov S, et al. Two distinct interstitial macrophage populations coexist across tissues in specific subtissular niches. Science. 2019;363(6432):eaau0964.
- Nickl B, et al. Anti-inflammatory role of Gpnmb in adipose tissue of mice. Sci Rep. 2021;11(1):19614.
- Weisberg SP, et al. Obesity is associated with macrophage accumulation in adipose tissue. J Clin Invest. 2003;112(12):1796–1808.
- Xu H, et al. Chronic inflammation in fat plays a crucial role in the development of obesity-related insulin resistance. J Clin Invest. 2003;112(12):1821–1830.
- Murano I, et al. Dead adipocytes, detected as crown-like structures, are prevalent in visceral fat depots of genetically obese mice. J Lipid Res. 2008;49(7):1562–1568.
- Clausen BE, et al. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 1999;8(4):265–277.
- Ogilvy S, et al. Transcriptional regulation of vav, a gene expressed throughout the hematopoietic compartment. Blood. 1998;91(2):419–430.
- Ahnstedt H, et al. Sex differences in adipose Tissue CD8+ T cells and regulatory T cells in middle-aged mice. Front Immunol. 2018;9:659.
- Maric I, et al. Sex and species differences in the development of diet-induced obesity and metabolic disturbances in rodents. Front Nutr. 2022;9:828522.
- Stranahan AM, et al. Sex differences in adipose tissue distribution determine susceptibility to neuroinflammation in mice with dietary obesity. Diabetes. 2022;72(2):245–260.
- La E, et al. Tight association between macrophages and adipocytes in obesity: implications for adipocyte preparation. Obesity (Silver Spring). 2014;22(5):1246–1255.
- Feng D, et al. High-fat diet-induced adipocyte cell death occurs through a cyclophilin D intrinsic signaling pathway independent of adipose tissue inflammation. Diabetes. 2011;60(8):2134–2143.
- Chung KJ, et al. A self-sustained loop of inflammation-driven inhibition of beige adipogenesis in obesity. Nat Immunol. 2017;18(6):654–664.
- Emont MP, et al. A single-cell atlas of human and mouse white adipose tissue. Nature. 2022;603(7903):926–933.
- Rabhi N, et al. Obesity-induced senescent macrophages activate a fibrotic transcriptional program in adipocyte progenitors. Life Sci Alliance. 2022;5(5):e202101286.
- Ganguly S, et al. Lipid-associated macrophages’ promotion of fibrosis resolution during MASH regression requires TREM2. Proc Natl Acad Sci U S A. 2024;121(35):e2405746121.
- Onogi Y, et al. Pro-inflammatory macrophages coupled with glycolysis remodel adipose vasculature by producing platelet-derived growth factor-B in obesity. Sci Rep. 2020;10(1):670.
- Félix I, et al. Single-cell proteomics reveals the defined heterogeneity of resident macrophages in white adipose tissue. Front Immunol. 2021;12:719979.
- Cinti S, et al. Adipocyte death defines macrophage localization and function in adipose tissue of obese mice and humans. J Lipid Res. 2005;46(11):2347–2355.
- Wang L, et al. Targeting ERβ in macrophage reduces crown-like structures in adipose tissue by inhibiting osteopontin and HIF-1α. Sci Rep. 2019;9(1):15762.
- Altintas MM, et al. Mast cells, macrophages, and crown-like structures distinguish subcutaneous from visceral fat in mice. J Lipid Res. 2011;52(3):480–488.
- Brestoff JR, et al. Intercellular mitochondria transfer to macrophages regulates white adipose tissue homeostasis and is impaired in obesity. Cell Metab. 2021;33(2):270–282.
- Sándor N, et al. CD11c/CD18 dominates adhesion of human monocytes, macrophages and dendritic cells over CD11b/CD18. PLoS One. 2016;11(9):e0163120.
- Lin X, et al. Omentin-1 modulates macrophage function via integrin receptors αvβ3 and αvβ5 and reverses plaque vulnerability in animal models of atherosclerosis. Front Cardiovasc Med. 2021;8:757926.
- Reyes R, et al. Different states of integrin LFA-1 aggregation are controlled through its association with tetraspanin CD9. Biochim Biophys Acta. 2015;1853(10 pt a):2464–2480.
- Lee K-J, et al. Hepatic TREM2+ macrophages express matrix metalloproteinases to control fibrotic scar formation. Immunol Cell Biol. 2023;101(3):216–230.
- Gliniak CM, et al. Adipose tissue fibrosis: the unwanted houseguest invited by obesity. J Endocrinol. 2023;259(3):e230180.
- Marcelin G, et al. Adipose tissue fibrosis in obesity: etiology and challenges. Annu Rev Physiol. 2022;84(1):135–155.
- Dahdah N, et al. Interrelation of adipose tissue macrophages and fibrosis in obesity. Biochem Pharmacol. 2024;225:116324.
- Chen Q, et al. Resident macrophages restrain pathological adipose tissue remodeling and protect vascular integrity in obese mice. EMBO Rep. 2021;22(8):e52835.
- Jiang Y, et al. Macrophages in organ fibrosis: from pathogenesis to therapeutic targets. Cell Death Discov. 2024;10(1):487.
- Marcelin G, et al. A PDGFRα-mediated switch toward CD9high adipocyte progenitors controls obesity-induced adipose tissue fibrosis. Cell Metab. 2017;25(3):673–685.
- Simões FC, et al. Macrophages directly contribute collagen to scar formation during zebrafish heart regeneration and mouse heart repair. Nat Commun. 2020;11(1):600.
- Schnoor M, et al. Production of type VI collagen by human macrophages: a new dimension in macrophage functional heterogeneity. J Immunol. 2008;180(8):5707–5719.
- Weir JBDV. New methods for calculating metabolic rate with special reference to protein metabolism. J Physiol. 1949;109(1-2):1–9.
- Mina AI, et al. CalR: a web-based analysis tool for indirect calorimetry experiments. Cell Metab. 2018;28(4):656–666.
- Ewels PA, et al. The nf-core framework for community-curated bioinformatics pipelines. Nat Biotechnol. 2020;38(3):276–278.
- FastQC: A Quality Control tool for High Throughput Sequence Data; 2010. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
- Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17(1):10–12.View this article via: CrossRef Google Scholar
- Dobin A, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29(1):15–21.
- Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011;12(1):323.
- Picard. Github Repository. Broad Institute; 2019. http://broadinstitute.github.io/picard/.
- Robinson MD, et al. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26(1):139–140.
- Elizarraras JM, et al. WebGestalt 2024: faster gene set analysis and new support for metabolomics and multi-omics. Nucleic Acids Res. 2024;52(w1):W415–W421.
-
Version history
- Version 1 (February 10, 2026): In-Press Preview
- Version 2 (March 23, 2026): Electronic publication










